Abstract
Spatial organization of metabolic enzymes allows substrate channeling, which accelerates processing of intermediates. Here, we investigated the effect of substrate channeling on the flux partitioning at a metabolic branch point, focusing on pyruvate metabolism in Saccharomyces cerevisiae. As a platform strain for the channeling of pyruvate flux, PYK1-Coh-Myc strain was constructed in which PYK1 gene encoding pyruvate kinase is tagged with cohesin domain. By using high-affinity cohesin-dockerin interaction, the pyruvate-forming enzyme Pyk1 was tethered to heterologous pyruvate-converting enzymes, lactate dehydrogenase and α-acetolactate synthase, to produce lactic acid and 2,3-butanediol, respectively. Pyruvate flux was successfully redirected toward desired pathways, with a concomitant decrease in ethanol production even without genetic attenuation of the ethanol-producing pathway. This pyruvate channeling strategy led to an improvement of 2,3-butanediol production by 38%, while showing a limitation in improving lactic acid production due to a reduced activity of lactate dehydrogenase by dockerin tagging.
Similar content being viewed by others
Introduction
In recent decades, metabolic engineering of microbial cells has received a great attention for the production of chemicals, fuels, and pharmaceuticals1,2,3. Amplification of the metabolic pathway toward desired compound and inhibition of the competing pathways are generally adopted engineering strategies to achieve high titer and yield of production. To this end, various methods of regulating gene expression levels and gene knockout/down systems have been applied to metabolic engineering2,4,5,6.
On the other hand, several efforts have been focused on the spatial organization of metabolic enzymes, leading to a substrate channeling effect, which has several potential advantages such as preventing the loss of intermediates to competing pathways, reducing the accumulation of toxic intermediates, and protecting unstable intermediates, as well as improving conversion rates7,8,9. Spatial proximity of the pathway enzymes can be achieved by several strategies such as making a fusion protein or a scaffold-based enzyme complex, and localization of the enzymes to subcellular compartments or organelles4,10. For example, expression of a fusion protein of 4-coumaroyl-CoA ligase and stilbene synthase in Saccharomyces cerevisiae resulted in up to 15-fold increase in resveratrol production compared to co-expression of these enzymes11. Synthetic protein scaffold was applied to the heterologous mevalonate pathway in Escherichia coli, resulting in a 77-fold increase in mevalonate titer with the optimized scaffold architecture9. In addition to protein scaffold, RNA- and DNA-based scaffold systems have been constructed and used to produce various target products12,13,14. The strategy of compartmentalization of metabolic pathways to specific organelles, especially mitochondria, using N-terminal targeting sequences, has also been successfully applied to the production of isobutanol, itaconic acid, and acetoin in S. cerevisiae, Aspergillus niger, and Candida glabrata, respectively15,16,17.
The feasibility of the high-affinity cohesin-dockerin interaction, found in cellulosome complex, has been demonstrated in various applications such as protein purification and enzyme immobilization18,19,20. Previously, three enzymes converting pyruvate to 2,3-butanediol, including α-acetolactate synthase (AlsS) and α-acetolactate decarboxylase (AlsD) from Bacillus subtilis and endogenous 2,3-butanediol dehydrogenase (Bdh1), were expressed as dockerin-fused proteins, and then assembled onto a scaffold containing multiple cohesin domains21. Although such a substrate channeling strategy was effective in channeling α-acetolactate produced by AlsS into 2,3-butanediol production pathway, it was not efficient to compete with the native ethanol production pathway for pyruvate availability. In this study, we aimed to test the effect of substrate channeling on the flux partitioning of pyruvate, an important metabolite at a metabolic branch point. In S. cerevisiae, pyruvate is mainly converted to ethanol in the presence of fermentable carbon sources such as glucose. Pyruvate kinase, which catalyzes the conversion of phosphoenolpyruvate (PEP) to pyruvate, is the enzyme for the last step of glycolysis. By tethering heterologous pyruvate-converting enzymes to pyruvate kinase (Pyk1) using cohesin-dockerin interaction, pyruvate flux was successfully redirected to desired pathways producing lactic acid or 2,3-butanediol.
Results
Construction of PYK1-Coh-Myc strain as a platform strain for the channeling of pyruvate flux
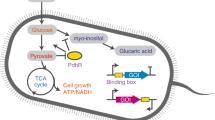
Pyruvate is a key metabolite of the central metabolism and serves as an important precursor for biosynthesis of several products such as carboxylic acid, alcohols, and amino acids. In S. cerevisiae, pyruvate produced by glycolysis is mainly converted to ethanol via acetaldehyde during fermentative growth (Fig. 1). Therefore, if pyruvate serves as a precursor for the production of target products, as in the case of producing lactic acid or 2,3-butanediol, it is very important to redirect the carbon flux from ethanol toward desired target products.
Pyruvate kinase (Pyk1) catalyzes the conversion of phosphoenolpyruvate (PEP) to pyruvate, the last step of glycolysis. Pyruvate is mainly converted to ethanol by the sequential actions of pyruvate decarboxylase (Pdc) and alcohol dehydrogenase (Adh) in S. cerevisiae. The PYK1-Coh-Myc strain expressing cohesin-tagged Pyk1 was constructed as a platform strain for the pyruvate channeling. (a) Overexpression of LdhA for lactate production or AlsS, AlsD, and Bdh1 for 2,3-butanediol production. Because cohesin-tagged Pyk1 cannot interact with native LdhA or AlsS, LdhA or AlsS competes with Pdc for the utilization of pyruvate, the common substrate. (b) Overexpression of LdhA-Doc for lactate production or AlsS-Doc, AlsD, and Bdh1 for 2,3-butanediol production. In PYK1-Myc-Coh strain, cohesin-dockerin interaction provides spatial proximity between cohesin-tagged Pyk1 and the dockerin-tagged LdhA or AlsS, resulting in pyruvate channeling to the interacting enzyme.
In this study, high-affinity cohesin-dockerin interaction was applied to recruit pyruvate-converting enzymes such as lactate dehydrogenase (LdhA) or α-acetolactate synthase (AlsS) to a pyruvate-forming enzyme, Pyk1 (Fig. 1). Although S. cerevisiae has another gene encoding pyruvate kinase, PYK2, Pyk1 is known as a main pyruvate kinase22. As a platform strain, we generated PYK1-Coh-Myc strain in which PYK1 gene in the chromosome is C-terminally tagged with cohesin domain and Myc epitope. As a control strain, PYK1-Myc strain where PYK1 is tagged only with Myc was also constructed.
Identification of the interaction between cohesin-fused Pyk1 and dockerin-fused enzyme by co-immunoprecipitation
To ensure the interaction between cohesin and dockerin domains, fused to Pyk1 and a pyruvate-converting enzyme, respectively, we performed co-immunoprecipitation (co-IP) assays using 2,3-butanediol dehydrogenase (Bdh1) as a testing interacting partner of Pyk1. PYK1-Coh-Myc strain was transformed with p415GPD-HA-Bdh1-Doc or p415GPD-HA-Bdh1, expressing N-terminal HA-tagged Bdh1 (HA-Bdh1) with or without C-terminal dockerin domain (Doc). As shown in Fig. 2, HA-Bdh1 with a C-terminal dockerin domain (HA-Bdh1-Doc) was successfully co-precipitated with Pyk1-Coh-Myc, while no interaction was detected between HA-Bdh1 and Pyk1-Coh-Myc protein. These results demonstrate that the dockerin domain of HA-Bdh1-Doc interacts specifically with the cohesin domain of Pyk1. Therefore, this approach connecting Pyk1 to pyruvate-converting enzymes based on cohesin-dockerin interaction could be applied for channeling of pyruvate produced by Pyk1 toward desired products.
Myc-tagged Pyk1 (Pyk1-Coh-Myc) was immunoprecipitated with anti-Myc antibody from PYK1-Coh-Myc strains expressing either HA-Bdh1 or HA-Bdh1-Doc. Cell lysates before immunoprecipitation (Input) and immunoprecipitates (IP) were analyzed by SDS-PAGE and immunoblotting (IB) with the indicated antibodies. HC indicates the position of the heavy chain of the anti-Myc antibody used for immunoprecipitation.
Lactate production using the pyruvate channeling system
In order to test whether co-localization of Pyk1 and pyruvate-converting enzymes through cohesin-dockerin interaction could bring to metabolic channeling of pyruvate to desired metabolic pathways, we first adopted this system to lactate production in S. cerevisiae. Lactic acid is a valuable chemical that can be used as a starting material for biodegradable polymers and as an acidulant and preservative in food industry. Although S. cerevisiae cannot produce lactate naturally, due to their robust characteristics such as high acid tolerance, there have been many attempts to engineer S. cerevisiae for lactate production by introducing heterologous lactate dehydrogenase23,24,25.
In this study, to produce lactate from pyruvate, lactate dehydrogenase LdhA from Leuconostoc mesenteroides subsp. mesenteroides ATCC 8293 was introduced into S. cerevisiae. This LdhA is specific for the production of D-lactate isomer26. First, we verified the effect of introducing native LdhA into PYK1-Myc or PYK1-Coh-Myc strain. Although the rates of glucose consumption and lactate production in PYK1-Coh-Myc strain were slightly lower than those of PYK1-Myc strain, there were no significant differences in the final production titers of lactate and ethanol between these two strains (Fig. 3a). Next, we investigated the substrate channeling effect on pyruvate distribution between lactate and ethanol by expressing dockerin-tagged LdhA (LdhA-Doc) in PYK1-Myc or PYK1-Coh-Myc strain (Fig. 3b). The PYK1-Myc control strain harboring p415GPD-ldhADoc produced 4.2 g/L lactate and 18.2 g/L ethanol after 36 h cultivation in SC-Leu medium containing 50 g/L glucose. On the other hand, PYK1-Coh-Myc strain harboring p415GPD-ldhADoc exhibited a 91% increase in lactate production (8.0 g/L) and a 9% decrease in ethanol production (16.5 g/L) compared with the control strain, demonstrating a successful substrate channeling effect between Pyk1 and LdhA (Fig. 3b). However, unfortunately, the final titer and yield of lactate decreased in both strains expressing LdhA-Doc compared with those in the strains expressing native LdhA (Fig. 3c). These results suggest that dockerin tagging might reduce the catalytic activity of LdhA, which limits the utilization of our substrate channeling strategy to improve lactate production. To confirm this, we determined the effect of dockerin tagging on the lactate dehydrogenase activity with purified enzymes, His6-LdhA and His6-LdhA-Doc. As expected, the specific activity of His6-LdhA-Doc (11.6 U/nmol) was lower than that of His6-LdhA (27.9 U/nmol) by more than two-fold.
Fermentation profiles of PYK1-Myc (open symbol) and PYK1-Coh-Myc (closed symbol) strains, harboring (a) p415GPD-ldhA or (b) p415GPD-ldhADoc. (c) The bar graphs show the production titers of lactate (left) and ethanol (right) after 36 h fermentation. Cells were grown in SC-Leu media containing 50 g/L glucose. Error bars indicate standard deviations of three independent experiments.
2,3-Butanediol production using the pyruvate channeling system
The effect of dockerin tagging on enzymatic activity might be different depending on enzymes. Therefore, to further confirm the effect of metabolite channeling for pyruvate redistribution, we attempted to use PYK1-Coh-Myc strain to produce 2,3-butanediol (Fig. 1). 2,3-Butanediol is a valuable platform chemical used in extensive industrial applications such as a liquid fuel, solvent, and monomer of synthetic rubber. 2,3-Butanediol production from pyruvate using heterologous acetoin production pathway in S. cerevisiae has been described previously27. In our previous study to verify substrate channeling using a protein scaffold, three enzymes converting pyruvate to 2,3-butanediol including α-acetolactate synthase (AlsS) and α-acetolactate decarboxylase (AlsD) from B. subtilis and endogenous 2,3-butanediol dehydrogenase (Bdh1), were expressed as dockerin-fused proteins, and then assembled onto a scaffold containing multiple cohesin domains21. Because present study aimed to demonstrate the effect of metabolite channeling on the flux partitioning at a metabolic branch point, we focused on AlsS, a pyruvate converting enzyme. Therefore, AlsS was expressed as dockerin-fusion enzyme (AlsS-Doc) that enables its recruitment to Pyk1 via cohesin-dockerin interaction (Fig. 1).
We tested the effect of introducing native AlsS together with AlsD and Bdh1 into PYK1-Myc or PYK1-Coh-Myc strain. As shown in Fig. 4a, there were no significant differences between these two strains, including glucose consumption rate and production of metabolites. After 30 h fermentation, PYK1-Myc and PYK1-Coh-Myc strains overexpressing native AlsS, AlsD, and Bdh1 from p413-SDB plasmid produced almost the same amounts of 2,3-butanediol (9.9 g/L and 10.1 g/L, respectively) and ethanol (8.3 g/L and 8.1 g/L, respectively) (Fig. 4a). These metabolite profiles during fermentation were also maintained in PYK1-Myc strain overexpressing AlsS-Doc, AlsD, and Bdh1 from p413-SDocDB, which produced 9.8 g/L 2,3-butanediol and 7.9 g/L ethanol after 30 h, suggesting that dockerin tagging does not affect AlsS activity (Fig. 4b). In contrast, PYK1-Coh-Myc strain overexpressing AlsS-Doc, AlsD, and Bdh1 produced 13.6 g/L 2,3-butanediol, which indicates about 38% increase compared with that produced in PYK1-Myc strain. Consistent with this increase in 2,3-butanediol titer, ethanol production decreased by 46% (4.3 g/L after 30 h fermentation) (Fig. 4b,c). These results demonstrate that the pyruvate flux was successfully redistributed from ethanol toward 2,3-butanediol through the spatial organization of Pyk1 and AlsS.
Fermentation profiles of PYK1-Myc (open symbol) and PYK1-Coh-Myc (closed symbol) strains, both harboring (a) p413-SDB or (b) p413-SDocDB. (c) The bar graphs show the production titers of 2,3-butanediol (left) and ethanol (right) after 30 h fermentation. Cells were grown in SC-His media containing 50 g/L glucose. Error bars indicate standard deviations of three independent experiments.
Discussion
As a proof of concept study, we investigated the effect of metabolite channeling on the flux partitioning at a metabolic branch point, focusing on pyruvate, a key intermediate of the central metabolism in S. cerevisiae. Especially, S. cerevisiae favors ethanol production from pyruvate in the presence of fermentable carbon sources even under aerobic conditions. Therefore, redirecting pyruvate flux to desired pathways in competition with ethanol production pathway is essential to produce pyruvate-derived chemicals. We constructed PYK1-Coh-Myc strain as a platform strain for pyruvate channeling and overexpressed pyruvate-converting enzymes with dockerin fusion. In the case of both lactate production and 2,3-butanediol production, pyruvate flux toward the target products was significantly enhanced, successfully competing with the native central metabolic pathway toward ethanol production even without genetic attenuation of the ethanol pathway enzymes such as pyruvate decarboxylase (Pdc) or alcohol dehydrogenase (Adh). As a result, it has been demonstrated that spatial organization of enzymes located at metabolic branch point could provide competitive advantages over the other pathways, within the frame of metabolite channeling effect.
Although cohesin tagging to Pyk1 and dockerin tagging to AlsS did not exert noticeable effects on enzyme activities, LdhA activity was severely reduced by dockerin tagging. Such a tagging-dependent inactivation of the protein function is one of the limitations of using substrate channeling strategies based on domain tagging. This problem might be solved by trying different protein-protein interaction domains or different tagging sites and linkers, which minimize the perturbation of the enzyme activity. The PYK1-Coh-Myc platform strain could also be applied to metabolic engineering strategies involving other pyruvate-converting enzymes whose activities are not largely diminished by dockerin tagging.
Methods
Strains and media
Yeast strains used in this study are listed in Table 1. S. cerevisiae CEN.PK2-1C was used as a parental strain. PYK1-Myc and PYK1-Coh-Myc strains were constructed by PCR-mediated homologous recombination based on Cre/loxP recombination system28. The integration cassettes were obtained by PCR amplification from pUG27-Myc and pUG27-Coh-Myc as templates, using a primer pair of 5′-CGGTGCTGGTCACTCCAACACTTTGCAAGTCTCTACCGTT GGTGGTTCTGGTGCTAGT-3′ and 5′-TTCAAAAAAATAATATCTTCATTCAATCATGATTCTTTTT GCATAGGCCACTAGTGGATC-3′ (sequences for homologous recombination were shown in italics). After confirmation of the correct integration of the cassette at the 3′ end of PYK1 target gene locus through PCR analysis using the confirmation primers (5′-TGTCCACTTCCGGTACCACC-3′ and 5′-ATGCAACACCTCATCGTT-3′), the marker gene was removed by transformation of Cre recombinase-expression vector, p414GPD-Cre.
Yeast cells were cultured in YPD medium (10 g/L yeast extract, 20 g/L bacto-peptone, and 20 g/L glucose) or in synthetic complete (SC) medium (6.7 g/L yeast nitrogen base without amino acids and 1.4 g/L amino acids dropout mixture lacking His, Trp, Leu, and Ura) supplemented with auxotrophic amino acids as required and 20 or 50 g/L glucose.
Plasmid construction
Plasmids used in this study are listed in Table 1. To construct C-terminal tagging vector, pUG27 plasmid containing loxP-his5+-loxP cassette was used as a starting plasmid. The DNA fragment sequentially consisting of linker sequence (GGSG), Myc epitope tag, and CYC1 terminator (TCYC1) was cloned into HindIII and SalI sites of pUG27, resulting in pUG27-Myc plasmid. The cohesin domain from Clostridium thermocellum CipA was obtained by PCR amplification using p416GPD-GST-[Coh]2 as template21. The PCR product was cloned into the BamHI site located between linker sequence and Myc epitope tag in pUG27-Myc plasmid, resulting in pUG27-Coh-Myc. In order to omit the procedure of galactose induction for the expression of Cre recombinase, the DNA fragment encoding cre recombinase was obtained by cutting the pSH63 plasmid with SpeI and XhoI and cloned into p414GPD plasmid, resulting in p414GPD-Cre. HA-tagged BDH1 was amplified by using PCR primer containing HA tag sequence and cloned into p415GPD, resulting in p415GPD-HA-BDH1. A plasmid p415GPD-HA-BDH1-Doc, expressing N-terminal HA-tagged Bdh1 with C-terminal dockerin fusion, was previously reported21. p415GPD-ldhA29, containing ldhA (LEUM_1756, D-lactate dehydrogenase) gene from L. mesenteroides ATCC 8293, was used for lactate production. The PCR product encoding ldhA without stop codon was prepared by PCR amplification and cloned into p415GPD-Doc21, resulting in p415GPD-ldhADoc. For the expression of N-terminal His-tagged proteins, E. coli expression vector pET28b was used. The DNA fragment encoding ldhA or dockerin-tagged ldhA (ldhA-Doc) was obtained by cutting the p415GPD-ldhA or p415GPD-ldhADoc with SpeI and XhoI, and cloned into NheI and XhoI sites of pET28b plasmid, resulting in pET28b-ldhA or pET28b-His-ldhA-Doc, respectively. The construction of p413-SDB, a multigene-expression vector for 2,3-butanediol pathway, was described previously30. The DNA fragment encoding dockerin-tagged alsS was obtained from the p413GPD-alsS-Doc plasmid21 and cloned into p414_PTDH3/TPYK1, resulting in p414_PTDH3-alsS-Doc-TPYK1. [alsS-Doc]-expression cassettes (PTDH3-alsS-Doc-TPYK1) flanked by MluI and NotI sites was obtained using the universal primers30 and cloned into AscI and NotI sites of p413-DB, resulting in p413-SDocDB.
Co-immunoprecipitation
The PYK1-Coh-Myc strain harboring p415GPD-HA-BDH1 or p415GPD-HA-BDH1-Doc was grown in SC-Leu medium containing 20 g/L glucose and harvested at early log phase. The cells were resuspended in 250 μl of IP150 buffer [50 mM Tris-HCl (pH 7.4), 150 mM NaCl, 2 mM MgCl2, and 0.1% NP-40)] supplemented with 0.1% protease inhibitor cocktail (Calbiochem) and 1 mM PMSF. After an equal volume of acid-washed glass beads was added to each sample, cells were lysed by vortexing 10 times for 1 min at full speed with 1 min intervals on ice. Whole-yeast lysates (700 μg of total protein) were incubated with 1 μg c-Myc antibody (Santa Cruz Biotechnology) for 3 h at 4 °C followed by addition of 15 μL protein G PLUS-Agarose beads (Santa Cruz Biotechnology). After further incubation for 90 min at 4 °C, samples were washed three times with IP150 buffer and the bound proteins were eluted by boiling in sample buffer for 10 min. Samples were resolved by SDS-PAGE and proteins were detected by immunoblotting with the use of antibodies against Myc (Santa Cruz Biotechnology) and HA epitope tag (Roche Life Science).
Protein expression and purification
Escherichia coli Rogetta gami2 (DE3) pLysS was used as a host strain to express N-terminal His-tagged proteins. The bacteria harboring pET28b-His-ldhA or pET28b-His-ldhADoc were grown to OD600 of 0.8 in LB medium supplemented with 50 μg/mL kanamycin at 37 °C, and induced with 1 mM isopropyl 1-thio-β-D-galactopyranoside (IPTG) at 30 °C for 4 h with shaking at 170 rpm. Cells were harvested, resuspended in binding buffer [20 mM Tris-HCl (pH 8.0), 500 mM NaCl, and 30 mM imidazole] supplemented with 0.1% protease inhibitor cocktail (Calbiochem) and 1 mM phenylmethylsulfonyl fluoride (PMSF), and lysed by sonication on ice for a total pulsing time of 10 min (pulse on for 15 s and off for 15 s). The cell lysates were centrifuged at 8,000 × g for 15 min at 4 °C and the supernatant samples were incubated with HisPur Ni-NTA resin (Thermo Scientific) for 2 h at 4 °C. After the Ni-NTA resin was washed twice with wash buffer [20 mM Tris-HCl (pH 8.0), 500 mM NaCl, and 45 mM imidazole], enzymes were eluted with elution buffer [20 mM Tris-HCl (pH 8.0), 500 mM NaCl, and 500 mM imidazole]. The expression and purity of the enzymes were confirmed by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and the concentrations of purified enzymes were determined by the Bradford assay (Bio-Rad) with a bovine serum albumin standard.
Lactate dehydrogenase activity assay
The enzyme activity of lactate dehydrogenase was determined as previously described31. Briefly, reduction of pyruvate to lactate was monitored by measuring the oxidation rate of NADH with a UV spectrophotometer at 340 nm. The enzyme activity was examined using 50 mM Tris-HCl (pH 7.0) buffer solution containing 10 mM sodium pyruvate and 0.2 mM β-NADH. One unit (U) of enzyme activity was defined as the amount that catalyzed the oxidation of 1 μmol of NADH per minute at pH 7.0.
Culture conditions
For the production of lactate, yeast cells harboring p415GPD-ldhA or p415GPD-ldhADoc were pre-cultured in SC-Leu medium containing 20 g/L glucose and diluted to OD600 of 0.3 in 7.5 mL of SC-Leu medium containing 50 g/L glucose in a 50 mL conical tube at 30 °C with shaking at 170 rpm. For the production of 2,3-butanediol, yeast cells harboring p413-SDB or p413-SDocDB were cultured in the same condition of lactate production in SC-His medium instead of SC-Leu medium.
Analytical methods
The concentration of metabolites in the culture medium was determined by high performance liquid chromatography (HPLC) with BioRad Aminex HPX-87H column as previously described32. Briefly, 800 μL of culture supernatants were collected and filtered through a 0.22 μm syringe filter. Analysis was performed with 5 mM H2SO4 as a mobile phase at a flow rate of 0.6 mL/min and refractive index (RI) detector. The column temperature and detector temperature were maintained at 60 °C and 35 °C, respectively.
Additional Information
How to cite this article: Kim, S. et al. Redirection of pyruvate flux toward desired metabolic pathways through substrate channeling between pyruvate kinase and pyruvate-converting enzymes in Saccharomyces cerevisiae. Sci. Rep. 6, 24145; doi: 10.1038/srep24145 (2016).
References
Keasling, J. D. Manufacturing molecules through metabolic engineering. Science 330, 1355–1358 (2010).
Lee, J. W. et al. Systems metabolic engineering of microorganisms for natural and non-natural chemicals. Nat. Chem. Biol. 8, 536–546 (2012).
Nielsen, J., Larsson, C., van Maris, A. & Pronk, J. Metabolic engineering of yeast for production of fuels and chemicals. Curr. Opin. Biotechnol. 24, 398–404 (2013).
Woolston, B. M., Edgar, S. & Stephanopoulos, G. Metabolic Engineering: Past and Future. Annu. Rev. Chem. Biomol. Eng. 4, 259–288 (2013).
Soma, Y., Tsuruno, K., Wada, M., Yokota, A. & Hanai, T. Metabolic flux redirection from a central metabolic pathway toward a synthetic pathway using a metabolic toggle switch. Metab. Eng. 23, 175–184 (2014).
Brockman, I. M. & Prather, K. L. Dynamic knockdown of E. coli central metabolism for redirecting fluxes of primary metabolites. Metab. Eng. 28, 104–113 (2015).
Geck, M. K. & Kirsch, J. F. A novel, definitive test for substrate channeling illustrated with the aspartate aminotransferase/malate dehydrogenase system. Biochemistry 38, 8032–8037 (1999).
Jorgensen, K. et al. Metabolon formation and metabolic channeling in the biosynthesis of plant natural products. Curr. Opin. Plant Biol. 8, 280–291 (2005).
Dueber, J. E. et al. Synthetic protein scaffolds provide modular control over metabolic flux. Nat. Biotechnol. 27, 753–759 (2009).
Conrado, R. J., Varner, J. D. & DeLisa, M. P. Engineering the spatial organization of metabolic enzymes: mimicking nature’s synergy. Curr. Opin. Biotechnol. 19, 492–499 (2008).
Zhang, Y. S. et al. Using unnatural protein fusions to engineer resveratrol biosynthesis in yeast and mammalian cells. J. Am. Chem. Soc. 128, 13030–13031 (2006).
Delebecque, C. J., Lindner, A. B., Silver, P. A. & Aldaye, F. A. Organization of intracellular reactions with rationally designed RNA assemblies. Science 333, 470–474 (2011).
Conrado, R. J. et al. DNA-guided assembly of biosynthetic pathways promotes improved catalytic efficiency. Nucleic Acids Res. 40, 1879–1889 (2012).
Lee, J. H. et al. Improved production of L-threonine in escherichia coli by use of a DNA scaffold system. Appl. Environ. Microbiol. 79, 774–782 (2013).
Avalos, J. L., Fink, G. R. & Stephanopoulos, G. Compartmentalization of metabolic pathways in yeast mitochondria improves the production of branched-chain alcohols. Nat. Biotechnol. 31, 335–341 (2013).
Blumhoff, M. L., Steiger, M. G., Mattanovich, D. & Sauer, M. Targeting enzymes to the right compartment: Metabolic engineering for itaconic acid production by Aspergillus niger . Metab. Eng. 19, 26–32 (2013).
Li, S. B., Liu, L. M. & Chen, J. Compartmentalizing metabolic pathway in Candida glabrata for acetoin production. Metab. Eng. 28, 1–7 (2015).
You, C. & Zhang, Y. H. P. Self-assembly of synthetic metabolons through synthetic protein scaffolds: One-step purification, co-immobilization, and substrate channeling. ACS Synth. Biol. 2, 102–110 (2013).
Liu, F., Banta, S. & Chen, W. Functional assembly of a multi-enzyme methanol oxidation cascade on a surface-displayed trifunctional scaffold for enhanced NADH production. Chem. Commun. 49, 3766–3768 (2013).
Kim, S., Baek, S. H., Lee, K. & Hahn, J. S. Cellulosic ethanol production using a yeast consortium displaying a minicellulosome and beta-glucosidase. Microb. Cell Fact. 12 (2013).
Kim, S. & Hahn, J. S. Synthetic scaffold based on a cohesin-dockerin interaction for improved production of 2,3-butanediol in Saccharomyces cerevisiae . J. Biotechnol. 192, 192–196 (2014).
Boles, E. et al. Characterization of a glucose-repressed pyruvate kinase (Pyk2p) in Saccharomyces cerevisiae that is catalytically insensitive to fructose-1,6-bisphosphate. J. Bacteriol. 179, 2987–2993 (1997).
Ishida, N. et al. Efficient production of L-lactic acid by metabolically engineered Saccharomyces cerevisiae with a genome-integrated L-lactate dehydrogenase gene. Appl. Environ. Microbiol. 71, 1964–1970 (2005).
Tokuhiro, K. et al. Double mutation of the PDC1 and ADH1 genes improves lactate production in the yeast Saccharomyces cerevisiae expressing the bovine lactate dehydrogenase gene. Appl. Microbiol. Biotechnol. 82, 883–890 (2009).
Lee, J. Y., Kang, C. D., Lee, S. H., Park, Y. K. & Cho, K. M. Engineering cellular redox balance in Saccharomyces cerevisiae for improved production of L-lactic acid. Biotechnol. Bioeng. 112, 751–758 (2015).
Li, L. et al. Characterization of the major dehydrogenase related to D-lactic acid synthesis in Leuconostoc mesenteroides subsp mesenteroides ATCC 8293. Enzyme Microb. Technol. 51, 274–279 (2012).
Kim, S. J., Seo, S. O., Jin, Y. S. & Seo, J. H. Production of 2,3-butanediol by engineered Saccharomyces cerevisiae . Bioresour. Technol. 146, 274–281 (2013).
Gueldener, U., Heinisch, J., Koehler, G. J., Voss, D. & Hegemann, J. H. A second set of loxP marker cassettes for Cre-mediated multiple gene knockouts in budding yeast. Nucleic Acids Res. 30 (2002).
Baek, S. H., Kwon, E. Y., Kim, Y. H. & Hahn, J. S. Metabolic engineering and adaptive evolution for efficient production of D-lactic acid in Saccharomyces cerevisiae . Appl. Microbiol. Biotechnol. 100, 2737–2748 (2016).
Kim, S. & Hahn, J. S. Efficient production of 2,3-butanediol in Saccharomyces cerevisiae by eliminating ethanol and glycerol production and redox rebalancing. Metab. Eng. 31, 94–101 (2015).
Jun, C. et al. Discovery and characterization of a thermostable D-lactate dehydrogenase from Lactobacillus jensenii through genome mining. Process Biochem. 48, 109–117 (2013).
Kim, S., Lee, K., Bae, S. J. & Hahn, J. S. Promoters inducible by aromatic amino acids and g-aminobutyrate (GABA) for metabolic engineering applications in Saccharomyces cerevisiae . Appl. Microbiol. Biotechnol. 99, 2705–2714 (2015).
Mumberg, D., Muller, R. & Funk, M. Yeast vectors for the controlled expression of heterologous proteins in different genetic backgrounds. Gene 156, 119–122 (1995).
Acknowledgements
This work was supported by the National Research Foundation of Korea (NRF) Grant funded by the Korean Government (NRF-2015R1A2A2A01005429).
Author information
Authors and Affiliations
Contributions
S.K. and J.-S.H. designed the experiments and wrote the manuscript. S.K. and S.-J.B. performed the experiments and analyzed the data. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Kim, S., Bae, SJ. & Hahn, JS. Redirection of pyruvate flux toward desired metabolic pathways through substrate channeling between pyruvate kinase and pyruvate-converting enzymes in Saccharomyces cerevisiae. Sci Rep 6, 24145 (2016). https://doi.org/10.1038/srep24145
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep24145
This article is cited by
-
l-Lactic acid production from glucose and xylose with engineered strains of Saccharomyces cerevisiae: aeration and carbon source influence yields and productivities
Microbial Cell Factories (2018)
-
The role of dynamic enzyme assemblies and substrate channelling in metabolic regulation
Nature Communications (2018)
-
Metabolic engineering of Zymomonas mobilis for 2,3-butanediol production from lignocellulosic biomass sugars
Biotechnology for Biofuels (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.