ABSTRACT
To determine the possible role of the epigenetic mechanisms in carcinogenesis of the hepatocellular carcinoma, we methylation-profiled the promoter CpG islands of twenty four genes both in HCC tumors and the neighboring non-cancerous tissues of twenty eight patients using the methylation-specific PCR (MSP) method in conjunction with the DNA sequencing. In comparison with the normal liver tissues from the healthy donors, it was found that while remained unmethylated the ABL, CAV, EPO, GATA3, LKB1, NEP, NFL, NIS and p27KIP1 genes, varying extents of the HCC specific hypermethylation were found associated with the ABO, AR, CSPG2, cyclin a1, DBCCR1, GALR2, IRF7, MGMT, MT1A, MYOD1, OCT6, p57KIP2, p73, WT1 genes, and demethylation with the MAGEA1 gene, respectively. Judged by whether the hypermethylated occurred in HCC more frequently than in their neighboring normal tissues, the hypermethylation status of the AR, DBCCR1, IRF7, OCT6, and p73 genes was considered as the event specific to the late stage, while that the rest that lacked such a distinguished contrast, as the event specific to the early stage of HCC carcinogenesis. Among all the clinical pathological parameters tested for the association with, the hypermethylation of the cyclin a1 gene was more prevalent in the non-cirrhosis group (P=0.021) while the hypermethylated p16INK4a gene was more common in the cirrhosis group (P=0.017). The concordant methylation behaviors of nineteen genes, including the four previously studied and their association with cirrhosis has been evaluated by the best subgroup selection method. The data presented in this report would enable us to shape our understanding of the mechanisms for the HCC specific loss of the epigenetic stability of the genome, as well as the strategy of developing the novel robust methylation based diagnostic and prognostic tools.
Similar content being viewed by others
INTRODUCTION
Hepatocellular carcinoma(HCC) is one of the commonest cancerous diseases, rating the fifth in occurrence and the third in mortality worldwide1. As it is geographically biased toward the several parts of Asia and Africa, China in particular, it presents one of the major health threat in China2, 3. The dismal prognostic future of the patients is largely attributive to the rapidly advancing nature and difficulties in early diagnosis of HCC. Therefore, there are urgent needs for the robust diagnostic, prognostic and even therapeutic approaches that can only be brought about by the much improved understanding of the fundamental aspects at various biological levels of the events during the malignant transformation of the normal hepatocytes. The decades of intensive molecular genetic analyses have yielded a considerable amount of information on the potential genetic defects associated with the natural course of carcinogenesis of hepatocytes4, 5, 6. Until recently, the epigenetic mechanisms without the changes in DNA sequence has been found capable of profoundly affecting the transcription status of both genes and repetitively sequences, that subsequently confers the growing advantage to tumor cells over their normal counterparts7, 8, 9. The covalent addition of the methyl group at the C5 of cytosine in the CpG dinucleotide is essentially the only form of the covalent modification of DNA in high eukaryotes, having a number of biochemical as well as biological implications It can eliminate the sequence specific binding of the transcription factors to the cognate cis-elements and promote the association of the methyl CpG specific binding proteins to the methyl CpG, with a cascade of reactions leading to the chromatin condensation and transcription silencing10. Over 50% of the protein coding genes have at least one CpG island within or near their promoters. Expressions of these genes are subjected to the controls over the methylation state of the CpG islands. Aberrant DNA methylation pattern changes the gene transcription and has been etiologically linked to the occurrence of a number of genetic diseases including cancers10. The enzymes responsible for DNA methylation are the DNA methyl transferase I and IIIA as well as IIIB. The former is mainly responsible for the maintenance of the methylation status of genomes after DNA replication, whereas the later two act principally in the de novo DNA methylation in the early development of high eukaryotic organisms10. Elevated expression of these three methyl transferase genes were reported in the majority of cancers tested, which may partly account for the increased local hypermethylation10, 11, 12. However, recent evidences demonstrated that the histone modification, methylation of histone H3 in particular, might occur prior to the establishment of the DNA hypermethylation pattern that contribute to the long-term silencing of gene transcription13.
Changes in the DNA methylation patterns demonstrated in all the cancers examined, consist of the global level hypomethylation in parallel with the local hypermethylation12, 14. The genome-wide hypome-thylation can result in active transcription of the transposon like repetitive sequences (such as the Alu and LINE repeats in mammals) that have been linked to the increased genome instability, a predominant hallmark of cancer cells15, 16, 17. The hypermethylated status of the promoter CpG islands has been linked to the expression silencing of the tumor suppressor genes and implicated as the 2nd hit, reminiscent to the loss of heterozygosity or other type of genetic deletion for total inactivation of the tumor suppressor genes in cancers 18, 19, 20, 21. The loss of the genetic imprinting attributed to the changes of DNA methylation, such as reactivation of the IGF-2 gene, has been linked to the rapid proliferation of tumor cells in several types of human tumors22. The reverse process, i.e.,demethylatin can also result in the transcription activation of the otherwise inert genes, including c-myc, and c-ras, even though all of these genes lack the typical CpG island within or near to the promoters23. The association between the hypomethylation of the promoter CpG island and over-expression state of the genes such as SURVIVIN24 and hTERC that encodes the RNA component of the telomerase25 have been recently reported.
Methylation profiling of the promoter CpG islands of the known genes has been an important information gathering process for new insights in our understanding of the mechanisms of the DNA methylation in both initiation and progression of the carcinogenesis, as well as the new clues for development of the relevant diagnostic and prognostic methods and even for therapeutics against cancers. The recent collective efforts have identified a list of over one hundred genes, the promoter CpG islands of which change in various tumors (http://www.missouri.edu/hypermet/list_of_promoters.htm). However, as the majority of the studies to date had only targeted one or a few genes in rather small patient groups, the concurrent hypermethylation behavior of multiple genes has only been addressed in few tumor types, except for very few examples26. Prior to our recent work27 where methylation profile of the promoter CpG islands of twenty genes in twenty nine HCC patient samples were presented, there had been only two reports describing the HCC specific change in the methylation profiling of three tumor suppressor genes: p16INK4a, cyclin-dependent kinase inhibitor 4b (p15INK4b) and the alternative reading frame of the cyclin-dependent kinase inhibitor 4a (p14ARF), respectively. In that study we found that sixteen genes adenomatosis polyposis coli (APC), apoptotic protease activating factor (APAF1), breast cancer 1 (BRCA1), cadherin type 1 (CDH1), death-associated protein kinase 1 HCC (DAPK1), mutL homolog 1 (hMLH1), Telomerase RNA component (hTERC), p14ARF, p15INK4b, phosphatase and tensin homolog (PTEN), ras association domain family 1 protein isoform 1c (RASSF1c), retinoblastoma 1 (RB1), retinoic acid receptor, beta (RAR-b), SURVIVIN, tissue inhibitor of metalloproteinase 3 (TIMP3) and von Hippel-Lindau syndrome (VHL) remained unmethylated in all the sample tested, whereas the following four genes: Caspase 8 (CASP8), H-cadherin (CDH13), p16INK4a and RASSF1a genes displayed the HCC associated hypermethylation to varying extents. The lack of the HCC specific changes in the sixteen of the twenty genes whose hypermethylation state of the promoter CpG island has been linked to many other types of cancer27 was indeed a surprise. In order to establish the concordant methylation behavior of the genes displaying the HCC specific changes, we further extended our study to other twenty four genes to assess the extent of the methylation mediated mechanisms in the HCC and found fifteen genes had displayed the HCC specific changes. By the stringent mathematic analyses of the concordant methylation behaviors of the nineteen genes (including the four in the previous study27), the subsets of the two to nine genes have been established in HCC and its cirrhosis/non-cirrhosis subgroups, which may provide the useful clues for the DNA methylation based diagnostic and prognostic assays for HCC.
MATERIALS AND METHODS
Tissue samples and DNA extraction
With the informed consent of all patients and donors and approval of the ethics committee, the samples of tumor and adjacent non-cancerous tissues were collected from HCC patients (n = 28) during surgery at The Qidong County Hospital, The Oriental Institute for Liver Diseases and Guangxi Provincial Hospital, respectively. In addition, normal liver tissues (n = 4) were obtained from liver donors at the Liver Transplantation Unit in The First Affiliated Hospital, College of Medicine, Zhejiang University. The pathological classification of HCC tissues was carried out and the stage of each HCC was determined according to criteria outlined by the Liver Cancer Study Group of Japan28. Special efforts were made to include the corresponding non-cancerous tissues, samples of which were overlooked in all previous studies of HCC24. Furthermore, pathological examination of the tissues had been carried out to eliminate samples in which contamination of undesirable tissues exceeded 20%. To emphasize the HCC specificity of this study, normal liver tissues from four male healthy donors were collected from the Liver Transplantation Unit of the Zhejiang First Hospital as the normal liver control.
Total genomic DNA was extracted from frozen tissue specimens (50-100mg) according to standard protocol with some modifications 27. Frozen pulverized powders of the specimens were re-suspended with 2 ml lysis buffer: 50 mM Tris-HCl pH 8.0, 50 mM EDTA, 1% SDS, 10 mM NaCl plus 100 mg/ ml boiling-treated RNase A (Sigma). Following one hour of incubation at 37 °C, Proteinase K (Roche, USA) was added to the cellular lysates for a final concentration of 100 mg/ ml and the digestion was carried out at 55°C for 2 h. Organic extractions with a half volume of Phenol/Chloroform/Isoamyl alcohol (1:1:0.04) were repeatedly carried out until no visible interphase remained after centrifugation. DNA was precipitated from the aqueous phase in the presence of 0.3 M NaOAc pH 7.0 and two and a half volumes of ethanol. The DNA pellet was washed once with 70% ethanol and dissolved at 65 °C for 30 min with 0.2 - 0.4 ml TE (10 mM Tris-HCl pH 7.4 and 1 mM EDTA), followed by storage at 4 °C until further use. The DNA concentrations were calculated according to their OD260nm readings.
Bisulphate treatment of DNA and methylation specific PCR (MSP)
The primer pairs for the methylation specific PCR in this report were either adopted or designed according to the same principle with assistance of the software packages for the CpG islands identification (http://www.uscnorris.com/cpgislands) and the primer design (http://micro-gen.ouhsc.edu/cgi-bin/primer3_www.cgi).
The methylation status of the promoter CpG islands of twenty four genes in all sample DNA were analyzed by MSP on the sodium-bisulfite converted DNA 27. In detail, 10 mg DNA in 50 ml TE was incubated with 5.5 ml of 3 M NaOH at 37 °C for 10 min, followed by a 16 hour treatment at 50 °C after adding 30 ml of freshly prepared 10 mM hydroquinone and 520 ml of freshly prepared 3.6 M sodium-bisulfite at pH 5.0. The DNA was desalted using a home-made dialysis system with 1% agarose (detailed protocol will be provided upon request). The DNA in the desalted sample (approximately 100 ml in volume) was denatured at 37 °C for 15 min with 5.5 ml of 3 M NaOH followed by ethanol precipitation with 33 ml 10 M NH4OAC and 300 ml ethanol. After washing with 70% ethanol, the gently dried DNA pellet was dissolved with 30 ml TE at 65 °C for 10 min. The DNA sample was finally stored at −20 °C until further use. PCR reaction was carried out in a volume of 15 ml with 50 ng or less template DNA with FastStart Taq polymerase (Roche, Germany) as follows. After an initial heat denaturing step 4 min treatment at 94°C, 30 cycles of 92°C for 15 sec, varying temperatures with primer pairs for 15 sec and 72 °C for 20 sec, was carried out. The PCR products were separated by 1.2 % ethidium bromide containing agarose gel electrophoresis with 1 X TAE and visualized under UV illumination. To verify the PCR results, representative bands from each target were gel-purified and cloned into T-vector (Promega, USA) followed by automatic DNA sequencing provided by BoCai (Shanghai, China). Only verified results were presented in this report.
Statistics
The methylation data were dichotomized as 1 for the co-existence of the methylated and unmethylated alleles, 2, for methylated allele only and 0 for the unmethylated for both alleles to facilitate statistical analysis using contingency tables. The methylation profiles of each individual gene (in percentage) classified by the cirrhosis status of the patients were presented both in table and in plot. The statistic analyses for the association between the methylation profile of the gene and each of the clinical-pathological parameters were carried out with the statistics package (http://www.R-project.org/), where both Pearsong's Chi-square test with Upton's adjustment and Fisher exact test (http://www.R-project.org/) used to deal with the sample cells with the low expected values. The relative frequency with a 95% confidence interval (P<0.05) for a binomial distribution was calculated for the whole as well as each subtypes of astrocytoma patients.
The concordant methylation behaviors of the genes were established by comparing the relevant occurrence of various subsets containing two, three and four genes respectively, with the best sub-set selection method29.
RESULTS AND DISCUSSIONS
Clinical-pathological considerations
The geographical distribution of HCC varies dramatically in China. In areas such as Qidong county near Shanghai as well as Guangxi province, the annual HCC incidence is as high as 70 - 96/10,0000 and approximately seven to eight fold higher than that in the low incidence areas in China30. In view of the high likelihood of the potential geographic impact, we deliberately recruited nineteen patients in the Shanghai area, including eight patients from Qidong County, and nine patients from Guangxi province. All the patients were hepatitis B virus (HBV) infected by both immuno-serological assays and the PCR test for existence of the HBVX gene in tumor tissue (result not shown). No geographical variation is detected between these two patient groups. For instance, male patients accounted for 78.9% and 80%, cirrhosis occurred in 52.6 % and 50 % of patients and diagnoses of grade I were 28% and 30%, grade II were 44% and 40 % and grade III were 28% and 30 % of the Shanghai and Guangxi patient groups, respectively27. Therefore, the remaining analyses were carried out without any consideration of geographic impact.
Methylation profiling
The methylation specific PCR (MSP) on the bisulphate treated DNA has been widely used for its simplicity as well as speed. In the previous work, we verified the MSP data by sequencing the representative PCR products and validated the data of two among twenty genes studied with a non PCR mediated method, i.e., Southern analyses of the DNA digests by the methylation sensitive enzymes27. In this extended study, therefore, the MSP in conjunction with the DNA sequencing of the batch-treated genomic DNA from the same patient group and the normal control was adopted.
The in vivo malignant transformation of normal cells is a multiple-stage process, involving at least a dozen of genetic and epigenetic changes. In this connection, there is a prevalent notion that the morphologically “normal” cells adjacent to the cancerous tissues may have already suffered from the genetic or/and epigenetic changes that are specific to the early phase of malignant transformation. To take this favorable assumption into account, we deliberately recruited, both in this report and the previous, the neighboring non-cancerous liver tissues in addition to HCC tissues into study, which have been often excluded in other similar studies(27 and references within).
The concerns were fully justified as to the cross-contamination of the normal tissue in the HCC samples, or vice versa. However, this seemed unfounded in our studies. First, over forty four genes (twenty four genes were in this study and twenty genes were in the previous report27), were chemically treated in batch and methylated profiled in group. The obtained well diversified methylation patterns of these forty four genes are fully incompatible with the assumption that there may be the severe cross-contamination of the undesirable tissues in the designated samples. Second, MSP is a sensitive method capable of detecting a very low level of contamination of the undesirable DNA. Thirdly, in addition to the sequencing verification, the HCC specific methylation profile of the p14ARF and p15INK4b genes had been confirmed by a non-PCR based Southern analysis of the digested DNA with the methylation sensitive restriction enzymes27.
Another issue of importance is concerning the methylation heterozygosity in any given samples. As shown the Fig 1 and Tab 2, although some samples were homozygously methylated or unmethylated, the majority had the co-existed methylated and unmethylated alleles. Inclusion of the normal liver tissues in our study enabled us to establish the normal methylation pattern of all the forty four genes, that except for the MAGEA131 being homozygously methylated, the rest forty three were homozygously demethylated27 (Fig 1 and Tab 2). Therefore, any changes in the methylation pattern from the normal liver tissue's to the non-cancerous neighboring tissues or HCC tissues should indicate the involvement of the DNA methylation mediated events during the HCC carcinogenesis, that have been scored as the positive methylation change. Whether the alleles lacking in HCC specific methylation change may lose its function via the genetic mechanisms remains to be addressed in the future.
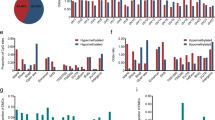
Methylation profiles of the promoter CpG islands of twenty four genes in HCC. Both electrophoretic patterns of the representative PCR products of each of twenty four targets (indicated respectively, at the top of figures) and the sequencing verification of the one representative PCR product were presented. To indicate the methylation status, the sequenced data are aligned with the wild-type sequence. *, size markers, the bands of 250 bp and 100 bp were shown. U, the unmethylated; M, the hypermethylated. panels: 1, ABL; 2, ABO; 3, AR; 4, CAV; 5, CPSG2; 6, cyclin a1; 7, DBCCR1; 8, EPO; 9, GALR2; 10, GATA3; 11, IRF7; 12, LKB1; 13, MAGEA1; 14, MGMT; 15, MT1A; 16, MYOD1; 17, NEP; 18, NFL; 19, NIS; 20, OCT6; 24, 21, p27KIP1; 22, p57KIP2; 23, p73 and 24, WT1.
The in vivo malignant transformation of normal cells is a multiple-stage process, which involves multiple genetic and epigenetic changes. In this connection, it has been generally accepted that the morphologically “normal” cells adjacent to the cancerous tissues may have already been abnormal at both genetic and epigenetic levels, reflecting the early-phase-specific alteration of the malignant transformation. We, hence, recruited the neighboring non-cancerous liver in addition to HCC tissues, to address the stage-specific nature of the methylation changes that are likely to be reflected in the neighboring non-cancerous and HCC tissues, respectively.
Methylation profiling of twenty four genes with or without the proven roles in carcinogenesis of the human cancers
The consequence of the tumor-specific defects in the epigenetic homeostasis is global, and some detectable changes may have etiological role to play whereas others may simply the by-stander by nature. Therefore, from the list of the promoter CpG island containing genes (http://www.missouri.edu/hypermet/list_of_promoters.htm) we deliberately selected the targets that lack any proven etiological role in carcinogenesis of other tissue origins, The former group consisted of the genes encoding the growth factor for the erythropoiesis (EPO) 32, a ubiquitously expressed transcription factor (OCT6)33, the blood cell typing antigen (ABO) 34, and the myogenetic or erythropoietic lineage-specific transcription factors (MYOD1 of GATA3)35 36as well as the light chain of neurofilament (NFL)37. In the second group, there were four tumor suppressor genes, including two cyclin-dependent kinase inhibitors: p27KIP1 38and p57KIP239; p53 analogue, p7338, 40, as well as the Wilms tumor 1 gene, WT1. There were seven genes encoding the surface proteins or nuclear receptors acting actively in the intercellular interactions: galanin receptor 2(GALR2)42, melanoma specific antigen A1 (MAGEA1), the membrane metallo-endopeptidase (NEP) 43, solute carrier family 5 (NIS) 44, caveolin 1 (CAV) 45, chondroitin sulfate proteoglycan 2 (CSPG2) 46 and androgen receptor (AR)47. Three genes implicated in signal transduction, cyclin a148, the interferon regulatory factor 7 (IRF7), and a serine/threonine kinase 11 (Peutz-Jeghers syndrome) gene (LKB1)18 were selected. The proto-oncogenes in this group were: the gene encoding v-abl homologue 1 (ABL)49 and for the deleted in bladder cancer chromosome region candidate 1 (DBCCR1)50. The final two in the list were the genes that may be responsible for detoxification of liver cells: O-6-methylguanine-DNA methyltransferase (MGMT) 18and metallothionein 1 A gene (MT1A)51.
As shown in Fig 1 and Tab 2, within the group of genes maintaining unmethylated in all three types of samples tested, there are the genes devoid of any demonstrated association with tumors: EPO(panel 8), GATA3(panel 10), NFL(panel 18) and NIS(panel 19), as well as the genes having the demonstrated hypermethylation mediated gene silencing in the human tumors of the non-HCC origins: ABL(panel 1), CAV(panel 4), LKB1(panel 12), NEP(panel 17), and p27KIP1(panel 21) genes. The simplistic explanation would be that either inactivation of these genes in HCC via the genetic based mechanisms, or HCC formation does not require the functional inactivation of these genes.
The genes with the HCC specifically altered methylation profiles
Fourteen genes: ABO(panel 2), AR(panel 3), CSPG2(panel 5), cyclin a1(panel 6), DBCCR1(panel 7), GALR2 (panel 9), IRF7(panel 11), MGMT(panel 14), MT1A(panel 15), MYOD1(panel 16), OCT6(panel 20), p57KIP2(panel 22), p73(panel 23), and WT1(panel 24) were unmethylated in all four cases of the normal liver tissues, and methylated to various extents in the patient's samples. Among them, there were the genes devoid of any obvious tumor association: 1, ABO (transferase A for the ABO blood typing), 2, MYOD1 (the myogenic specific transcription factor), and 3, OCT6 (a common transcription factor with POU domain). The hypermethylation frequency was as higher as 32%(9/28) and 50%(14/28) for ABO gene, 54%(15/28) and 54%(15/28) for MYOD1, as well as 50%(14/28) and 82%(23/28) for OCT6 gene in the non-cancerous neighboring liver and HCC tissue, respectively. Despite the higher incidence of changes in the methylation profile, it is not possible to discard the possible by-stander nature of such alterations (Fig 1 and Tab 2). There were suggestions for the possible aging related increase in the promoter CpG island of the genes including OCT6 and MYOD152, 53, However, it seems not the case in this study as there was no obvious correlation between the age of the HCC patients and the occurrence of the methylated status of the promoter CpG island. Alternatively, likely loss of these three gene expression (extrapolated from the hypermethylated status of the relevant promoter CpG islands) indeed play some parts in the HCC formation in the unknown manners, which may deserve the further investigation.
The rest are the genes implicating in the human tumors of other tissue origins via the methylation mediated gene silencing(http://www.missouri.edu/hypermet/list_of_promoters.htm): AR(panel 3), CSPG2(panel 5), cyclin a1(panel 6), DBCCR1(panel 7), GALR2(panel 9), IRF7(panel 11), MGMT(panel 14), MT1A(panel 15), p57KIP2(panel 22), p73(panel 23), and WT1 (panel 24)} or demethylation mediated activation: MAGEA1(panel 13). Among these targets, the MGMT gene was rarely methylated ( 3.57%, 1/28). It encodes the enzyme capable of removing the alkylating adducts from the O(6) position of guanine and protects the cells from cytotoxic and mutagenic effects 20. The MT1A gene that encodes the protein responsible for the cell detoxification targeting at various adversary stimuli including the heavy metals, has been reported being inactivated by the hypermethylated promoter CpG island in rat hepatoma 51. It was only marginally hypermethylated in HCC (10.71%, 3/28), indicating that the DNA methylation mediated mechanism should not play any significant role in its inactivation, if there is any, in HCC. The following three genes: CSPG2 (17.86%), GALR2(21.43%) and p57KIP2(14.29%) were moderately hypermethylated in HCC, expression of which may not be significantly affected by the DNA hypermethylation of their promoter CpG island in HCC.
The AR gene encodes the androgen receptor that play a key role in the signal transduction pathways of cells to respond to the male steroid hormone, androgen and was reported to be inactivated via the epigenetic mechanism in prostate cancers54. HCC seems an androgen-dependent tumor as it occurs five times more in males than that in the females. In this connection, a recent study reported that the AR gene was rarely expressed in the poorly differentiated HCC affecting the males55. This does correlate well with the observation in this study that there was high occurrence (82.14%, 20/28) of the hypomethylation of the AR gene in the male HCC patient tissue. Although it was hypermethylated more frequently in the female (83.3%, 5/6) than the male patient group (68%, 15/22), the implication may be different. Loss of the hypermethylation of the AR gene in the females may symbolize the tumor associated defects in epigenetic control in general.
The DBCCR1 gene identified at the region (q32-q33) within chromosome 9q with high LOH in human bladder carcinoma is the most frequent genetic alteration in transitional cell carcinoma (TCC) of the bladder. Its loss of expression correlated well the hypermethylation of the promoter CpG island at a frequency as high as 52% (36/69) bladder carcinoma50. In this study, the promoter CpG island has been detected at a comparable frequency (53.57%), indicating its possible role in the HCC formation.
The p73 gene encodes a protein structurally and functionally homologous to TP53, and maps to chromosomal band 1p36.33, where loss of heterozygosity has been observed in up to 90% of oligodendrogliomas and in 10-25% of diffuse astrocytoma56, 57. In this report, we found that the p73 gene was prevalently methylated (68 %) in HCC tissues. However, whether its hypermethy- lation correlates with its functional inactivation remain to be determined.
Both genetic defects and epigenetic abnormalities concerning the WT1 gene have been implicated in the formation of Wilm's tumor58. In this study, we also found the WT1 gene hypermethylated in 29% of the HCC cases, implying its possible involvement in the formation of HCC.
Resistance of tumors to the cytotoxic chemotherapies may result from the disrupted apoptosis programs and remains a major obstacle in cancer treatment. In this connection, the IRF7 gene was also analyzed, the analogue (IRF1) of which has been implicated in the IFN gammar mediated apoptosis with a profound impart to the chemo-sensitivity of the tumor cells59, 60. The IRF7 expression was negatively regulated by the promoter methylation61. In this study, the IRF7 gene was hypermethylated in 46% of the HCC cases being studied, indicating the possible involvement of its inactivation by the hypermethylation in HCC.
Hypomethylation of the promoter CpG island of the MAGEA1 gene may be essential to its HCC specific expression
Although the hypermethylation mediated gene silencing of the tumor suppressor genes has caught the major attention, the local hypomethylation has also recently been linked to reactivation of transcription of the genes that are hypermethylated and silenced in the normal tissues 9, 62. Therefore, we have also assessed whether the switching-on of the otherwise hypermethylated genes in the normal cells is linked to the demethylated state of the promoter CpG island in the HCC tissues. The MAGEA1 gene is such an example, expression of which is off in the normal hepatocytes and on in HCC31. Correlating well with such an expression profile, we in deed found that the promoter CpG island of the MAGEA1 (panel 13, Fig 1) was fully methylated in the normal liver tissue, became unmethylated at a similar frequency in 18/28 (64%) and 21/28 (75%) in each of the paired HCC tissue samples, respectively. Judged by the lack of the significance difference (c2=0.76, P=0.382), the demethylation of the promoter CpG island may more likely to be an early phase event of HCC carcinogenesis.
The stage-specific nature of the HCC associated changes in the methylation profiles of the promoter CpG islands of genes
It has been well recognized that the so-called non-cancerous cells defined under microscope may have already suffered some genetic lesions as have the corresponding cancerous tissues, theoutcome of the earlier events of carcinogenesis. Inclusion of the neighboring non-cancerous tissue in this study made it possible to analyzed our results from the stage-specific perspective of carcinogenesis. As shown in Tab 3, the following genes exhibited a similar frequency in changes of methylation pattern in the HCC tissues and the neighboring non-cancerous tissues from the pattern in the normal liver tissues from health volunteers: ABO (9/28, 32%; 14/28, 50%; x2= 1.845, P=0.174), CSPG2 (1/28, 4%; 5/28, 18%, x2= 2.987, P=0.084), GALR2(2/28, 7%; 6/28, 21%; x2= 2.292, P=0.13), MT1A (2/28, 7%, 3/28, 11%; x2=0.216, P=0.642), MYOD1( 15/28, 54%; 15/28, 54%; P=1), p57KIP2 (2/28, 7%; 4/28, 14%; x2=0.707, P=0.388), and WT1(4/28, 14%; 8/28, 29%; x2=1.697,P=0.193) as well as the MAGEA1 with having the HCC specific demethylation (18/28, 64%; 21/28, 75%; x2=0.76, P=0.383). Three of the four genes studied previously27 displayed a similar pattern. They were: the RASSF1a (24/29, 79%; 29/29, 100%; P=0.085), CDH13 (5/29, 17%; 6/29, 20%, x2= 0.112, P=0.738); and CASP8 (21/21, 100%; 21/21, 100%; P=1). The rest fell into the second category. They are the genes encoding AR (12/28, 43%; 20/29, 71%; x2=4.667, P=0.031), cyclin a1 (8/ 28, 29%; 15/28, 54%; x2=3.615, 0.057), DBCCR1 (3/28, 11%; 16/28, 57%; x2=13.462, p<0.001), IRF7(2/28, 7%; 12/28, 43%; x2=9.524, P=0.002), OCT6(14/28, 50%; 23/28, 82%; x2=6.452, P=0.011), and p73 (10/28, 36%; 19/28, 68%; x2=5.793, P=0.016). The corresponding figures of the p16INK4a in the previously study 27 were 6/29, 20%, and 16/29, 55%(x2=7.323, P=0.007). If the neighboring non-cancerous tissue may indeed have already suffered from the early stage genetic and/or epigenetic lesions during carcinogenesis, the HCC specific methylation changes of the ABO, CSPG2, GALR2, MT1A, MYOD1, WT1, MAGEA1, RASSF1a, CDH13 and CASP8 genes occurred at the early stage of the HCC carcinogenesis. On the contrary, the changes in the methylation pattern of the AR, cyclin a1, DBCCR1, IRF7, OCT6, p73, and p16INK4a genes was likely to be the late phase event of HCC carcinogenesis. The implication of this new analysis in the HCC remains to be explored in the future.
Association studies of the methylation profiles of the genes with the clinical-pathological parameters of HCC
After completing the information-gathering of the methylation profiles of forty four genes ( twenty four genes in this report and twenty genes in the previous report 27) it naturally follows to identify the association between the HCC changes in the methylation of each target with any given clinical-pathological parameters. By a stringent statistic evaluation with both χ2 and P tests, we looked for the association between the HCC specific methylation changes of each of the nineteen genes (including four genes in the previous study, 1) and various clinical-pathological parameters. Probably due to the relatively smaller size of patient samples, few association survived from such a scrutiny.
HCC is classified into two major subgroups, those associated with cirrhosis and those without, which differ from both pathological processes as well as the etiological profiles63, 64. Cirrhosis is a significant pre-HCC pathologic lesion and can be easily diagnosed by the non-invasive ultrasound method. In this connection, the hypermethylated cyclin a1 gene occurred more frequently in the non-cirrhosis HCC group (10/13, 76.9%) than in the cirrhosis HCC group (5/15, 20%, x2= 3.615, P=0.057). On the contrary, the hypermethylated p16INK4a gene was more prevalent in cirrhosis HCC patients (12/16, 75%) than the non-cirrhosis HCC patients (4/13, 30.3 %, x2=7.323, P=0.007). Although the underlying mechanisms whereby such a cirrhosis based differential methylation profile has been brought about, it's value for the prognostic evaluation of the clinical treatment of the HCC patients should not be overlooked. Furthermore, it is anticipated that more associations between the methylation of any given targets and the clinical-pathological indicators will be discovered when more HCC patients are subjected to such a study in future.
The concordant methylation behavior of the promoter CpG islands of the genes in HCC
As far as the number of the genes is concerned, our present studies are likely to be most comprehensive in the HCC field, to our best knowledge, which enable us to look into the concordant methylation behavior of the nineteen genes in the HCC group and in various sub-groups for the first time. Both occurrence and frequencies of the HCC specific changes in various subsets of two upward to nine genes among the nineteen genes (including four genes in previous study27) with the best sub-set selection method29. Although, CASP8 was hypermethylated in 21/29 samples tested, the remaining eight samples failed to be informative, we excluded the CASP8 gene from this analysis. As shown in Tab 5, the frequency of the two gene (the hypermethylated OCT6 and RASSF1a) was 82% and of three gene (the former two plus the hypermethylated p73) was 68% and of four gene subsets (the former three plus the demethylated MAGEA1) was 54% in the HCC patient group. In the cirrhosis-HCC groups, the frequency of the two genes (the hypermethylated RASSF1a and OCT6) was 87%, and three gene sub-sets (the former two genes plus the p16INK4a or p73) was 73%. The corresponding pattern of the non-cirrhosis patient group was different, the frequency of the two genes (the hypermethylated RASSF1a plus the hypomethylated MAGEA1 or the hyperme-thylated OCT6 or the hypermethylated cyclin a1) was 77% and of the three gene sub-sets (the hypermethylated RASSF1a plus hypermethylated OCT6 and cyclin a1) was 69%.
SUMMARIES
Tumor associated changes in the methylation profiles of the promoter CpG islands of, chiefly, the suppressor genes have been well documented, suggesting the possible role of the epigenetic mechanisms for gene inactivation, as an alternative to the genetic lesions, including deletion and mutation in tumors7, 65, 66. Therefore, methylation profiling would be a useful information gathering process for a better understanding of carcinogenesis as well as the better diagnostic, prognostic and even therapeutic measures against tumors. In this study, we expand further the list of targets from twenty27 to forty four known genes for methylation profiling in the normal healthy liver, HCC tissues and the paired normal tissues, representing a most comprehensive survey in the HCC.
Finally, the concordant methylation profiles of these nineteen genes summarized above (d-f, Tab 5) may be useful as the prognostic, possibly the diagnostic biomarkers for HCC. They may serve the good epigenetic markers to detect tumor cells from biopsies, serum, and so forth, if both sensitivity and specificity of these assays can be satisfactorily established. It may also be useful to determine the methylation status of these markers in circulating tumor cells in blood or predict the sensitivity to chemotherapy, and overall therapeutic outcomes of the HCC patients differing in cirrhosis status, grading and other clinical-pathological profiles.
References
Ferlay J, F Bray, P Pisani and D M Parkin . GOBOCAB 2000 Cancer Incidence, Mortality and Prevalence Worldwide,. Lyon: IARC Scienctific Publications, IARC Press, 2001
Parkin D, P Pisani and J Ferlay . Global cancer statistics. CA Cancer J. Clin., 1999; 49:33–64.
Cancer Incidence and Mortality in China, 1993–1997 (Selected Cities and Counties). Beijing: China Publishing House of Medical Sciences and Technologies, 1998
Feitelson M A, B Sun, N L Satiroglu Tufan, J Liu, J Pan and Z Lian . Genetic mechanisms of hepatocarcinogenesis. Oncogene 2002; 21(16):2593–604.
Kim J W and X W Wang . Gene expression profiling of preneoplastic liver disease and liver cancer: a new era for improved early detection and treatment of these deadly diseases? Carcinogenesis 2003; 24(3):363–9.
Wang X W, S P Hussain, T I Huo, et al. Molecular pathogenesis of human hepatocellular carcinoma. Toxicology 2002; 181–182:43–7.
Baylin S and T H Bestor . Altered methylation patterns in cancer cell genomes: cause or consequence? Cancer Cell 2002; 1(4):299–305.
Esteller M . Relevance of DNA methylation in the management of cancer. Lancet Oncol 2003; 4(6):351–8.
Esteller M and J G Herman . Cancer as an epigenetic disease: DNA methylation and chromatin alterations in human tumours. J Pathol 2002; 196(1):1–7.
Jones P A and D Takai . The role of DNA methylation in mammalian epigenetics. Science 2001; 293(5532):1068–70.
Jaenisch R and A Bird . Epigenetic regulation of gene expression: how the genome integrates intrinsic and environmental signals. Nat Genet 2003; 33 Suppl:245–54.
Jones P A . Epigenetics in carcinogenesis and cancer prevention. Ann N Y Acad Sci 2003; 983:213–9.
Geiman T M and K D Robertson . Chromatin remodeling, histone modifications, and DNA methylation-how does it all fit together? J Cell Biochem 2002; 87(2):117–25.
Feninberg A . Cancer epigenetics takes center stage. Proc. Natl. Acad. Sci. U S.A. 2001; 98(2):392–4.
Eden A, F Gaudet, A Waghmare and R Jaenisch . Chromosomal instability and tumors promoted by DNA hypomethylation. Science 2003; 300(5618):455.
Gaudet F, J G Hodgson, A Eden, et al. Induction of tumors in mice by genomic hypomethylation. Science 2003; 300(5618):489–92.
Chen R Z, U Pettersson, C Beard, L Jackson-Grusby and R Jaenisch . DNA hypomethylation leads to elevated mutation rates. Nature 1998; 395(6697):89–93.
Esteller M, M F Fraga, M Guo, et al. DNA methylation patterns in hereditary human cancers mimic sporadic tumorigenesis. Hum Mol Genet 2001; 10(26):3001–7.
Zochbauer-Muller S, K M Fong, A K Virmani, J Geradts, A F Gazdar and J D Minna . Aberrant promoter methylation of multiple genes in non-small cell lung cancers. Cancer Res 2001; 61(1):249–55.
Rosas S L, W Koch, M G da Costa Carvalho, et al. Promoter hypermethylation patterns of p16, O6-methylguanine-DNA-methyltransferase, and death-associated protein kinase in tumors and saliva of head and neck cancer patients. Cancer Res 2001; 61(3):939–42.
Foster S A, D J Wong, M T Barrett and D A Galloway . Inactivation of p16 in human mammary epithelial cells by CpG island methylation. Mol Cell Biol 1998; 18(4):1793–801.
Cui H, P Onyango, S Brandenburg, Y Wu, C L Hsieh and A P Feinberg . Loss of imprinting in colorectal cancer linked to hypomethylation of H19 and IGF2. Cancer Res 2002; 62(22):6442–6.
Cho B, H Lee, S Jeong, et al. Promoter hypomethylation of a novel cancer/testis antigen gene CAGE is correlated with its aberrant expression and is seen in premalignant stage of gastric carcinoma. Biochem Biophys Res Commun 2003; 307(1):52–63.
Hattori M, H Sakamoto, K Satoh and T Yamamoto . DNA demethylase is expressed in ovarian cancers and the expression correlates with demethylation of CpG sites in the promoter region of c-erbB-2 and survivin genes. Cancer Lett 2001; 169(2):155–64.
Hoare S F, L A Bryce, G B Wisman, et al. Lack of telomerase RNA gene hTERC expression in alternative lengthening of telomeres cells is associated with methylation of the hTERC promoter. Cancer Res 2001; 61(1):27–32.
Toyota M, N Ahuja, M Ohe-Toyota, J G Herman, S B Baylin and J P Issa . CpG island methylator phenotype in colorectal cancer. Proc Natl Acad Sci U S A 1999; 96(15):8681–6.
Yu J, M Ni, J Xu, et al. Methylation profiling of twenty promoter-CpG islands of genes which may contribute to hepatocellular carcinogenesis. BMC Cancer 2002; 2(1):29.
Liver Cancer Study Group of Japan. TNM classification for hepatocellular carcinoma by Liver Cancer Study group. World J Surg 1989; 13:212.
Miller A J . Subset Selection in Regression. New York Champman and Hall 1990.
Liu Q, Wang, W, & eds. Liver Cancer. Beijing: People's Medical Publishing House, 2000
Mou D C, S L Cai, J R Peng, et al. Evaluation of MAGE-1 and MAGE-3 as tumour-specific markers to detect blood dissemination of hepatocellular carcinoma cells. Br J Cancer 2002; 86(1):110–6.
Yin H and K Blanchard . DNA methylation represses the expression of the human erythopoietin gene by two diffferent mechanisms. Blood 2000; 95(1):111–9.
Sauter P and P Matthias . Coactivator OBF-1 makes selective contacts with both the POU-specific domain and the POU homeodomain and acts as a molecular clamp on DNA. Mol Cell Biol 1998; 18(12):7397–409.
Iwamoto S, D A Withers, K Handa and S Hakomori . Deletion of A-antigen in a human cancer cell line is associated with reduced promoter activity of CBF/NF-Y binding region, and possibly with enhanced DNA methylation of A transferase promoter. Glycoconj J 1999; 16(10):659–66.
Chen B, P Dias, J J Jenkins, 3rd, V H Savell and D M Parham . Methylation alterations of the MyoD1 upstream region are predictive of subclassification of human rhabdomyosarcomas. Am J Pathol 1998; 152(4):1071–9.
Hutchins A S, A C Mullen, H W Lee, et al. Gene silencing quantitatively controls the function of a developmental trans-activator. Mol Cell 2002; 10(1):81–91.
Reeben M, S Myohanen, M Saarma and H Prydz . Sequencing of the rat light neurofilament promoter reveals differences in methylation between expressing and non-expressing cell lines, but not tissues. Gene 1995; 157(1-2):325–9.
Kibel A S, M Christopher, D A Faith, G S Bova, P J Goodfellow and W B Isaacs . Methylation and mutational analysis of p27(kip1) in prostate carcinoma. Prostate 2001; 48(4):248–53.
Li Y, H Nagai, T Ohno, et al. Aberrant DNA methylation of p57(KIP2) gene in the promoter region in lymphoid malignancies of B-cell phenotype. Blood 2002; 100(7):2572–7.
Watanabe T, H Huang, M Nakamura, et al. Methylation of the p73 gene in gliomas. Acta Neuropathol (Berl) 2002; 104(4):357–62.
Laux D E, E M Curran, W V Welshons, D B Lubahn and T H Huang . Hypermethylation of the Wilms' tumor suppressor gene CpG island in human breast carcinomas. Breast Cancer Res Treat 1999; 56(1):35–43.
Wang S, T Hashemi, S Fried, A L Clemmons and B E Hawes . Differential intracellular signaling of the GalR1 and GalR2 galanin receptor subtypes. Biochemistry 1998; 37(19):6711–7.
Usmani B A, R Shen, M Janeczko, et al. Methylation of the neutral endopeptidase gene promoter in human prostate cancers. Clin Cancer Res 2000; 6(5):1664–70.
Venkataraman G M, M Yatin, R Marcinek and K B Ain . Restoration of iodide uptake in dedifferentiated thyroid carcinoma: relationship to human Na+/I-symporter gene methylation status. J Clin Endocrinol Metab 1999; 84(7):2449–57.
Cui J, L R Rohr, G Swanson, V O Speights, T Maxwell and A R Brothman . Hypermethylation of the caveolin-1 gene promoter in prostate cancer. Prostate 2001; 46(3):249–56.
Toyota M, C Ho, N Ahuja, et al. Identification of differentially methylated sequences in colorectal cancer by methylated CpG island amplification. Cancer Res 1999; 59(10):2307–12.
Sasaki M, Y Tanaka, G Perinchery, et al. Methylation and inactivation of estrogen, progesterone, and androgen receptors in prostate cancer. J Natl Cancer Inst 2002; 94(5):384–90.
Muller C, C Readhead, S Diederichs, et al. Methylation of the cyclin A1 promoter correlates with gene silencing in somatic cell lines, while tissue-specific expression of cyclin A1 is methylation independent. Mol Cell Biol 2000; 20(9):3316–29.
Rachmilewitz E A . The role of methylation in CML. Przegl Lek 2000; 57 Suppl 1:25–6.
Habuchi T, M Luscombe, P A Elder and M A Knowles . Structure and methylation-based silencing of a gene (DBCCR1) within a candidate bladder cancer tumor suppressor region at 9q32-q33. Genomics 1998; 48(3):277–88.
Ghoshal K, S Majumder, Z Li, X Dong and S T Jacob . Suppression of metallothionein gene expression in a rat hepatoma because of promoter-specific DNA methylation. J Biol Chem 2000; 275(1):539–47.
Yuasa Y . DNA methylation in cancer and ageing. Mech Ageing Dev 2002; 123(12):1649–54.
Richardson B . Impact of aging on DNA methylation. Ageing Res Rev 2003; 2(3):245–61.
Yamanaka M, M Watanabe, Y Yamada, et al. Altered methylation of multiple genes in carcinogenesis of the prostate. Int J Cancer 2003; 106(3):382–7.
Tavian D, G De Petro, A Pitozzi, N Portolani, S M Giulini and S Barlati . Androgen receptor mRNA under-expression in poorly differentiated human hepatocellular carcinoma. Histol Histopathol 2002; 17(4):1113–9.
Dong S, J C Pang, J Hu, L F Zhou and H K Ng . Transcriptional inactivation of TP73 expression in oligodendroglial tumors. Int J Cancer 2002; 98(3):370–5.
Loiseau H, J Arsaut and J Demotes-Mainard . p73 gene transcripts in human brain tumors: overexpression and altered splicing in ependymomas. Neurosci Lett 1999; 263(2-3):173–6.
Satoh Y, T Nakagawachi, H Nakadate, et al. Significant Reduction of WT1 Gene Expression, Possibly Due to Epigenetic Alteration in Wilms' Tumor. J Biochem (Tokyo) 2003; 133(3):303–8.
Tomita Y, V Bilim, N Hara, T Kasahara and K Takahashi . Role of IRF-1 and caspase-7 in IFN-gamma enhancement of Fas-mediated apoptosis in ACHN renal cell carcinoma cells. Int J Cancer 2003; 104(4):400–8.
Detjen K M, D Murphy, M Welzel, K Farwig, B Wiedenmann and S Rosewicz . Downregulation of p21(waf/cip-1) mediates apoptosis of human hepatocellular carcinoma cells in response to interferon-gamma. Exp Cell Res 2003; 282(2):78–89.
Lu R, W C Au, W S Yeow, N Hageman and P M Pitha . Regulation of the promoter activity of interferon regulatory factor-7 gene. Activation by interferon snd silencing by hyperme-thylation. J Biol Chem 2000; 275(41):31805–12.
Ehrlich M . DNA methylation in cancer: too much, but also too little. Oncogene 2002; 21(35):5400–13.
Su Q and P Bannasch . Relevance of hepatic preneoplasia for human hepatocarcinogenesis. Toxicol Pathol 2003; 31(1):126–33.
Kopp-Schneider A . Biostatistical evaluation of focal hepatic preneoplasia. Toxicol Pathol 2003; 31(1):121–5.
Baylin S B and J G Herman . DNA hypermethylation in tumorigenesis: epigenetics joins genetics. Trends Genet 2000; 16(4):168–74.
Bestor T H . Unanswered questions about the role of promoter methylation in carcinogenesis. Ann N Y Acad Sci 2003; 983:22–7.
Acknowledgements
Thanks are due to Jian Guo CHEN, Li Sheng ZHANG, Meng Chao WU and Su Shen ZHEN for providing the patient's samples, Jian Ren GU for his support at the early stage of this project and Jian Xin and Jianyin Guo for their assistance in the statistic treatment of the data. This work was supported by the National High Technology Research and Development Program of China (863 Program) (2001AA217011, 2002AA2Z3352), the Major State Basic Research Development Program of China (973 Program) (G1998051004), and the Science Foundation of Shanghai Municipal Government (02DJ14056) to JingDe ZHU.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
YU, J., ZHANG, H., MA, Z. et al. Methylation profiling of twenty four genes and the concordant methylation behaviours of nineteen genes that may contribute to hepatocellular carcinogenesis. Cell Res 13, 319–333 (2003). https://doi.org/10.1038/sj.cr.7290177
Issue Date:
DOI: https://doi.org/10.1038/sj.cr.7290177
Keywords
This article is cited by
-
Down-regulated GATA-1 up-regulates interferon regulatory factor 3 in lung adenocarcinoma
Scientific Reports (2017)
-
DNA methylation: potential biomarker in Hepatocellular Carcinoma
Biomarker Research (2014)
-
Genome-wide methylation profiling of the different stages of hepatitis B virus-related hepatocellular carcinoma development in plasma cell-free DNA reveals potential biomarkers for early detection and high-risk monitoring of hepatocellular carcinoma
Clinical Epigenetics (2014)
-
Analysis of Genetic Damage and Gene Polymorphism in Hepatocellular Carcinoma (HCC) Patients in a South Indian Population
Digestive Diseases and Sciences (2013)
-
Involvement of MyoD and c-myb in regulation of basal and estrogen-induced transcription activity of the BRCA1 gene
Breast Cancer Research and Treatment (2011)