-
PDF
- Split View
-
Views
-
Cite
Cite
Curtis J. Donskey, Antibiotic Regimens and Intestinal Colonization with Antibiotic-Resistant Gram-Negative Bacilli, Clinical Infectious Diseases, Volume 43, Issue Supplement_2, September 2006, Pages S62–S69, https://doi.org/10.1086/504481
Close - Share Icon Share
Abstract
The intestinal tract provides an important reservoir for antibiotic-resistant gram-negative bacilli, including Enterobacteriaceae species, Pseudomonas aeruginosa, and Acinetobacter baumannii. Selective pressure exerted by antibiotics plays a crucial role in the emergence and dissemination of these pathogens. Many classes of antibiotics may promote intestinal colonization by health care-associated gram-negative bacilli, because the organisms are often multidrug resistant. Antibiotics may inhibit colonization by gram-negative pathogens that remain susceptible, but the benefits of this effect are often limited because of the emergence of resistance. Antibiotic formulary alterations and standard infection control measures have been effective in controlling outbreaks of colonization and infection with antibiotic-resistant gram-negative pathogens. Additional research is needed to clarify the role of strategies such as selective decontamination of the digestive tract and decontamination of environmental surfaces and of patients' skin and wounds.
The intestinal tract provides an important reservoir for antibiotic-resistant gram-negative bacilli, including Enterobacteriaceae species, Pseudomonas aeruginosa, and Acinetobacter species [1–9]. Although most patients who are colonized with these organisms remain asymptomatic, infections may occur because of translocation across the intestinal lining or as a consequence of fecal contamination of wounds or devices [2, 9]. Several studies have shown that intestinal colonization by gram-negative bacilli often precedes the onset of infection [1–4]. Fecal shedding onto patients' skin and environmental surfaces contributes to nosocomial transmission of antibiotic-resistant gram-negative pathogens [9]. Finally, the intestinal tract provides an important site for transfer of genes conferring antibiotic resistance [10].
Selective pressure exerted by antibiotics plays a crucial role in the emergence and dissemination of antibiotic-resistant microorganisms. This review is an examination of the effects of antibiotic treatment on colonization of the intestinal tract with antibiotic-resistant gram-negative bacilli. Findings from studies involving animal models and healthy human volunteers are used to illustrate general concepts regarding the effects of antibiotics on colonization with pathogens, and the applicability of these concepts to clinical settings and implications for control of antibiotic-resistant gram-negative bacilli are discussed.
Colonization Resistance
The indigenous bacteria of the colon provide an important host-defense mechanism by inhibiting colonization by potentially pathogenic microorganisms. This defense mechanism, termed “colonization resistance,” can be applied to the prevention of overgrowth by indigenous potential pathogens and the inhibition of colonization by exogenously introduced organisms [11]. Escherichia coli, a member of the indigenous colonic microflora, is normally maintained at relatively low population densities by the predominant anaerobic microflora [12]. Healthy humans ingesting small numbers of P. aeruginosa (102 cfu) do not develop detectable levels of organisms in stool; however, larger inocula (⩾106 cfu) have been found to result in shedding in stool for 1–6 days [13]. Foods such as salads may contain relatively large numbers of P. aeruginosa or Enterobacteriaceae species (103-104 cfu/serving) [14], a finding that could, in part, explain why P. aeruginosa has been detected in stool of patients with cancer who have not received prior antibiotic treatment [3, 14]. Experimental ingestion of exogenous E. coli by healthy humans does not typically result in persistent colonization [12, 15]; however, travelers to Mexico frequently acquire colonization by antibiotic-resistant strains of E. coli in the absence of prophylactic or therapeutic antibiotic treatment [16]. Acquisition in this setting could be due to repeated ingestion of large numbers of organisms and/or special properties of the ingested organisms (e.g., the ability to adhere to the mucosa) [17].
Antibiotics and Colonization with Gram-Negative Bacilli
Antibiotic selective pressure. Bacteria possess a remarkable ability to develop and acquire resistance to antibiotics. Among gram-negative bacilli, common mechanisms of resistance include modification of drug targets; production of inactivating enzymes, such as β-lactamases and aminoglycoside-modifying enzymes; efflux pumps; and alterations in outer membrane proteins that are associated with decreased permeability to antibiotics [18]. In general, antibiotic exposure is not thought to directly induce these resistance mechanisms. Rather, antibiotic therapy promotes proliferation of antibiotic-resistant gram-negative bacilli by exerting selective pressure (i.e., inhibition of competing microflora but not of resistant organisms). In individual patients, selective pressure may facilitate the emergence of new resistant mutants or of preexisting subpopulations of resistant organisms. For example, ceftazidime therapy may eliminate susceptible gram-negative bacilli while allowing expansion of the population of a new mutant of Klebsiella pneumoniae that harbors an extended-spectrum β-lactamase (ESBL) or of a preexisting subpopulation of Enterobacter species that constitutively hyperproduces chromosomal cephalosporinases [19–21]. Numerous clinical studies have documented the emergence of resistant gram-negative bacilli in association with the use of agents for treatment of gram-negative pathogens [4–7, 19–28]. Although this review focuses on the intestinal tract as a site for emergence of antibiotic-resistant gram-negative bacilli, resistant organisms also emerge frequently from other sites.
Once antibiotic-resistant gram-negative pathogens have emerged, antibiotics play a crucial role in their subsequent spread from patient to patient [9]. Antibiotic therapy may markedly reduce the number of exogenous bacteria that must be ingested to establish intestinal colonization [9, 11, 13]. Antibiotic-associated overgrowth of nosocomial pathogens and antibiotic-associated diarrhea contribute to increased shedding of organisms onto patients' skin or environmental surfaces [29, 30]. Organisms on skin or surfaces may then be acquired on health care workers' hands [9].
Because health care-associated gram-negative bacilli are often resistant to multiple classes of antibiotics, many different antibiotics may potentially facilitate colonization and dissemination of these pathogens. For example, third-generation cephalosporins, trimethoprim-sulfamethoxazole, ciprofloxacin, and aminoglycosides have all been associated with ESBL-producing gram-negative bacilli [7, 31]. In an outbreak of colonization and infection with ESBL-producing gram-negative bacilli in nursing homes, most patients had not received prior ceftazidime [7]. Rather, receipt of ciprofloxacin or trimethoprim-sulfamethoxazole was an independent risk factor for colonization; determinants of resistance to ceftazidime and trimethoprim-sulfamethoxazole were linked on a plasmid, whereas ciprofloxacin resistance was not directly linked to ceftazidime resistance [7]. Similarly, piperacillin-tazobactam, imipenem, aminoglycosides, vancomycin, and broad-spectrum cephalosporins have all been associated with piperacillin- and tazobactam-resistant P. aeruginosa [32].
Although antibiotics may promote proliferation of antibiotic-resistant gram-negative pathogens, these agents may also provide a protective effect if they have inhibitory activity [9, 33–36]. For example, Kaye et al. [33] found a trend toward reduced isolation of E. coli resistant to ampicillin and sulbactam in patients exposed to piperacillin-tazobactam, an agent with activity against many isolates that are resistant to ampicillin and sulbactam. In another study, fluoroquinolone exposure was found to be protective against isolation of broad-spectrum cephalosporin-resistant Enterobacter species. [34]. Paterson et al. [36] used oral norfloxacin to inhibit intestinal colonization by fluoroquinolone-susceptible ESBL-producing E. coli during an outbreak among patients undergoing liver transplantation. However, increasing rates of fluoroquinolone resistance among gram-negative bacilli, including ESBL producers, is likely to limit the protective effect of these agents [37, 38]. As was noted above, prior receipt of ciprofloxacin has been shown to be an independent risk factor for colonization with ESBL-producing gram-negative bacilli resistant to fluoroquinolones [7].
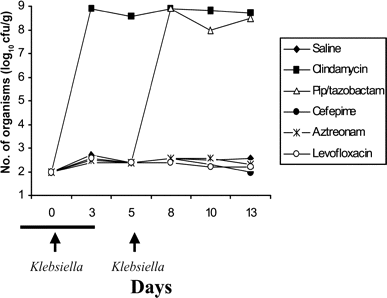
Animal models. Mouse models provide a useful means of directly comparing the effects of antibiotics on intestinal colonization with nosocomial pathogens. During treatment, antibiotics that are excreted into the intestinal tract may potentially inhibit gram-negative pathogens and competing indigenous microflora. After completion of treatment, the indigenous microflora recover over several days. Susceptibility to colonization by resistant gram-negative bacilli may persist during the recovery period. Figure 1 summarizes the findings of my group's mouse model studies examining establishment of colonization by ESBL-producing K. pneumoniae [39–41] (author's unpublished data). ESBL-producing K. pneumoniae (104 cfu) was administered orally once during and once 2 days after completion of subcutaneous antibiotic treatment. Antibiotics that disrupted the anaerobic microflora and possessed minimal activity against the K. pneumoniae isolate (e.g., clindamycin and linezolid) promoted colonization. An antibiotic that disrupted the anaerobic microflora and possessed significant activity against the K. pneumoniae isolate (piperacillin-tazobactam [MIC, 4 µg/mL]) inhibited the establishment of colonization during treatment but promoted overgrowth when exposure occurred during the period of recovery of the indigenous microflora. Antibiotics that did not disrupt the anaerobic microflora (e.g., cefepime, aztreonam, levofloxacin, and daptomycin) did not promote colonization. These findings are very similar to findings from mouse studies that examined the effects of antibiotics on colonization by vancomycin-resistant enterococci (VRE) and Clostridium difficile [9, 42]. Of note, however, Hentges et al. [43] found that oral streptomycin promoted Pseudomonas aeruginosa intestinal colonization and translocation across the intestinal lining to a greater degree than did oral clindamycin; oral streptomycin inhibits facultative anaerobes and obligate anaerobes, whereas clindamycin selectively inhibits anaerobes.
Effect of antibiotic treatment on establishment of intestinal colonization with extended-spectrum β-lactamase-producing Klebsiella pneumoniae in mice. Mice received subcutaneous antibiotic treatment from day -2 to day 3 (solid bar), and oral extended-spectrum β-lactamase-producing K. pneumoniae (10,000 cfu) was administered once during treatment and once 2 days after completion of treatment. Densities of extended-spectrum β-lactamase-producing K. pneumoniae in stool are shown. If extended-spectrum β-lactamase-producing K. pneumoniae were not detected, the lower limit of detection was assigned (2 log10 cfu/g). Pip, piperacillin.
Although animal models have limitations, these studies may raise important issues that deserve further study in patients. First, animal studies may implicate antibiotics, such as clindamycin and linezolid, that have not been associated with antibiotic-resistant gram-negative bacilli in clinical studies. It is plausible that these antibiotics may promote overgrowth of gram-negative pathogens in patients, because both promote overgrowth of Enterobacteriaceae species in healthy humans and because clindamycin promotes the emergence of new gram-negative bacilli [44]. Second, the promotion of piperacillin- and tazobactam-susceptible K. pneumoniae by piperacillin-tazobactam in mice exposed after the treatment period (figure 1) illustrates that adverse effects of antibiotics often extend beyond the period of treatment. Although recovery of the microflora of mice or healthy humans occurs within 1–2 weeks after antibiotic therapy is stopped [40], longer delays in recovery of protective indigenous microflora may occur in patients who have received multiple or prolonged courses of antibiotics. Finally, animal model studies suggest that antibiotics that do not disrupt the anaerobic microflora may be less likely to promote colonization by resistant gram-negative bacilli [39–41]. More data are needed to evaluate whether selective use of such agents may offer any advantage to patients.
Healthy volunteers. Many studies have been performed to evaluate the effect of different antibiotics on the indigenous intestinal microflora of humans [44]. Most studies have been performed in healthy volunteers, and clinical studies usually have involved monotherapy regimens in patients with mild to moderate severity of illness. As has been noted previously, these studies demonstrate that antibiotics that inhibit anaerobes without inhibiting Enterobacteriaceae species (e.g., clindamycin, linezolid, and oral vancomycin) may promote overgrowth of indigenous Enterobacteriaceae species and the emergence of new antibiotic-resistant gram-negative bacilli [44]. However, antibiotics that cause relatively little disruption of the anaerobic microflora (e.g., trimethoprim-sulfamethoxazole, cefadroxil, and ciprofloxacin) have also been shown to promote the emergence of antibiotic-resistant gram-negative bacilli [11, 44].
Although studies of healthy volunteers provide a useful reference regarding the potential impact of antibiotics on colonization with gram-negative bacilli, several factors may limit their applicability to clinical settings. First, patients are at increased risk of exposure to exogenous antibiotic-resistant gram-negative bacilli. The oropharynx of a hospitalized patient frequently becomes colonized with gram-negative bacilli that may be ingested. Nasogastric tubes may facilitate colonization of the oropharynx by P. aeruginosa that form biofilms on plastic surfaces [45]. Second, patients often receive proton pump inhibitors or histamine2 blockers that inhibit production of stomach acid [9]. These agents promote overgrowth of gram-negative bacilli in the stomach and facilitate passage of organisms into the small intestine [9]. Third, the colonic microflora of ill hospitalized patients may be altered in the absence of antibiotic therapy [46]. Finally, patients often receive multiple antibiotics concurrently or in sequence, which makes it difficult to determine the effects of individual agents.
One illustration of the discrepancy that may occur between healthy volunteers and patients is provided by studies of fluoroquinolone antibiotics. In healthy volunteers, most fluoroquinolones inhibit Enterobacteriaceae species but cause minimal disruption of intestinal anaerobes, and acquisition of fluoroquinolone-resistant gram-negative bacilli is uncommon [44, 47]. In contrast, numerous clinical studies have demonstrated that fluoroquinolone use may be associated with fluoroquinolone-resistant gram-negative pathogens [26, 28, 37, 38]. In addition to being at increased risk for exposure to fluoroquinolone-resistant gram-negative bacilli, hospitalized patients may have preexisting colonization with such organisms, which may expand in population during fluoroquinolone therapy. The fact that hospitalized patients often receive fluoroquinolones in combination with other antibiotics may contribute significantly to the potential for acquisition and overgrowth of fluoroquinolone-resistant gram-negative organisms. Joris et al. [48] illustrated this by giving low-dose ciprofloxacin monotherapy to healthy outpatient volunteers, followed by ciprofloxacin in combination with oral clindamycin. As shown in figure 2, no ciprofloxacin-resistant gram-negative bacilli were detected during ciprofloxacin monotherapy, but 3 of 5 subjects acquired new ciprofloxacin-resistant gram-negative bacilli when clindamycin was added [48].
Densities of indigenous and acquired facultative gram-negative bacilli in stool samples from 5 healthy volunteers receiving treatment with oral ciprofloxacin (20 mg/day) for 14 days followed by oral ciprofloxacin in combination with clindamycin (300 mg/day). Ciprofloxacin monotherapy eliminated indigenous Escherichia coli (dotted lines). No subjects acquired exogenous ciprofloxacin-resistant gram-negative bacilli during ciprofloxacin monotherapy, whereas 3 subjects did during the combination treatment period (solid circles). The acquired exogenous gram-negative bacilli included E. coli (2 strains) and Citrobacter freundii. Data are from Joris et al. [48].
Effect of antibiotics with activity against intestinal anaerobes. Antibiotics that inhibit intestinal anaerobes promote overgrowth of VRE in stools of colonized patients [29]. We tested the hypothesis that such antibiotic regimens may also promote overgrowth of coexisting gram-negative bacilli resistant to ceftazidime, ciprofloxacin, or piperacillin-tazobactam in stool of patients colonized with VRE [49]. As shown in figure 3A and 3B, therapy with antianaerobic antibiotic regimens was associated with an increased likelihood that an antibiotic-resistant gram-negative bacillus would be isolated, and, when present, the density of these organisms was higher during therapy than in the absence of antianaerobic therapy for at least 2 weeks. These findings suggest that efforts to limit the use of antianaerobic antibiotics could minimize the density of VRE and coexisting antibiotic-resistant gram-negative bacilli. However, it should be noted that many antibiotics with antianaerobic activity also have activity against gram-negative bacilli. For example, figure 3C shows the emergence of an isolate of E. coli resistant to piperacillin-tazobactam in stool of a patient during treatment with piperacillin-tazobactam, followed by elimination of the organism during meropenem therapy [49].
Effect of anti-anaerobic antibiotic regimens on the detection and density of antibiotic-resistant gram-negative bacilli in stool of patients colonized with vancomycin-resistant enterococci (VRE). A, Detection of gram-negative bacilli resistant to ceftazidime, ciprofloxacin, or piperacillin-tazobactam in stool of patients during therapy with anti-anaerobic antibiotic regimens and in the absence of such therapy for >1 month. B, Density of resistant gram-negative bacilli in stool during anti-anaerobic antibiotic therapy and in the absence of such therapy for >2 weeks (among patients with detectable antibiotic-resistant gram-negative bacilli). C, Effect of antibiotic therapy on the density of VRE (triangles) and piperacillin-tazobactam-resistant Escherichia coli (squares) in the stool of a patient. Antibiotic therapy included piperacillin-tazobactam, vancomycin, and metronidazole (regimen A); piperacillin-tazobactam and vancomycin (regimen B); and meropenem and linezolid (regimen C). The star indicates development of catheter-related bacteremia with an E. coli isolate with a susceptibility pattern identical to that of the stool isolate. Treatment with meropenem and linezolid (regimen C) resulted in suppression of the 2 pathogens to undetectable levels in stool (reprinted with permission from [49]).
Control Strategies For Antibiotic-Resistant Gram-Negative Bacilli
Infection control measures. Standard infection control measures play a crucial role in limiting the spread of antibiotic-resistant gram-negative bacilli. Because the hands of health care workers often become contaminated with gram-negative bacilli, efforts to improve adherence to hand hygiene are essential [50]. Alcohol-based hand-hygiene products are effective at eliminating gram-negative bacilli, and the use of these agents in combination with ongoing education has been associated with reductions in nosocomial infections [50]. For organisms resistant to multiple antibiotics or during outbreaks, contact precautions may be indicated [9]. Surveillance for stool carriage of antibiotic-resistant gram-negative bacilli may be helpful in some situations, because colonized patients often outnumber those with clinical isolates [6, 9]. Lucet et al. [6] used a multifaceted infection control intervention with no concurrent antibiotic formulary changes to control high rates of colonization and infection with endemic ESBL-producing organisms. Because multiple nosocomial pathogens resistant to antibiotics often coexist in the same patient populations [9], efforts to improve infection control practices may offer the additional benefit of limiting the spread of coexisting pathogens [6, 9, 51].
Reducing the burden of antibiotic-resistant gram-negative pathogens present on patients' skin and on environmental surfaces might potentially reduce transmission by decreasing the number of organisms acquired on health care workers' hands and by decreasing direct transmission from surfaces to patients [9]. A recent study found that patients harboring antibiotic-resistant gram-negative pathogens were less likely to have environmental contamination than were those harboring resistant gram-positive pathogens (4.9% vs. 24.7%) [52]. However, extensive environmental contamination with Acinetobacter baumannii has been described, and this organism survives for prolonged periods on surfaces [53]. In addition, P. aeruginosa may persistently contaminate moist areas, such as sinks [54]. Several quasi-experimental studies have suggested that environmental decontamination may be a useful adjunctive measure for control of these pathogens [53–58]. Borer et al. [59] found that using chlorhexidine to disinfect the skin of patients in intensive care units was an effective means of reducing skin contamination with A. baumannii. Similarly, Urban et al. [58] used polymyxin B to decontaminate wounds as an adjunctive control measure for A. baumannii.
Selective decontamination of the digestive tract. The goal of selective decontamination is to inhibit pathogens in the gastrointestinal tract without disturbing the anaerobic microflora [9]. Typically, nonabsorbed antibiotics are applied to the oropharynx and ingested orally with or without concurrent administration of intravenous antibiotics. In a recent study in the Netherlands, a selective decontamination regimen including parenteral cefotaxime and oropharyngeal and enteral tobramycin, colistin, and amphotericin was associated with a reduction in ventilator-associated pneumonia and mortality among patients in an intensive care unit [60]. Acceptance of selective decontamination in the United States has been limited, in part by concerns that the antibiotic regimens may promote overgrowth and infections with pathogens that are resistant to the agents being administered (e.g., VRE or methicillin-resistant Staphylococcus aureus) [60]. Jackson et al. [61] have shown that the selective decontamination regimen used in the Dutch study causes overgrowth and translocation of indigenous enterococci in rats. Therefore, caution is indicated if these regimens are to be used in settings in which VRE and methicillin-resistant S. aureus are endemic. As has been noted elsewhere, oral norfloxacin has been used to selectively decontaminate intestinal colonization by fluoroquinolone-susceptible ESBL-producing organisms [36]. Finally, oropharyngeal decontamination alone has been effective in preventing ventilator-associated pneumonia, suggesting that the combination of intravenous and orally ingested components of selective decontamination may not be necessary for prevention of this condition [62].
Measures to control antibiotic use. One strategy for the control of antibiotic-resistant gram-negative bacilli is to limit antibiotic use, in an effort to reduce antibiotic selective pressure. In a teaching hospital in Cleveland, we found that 30% of days of antibiotic therapy among patients who were not in intensive care were unnecessary, on the basis of standard guidelines or standard principles of infectious diseases [63]. These data demonstrate that significant reductions in antibiotic use in hospitals may be feasible. In addition, efforts to reduce the duration of antibiotic therapy have yielded promising results. For example, Chastre et al. [64] found that therapy for 8 days was as effective as therapy for 15 days for treatment of ventilator-associated pneumonia, and, among patients who developed recurrent infections, multidrug-resistant pathogens emerged less frequently in those who had received antibiotics for 8 days.
Another strategy to control antibiotic-resistant gram-negative bacilli is the implementation of formulary changes that involve restricting the use of antibiotics that have frequently been associated with particular pathogens. Third-generation cephalosporins have often been targeted for restriction, because they have been associated with ESBL-producing gram-negative bacilli as well as with multidrug-resistant P. aeruginosa and Acinetobacter species [32, 58, 65]. Substitution of piperacillin-tazobactam, cefepime, imipenem, or ticarcillin-clavulanate for third-generation cephalosporins has been associated with reductions in ESBL-producing organisms [65]. The rationale for such formulary alterations is that selective pressure due to third-generation cephalosporins is removed, and the other agents may suppress these pathogens in the intestinal tract or at other sites [9, 65]. Restriction of ciprofloxacin has also been associated with a decrease in ciprofloxacin resistance in P. aeruginosa isolates [66]. Unfortunately, substitution of one agent for another may lead to the emergence of resistance to the new agent [67]. Finally, it is noteworthy that the data supporting formulary alterations come primarily from quasi-experimental studies that should be interpreted cautiously because they are subject to a number of biases [68].
Conclusion
Antibiotic selective pressure has contributed to the emergence and spread of antibiotic-resistant gram-negative pathogens. The intestinal tract provides an important reservoir for dissemination of these pathogens. Adherence to standard infection control measures and good antibiotic stewardship are essential control strategies. Additional research is needed to clarify the potential utility of strategies such as selective decontamination of the digestive tract and decontamination of environmental surfaces and of patients' skin and wounds. Future directions for research should include efforts to develop novel technologies for control of antibiotic-resistant gram-negative pathogens. For example, we have demonstrated that oral administration of β-lactamase enzymes in conjunction with parenteral β-lactam antibiotics can degrade the portion of antibiotic that is excreted into the intestinal tract of mice, thereby preserving resistance against colonization by ESBL-producing K. pneumoniae [69]. Others have developed light-activated antimicrobial coatings for continuous disinfection of surfaces [70].
Acknowledgments
I thank Robert Bonomo and Marion Helfand for critical manuscript review.
Financial support. This work was supported by an Advanced Research Career Development Award from the Department of Veterans Affairs to C.J.D.
Potential conflicts of interest. C.J.D. has received research grant support from Elan, Merck, Cubist, and Ortho-McNeil and serves on the speakers' bureau of Elan and Ortho-McNeil. He is a consultant for Optimer Pharmaceuticals.




![Densities of indigenous and acquired facultative gram-negative bacilli in stool samples from 5 healthy volunteers receiving treatment with oral ciprofloxacin (20 mg/day) for 14 days followed by oral ciprofloxacin in combination with clindamycin (300 mg/day). Ciprofloxacin monotherapy eliminated indigenous Escherichia coli (dotted lines). No subjects acquired exogenous ciprofloxacin-resistant gram-negative bacilli during ciprofloxacin monotherapy, whereas 3 subjects did during the combination treatment period (solid circles). The acquired exogenous gram-negative bacilli included E. coli (2 strains) and Citrobacter freundii. Data are from Joris et al. [48].](https://oup.silverchair-cdn.com/oup/backfile/Content_public/Journal/cid/43/Supplement_2/10.1086_504481/1/m_43-Supplement_2-S62-fig002.gif?Expires=1716356332&Signature=JzqHImzOm50VdhGsRh0~jZoGc7w1~uyLYt4G11Or3-ll~2zkOzApkHlUXAc4x5ZBMIElENytPFhIm2MyJUv5pgUg6IcTRSgWctVW0cTQIqAydbKTSunvbdRqj2tWBM18Use0BioaG9jaRYdmolaIClyH43Rhx3zoY4rXtHwXfDf-0Z-8ntGdVCGmnpI1OSNT0LVcx-~3-44t4tGDD4XxCmEUxLw8MrJnwLfieTSX48qSISZgqkEe1NIKmF6NwIL3TaFI6f7XJ5sR0-VQM04DbEFJescy7G1FKPeyspKZBqb7XQP7ha5TbbDS0EOaeDzTyWrULltwy3mBOR00aUeazA__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)
![Effect of anti-anaerobic antibiotic regimens on the detection and density of antibiotic-resistant gram-negative bacilli in stool of patients colonized with vancomycin-resistant enterococci (VRE). A, Detection of gram-negative bacilli resistant to ceftazidime, ciprofloxacin, or piperacillin-tazobactam in stool of patients during therapy with anti-anaerobic antibiotic regimens and in the absence of such therapy for >1 month. B, Density of resistant gram-negative bacilli in stool during anti-anaerobic antibiotic therapy and in the absence of such therapy for >2 weeks (among patients with detectable antibiotic-resistant gram-negative bacilli). C, Effect of antibiotic therapy on the density of VRE (triangles) and piperacillin-tazobactam-resistant Escherichia coli (squares) in the stool of a patient. Antibiotic therapy included piperacillin-tazobactam, vancomycin, and metronidazole (regimen A); piperacillin-tazobactam and vancomycin (regimen B); and meropenem and linezolid (regimen C). The star indicates development of catheter-related bacteremia with an E. coli isolate with a susceptibility pattern identical to that of the stool isolate. Treatment with meropenem and linezolid (regimen C) resulted in suppression of the 2 pathogens to undetectable levels in stool (reprinted with permission from [49]).](https://oup.silverchair-cdn.com/oup/backfile/Content_public/Journal/cid/43/Supplement_2/10.1086_504481/1/m_43-Supplement_2-S62-fig003.gif?Expires=1716356332&Signature=PVHybvMBgQnnnCd10Ek-gkJBhqQkX80j8ZPUW49xWdBjgnpapaNo6VV~wJNAbuOk5XHAU~sPe6WNc~QZIOXOdNoaM246zk1dnGzzzMHXBaMsJ~sGgQI-niNQAvmzXzPHuU~2HpzvAmP02LvqAnd2-EysmhHJb1BlmkUWcFbJdbB1Kc8FGWz7NqsV4eMWF166DZlTyhRg9mCs0LY3v2d1ym-Byb6sWVmrLGLakBeqIlu69kg0O2zyQ2GEV1hezqBSrLvFeTHfVGwfCG01Ml3Q-~spDJZR-ZyY~cOE7BnSANRCxK5mn5M73iFGkKcWq9lf8CiF1xXrqfnYOo8j3rwD~g__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)

Comments