Abstract
Microorganisms living at many sites in the human body compose a complex and dynamic community. Accumulating evidence suggests a significant role for microorganisms in cancer, and therapies that incorporate bacteria have been tried in various types of cancer. We previously demonstrated that cupredoxin azurin secreted by the opportunistic pathogen Pseudomonas aeruginosa, enters human cancer cells and induces apoptotic death1,2,3,4. However, the physiological interactions between P. aeruginosa and humans and their role in tumor homeostasis are largely unknown. Here, we show that P. aeruginosa upregulated azurin secretion in response to increasing numbers of and proximity to cancer cells. Conversely, cancer cells upregulated aldolase A secretion in response to increasing proximity to P. aeruginosa, which also correlated with enhanced P. aeruginosa adherence to cancer cells. Additionally, we show that cancer patients had detectable P. aeruginosa and azurin in their tumors and exhibited increased overall survival when they did, and that azurin administration reduced tumor growth in transgenic mice. Our results suggest host–bacterial symbiotic mutualism acting as a diverse adjunct to the host defense system via inter-kingdom communication mediated by the evolutionarily conserved proteins azurin and human aldolase A. This improved understanding of the symbiotic relationship of bacteria with humans indicates the potential contribution to tumor homeostasis.
Similar content being viewed by others
Introduction
Microorganisms, particularly pathogenic bacteria, were used in the treatment of various types of human cancer over 100 years ago5,6 under the premise that a toxin produced by given pathogenic bacteria will inhibit the growth and spread of human cancer7. With the increasing knowledge of the role of the microbiota and the increasing incidence of cancer, treatment options using this approach have been reevaluated8,9,10,11,12,13.
We have been investigating the therapeutic role of microbes, such as the opportunistic pathogen Pseudomonas aeruginosa, in the management of human cancers1,2,3,4,14,15. Although P. aeruginosa is a major pathogen in cystic fibrosis (CF) causing significant morbidity and mortality16, CF patients have a lower incidence of melanoma and breast cancer than non-CF patients17,18. When supernatants of P. aeruginosa culture medium were co-incubated with tumor-derived J774A.1 macrophages, they induced apoptosis in a dose-dependent manner in J774A.12. The supernatants of the culture medium contained a high concentration of the P. aeruginosa-secreted redox protein azurin19,20.
Azurin is a 14 kDa periplasmic copper protein containing 128 amino acids (aa) that are found in several types of bacteria and blue-green algae21,22. One of its known biological functions is as an electron transfer protein in anaerobic energy production via nitrogen fixation23. In vitro studies have demonstrated that azurin induces apoptotic cell death in a variety of cancer cells, with minimal or no effect on their normal counterparts1,2,3,14,24,25,26. Upon preferential entry into cancer cells, azurin binds to the tumor suppressor protein p53 and induces caspase-mediated apoptosis1,3,27. Human xenotransplant studies in athymic mice showed that systemic administration of azurin inhibited tumor growth without significant adverse effects on the host1,3,28. In addition, we previously identified the active peptide fragment of azurin (i.e., p28 or Azu28)24,29, and the efficacy of this 28-aa cell-penetrating peptide p28 has been extensively investigated both experimentally and clinically30,31,32.
Although the clinical application and usefulness of azurin and its 28-aa peptide have been recognized, the physiological roles of P. aeruginosa in relationship with the host when it encounters cancer cells—which in turn inhibit cancer growth and spread—have not been investigated. The primary objective of this study is to demonstrate the interaction of P. aeruginosa with human cancer cells and its role in tumor homeostasis. Besides highlighting the importance of this specific bacterial-cancer interaction, we show for the first time the bidirectional regulation of bacterial-cancer communication in relation to the potential for P. aeruginosa-secreted azurin to inhibit tumor growth. Our observations suggest that these interactions may act as diverse adjuncts to the host defense system via inter-kingdom communication.
Results
High levels of P. aeruginosa azurin in the sera of CF patients
Based on the frequent colonization of CF patients with P. aeruginosa, the lower incidence of melanoma and breast cancer in CF than non-CF patients17,18 and previous descriptions of cancer-specific effects of azurin1,28,33,34,35,36, we hypothesized that azurin from P. aeruginosa plays a role in tumor development. To test our hypothesis that a bacterial protein, azurin, is detectable in human sera and that there is a significant elevation of azurin levels in CF patients, we first compared the serum levels of azurin in CF patients with chronic Pseudomonas infection to those in healthy volunteers with no disease (controls; see Supplementary Fig. 1a for demographic information) and found that azurin levels in CF patients were significantly higher than those in controls (Supplementary Fig. 1b). The serum level of azurin was correlated with the age of CF patients (Supplementary Fig. 1c). This clinical evidence showed that CF patients have significantly elevated levels of circulating azurin.
P. aeruginosa azurin secretion is stimulated by cancer cells
To investigate how host cells, either human cancer cells or their corresponding normal counterparts, affect P. aeruginosa azurin secretion, azurin secretion levels were measured in P. aeruginosa-host cell cultures. Azurin secretion was significantly higher, as were the transcription levels of the azurin-encoding gene azu, in the presence of various human cancer cell lines than in the presence of their normal counterparts (Fig. 1a, Supplementary Fig. 2). The evidence that the incidence rates of melanoma and breast cancer are lower in CF patients17,18 and that these two types of cancers showed high contrast in azurin secretion in our experimental systems1,3 led us to focus on these cancers for the remainder of this study. Azurin secretion by P. aeruginosa was positively correlated with the number of human breast cancer (Fig. 1b) or melanoma (Fig. 1c) cells, and azurin secretion was significantly lower when P. aeruginosa was incubated with normal breast cells or benign nevus (congenital melanocytic nevus, CMN) cells (Fig. 1a–c). These results suggest that P. aeruginosa preferentially secretes azurin in the presence of cancer cells and that this effect has a dose-dependent relationship with the population of cancer cells.
a Induction of azurin secretion by P. aeruginosa in the presence of host cells. Human cells and P. aeruginosa were co-incubated for 30 min at a concentration of 500,000 human cells/ml and P. aeruginosa (Pa) at an optical density (OD) = 0.3. Human cell lines derived from various tissues and tumors were used: melanoma Mel-2, congenital melanocytic nevus CMN, prostate cancer DU-145, normal prostate CRL-11611, ovarian cancer SK-OV3, normal ovary HOSE6-3, breast cancer MDA-MB-231, and normal breast MCF-10A. Secretion of azurin by P. aeruginosa into the culture supernatant was assessed by western blot analysis; the graph shows the observed band intensities determined by a densitometer UN-SCAN-IT gel version 5.1. The mean+SE values were calculated for skin, prostate, ovarian, and breast cell pairs. b, c Azurin secretion is cancer cell dose-dependent. P. aeruginosa secretes higher levels of azurin in the presence of human breast cancer MDA-MB-231 (b) and melanoma Mel-2 cells (c) than in the presence of melanocytes (CMN) or non-malignant MCF-10A breast cells, in a dose-dependent manner. Cancer and normal/non-malignant cells were co-incubated with P. aeruginosa for 30 min at concentrations ranging from 0 to 2,000,000 cells/ml and P. aeruginosa at OD = 0.3. Secretion of azurin by P. aeruginosa into the culture supernatant was assessed by western blot analysis; the graph shows the observed band intensities. The mean+SE values were calculated. d Soluble extracellular factors stimulate azurin secretion in a distance-dependent manner. Mel-2 and P. aeruginosa cells were separated by a permeable Transwell® insert membrane with a pore size of 0.4 μm. In this co-culture system, P. aeruginosa (OD = 0.3) and Mel-2 cells (3,000,000 cells/ml) were separated at distances ranging from 2 mm to 12 mm and incubated for 30 min at 37 °C (inset figure: schematic of the co-culture system. Green: P. aeruginosa, blue: Mel-2). Secretion of azurin by P. aeruginosa into the culture supernatant was assessed by western blot analysis; the graph shows the observed band intensities.
We next determined whether the bacteria-cancer interaction-mediated modulation of azurin secretion requires direct cell-to-cell contact between P. aeruginosa and cancer cells. Secreted azurin levels were highest at the shortest distance (2 mm) between the bacteria and cancer cells and quite low at distances greater than 12 mm, showing an inversely proportional correlation (Fig. 1d). Despite the lack of physical contact, azurin secretion was elicited, thereby indicating that a soluble factor secreted by Mel-2 cells may act as a stimulus. Changes in azurin levels were not due to P. aeruginosa contamination from top to bottom wells (Supplementary Fig. 3). These results suggest the existence of at least one soluble agent originating from cancer cells that exerts a concentration-dependent stimulatory effect on azurin secretion.
Cancer cells secrete aldolase A in response to azurin
The significant stimulation of azurin secretion by malignant host cells compared to that by their normal counterparts suggested that these cell groups differed in the secretion of a host factor. To identify any proteins that were differentially secreted in the presence of P. aeruginosa, we used a mass spectrometry-based proteomic approach. We found that human aldolase A was released extracellularly into the culture medium when human cancer cells or their normal counterparts were co-incubated with P. aeruginosa (Supplementary Fig. 4a–c).
Aldolase, also called fructose-bisphosphate aldolase (FBA), is a glycolytic enzyme involved in the Embden-Meyerhof-Parnas glycolytic pathway and gluconeogenesis and is highly conserved in bacterial, archaeal, and eukaryotic organisms. Human aldolase A secretion by human cancer cells in the presence of P. aeruginosa was significantly higher than that by their normal counterparts (Fig. 2a). Changes in aldolase A secretion were not due to alteration of the cell growth rate or induction of toxicity in cancer cells (Supplementary Fig. 5a, b). The secretion levels of aldolase A in culture medium were related to the secretion levels of azurin when human malignant and benign cells derived from various tumors and tissues were co-incubated with P. aeruginosa (Fig. 2b). To address the role of azurin on aldolase A secretion, three experimental approaches were taken: aldolase A secretion induced by (1) wild type (WT) and azu gene deleted mutant P. aeruginosa37, (2) the non-pathogenic E. coli expressing azurin gene, and (3) recombinant azurin protein. First, aldolase A secretion was measured when cancer cells were co-incubated with WT and azunull mutant P. aeruginosa. Aldolase A secretion induced by WT P. aeruginosa was significantly higher than azunull mutant, but azunull mutant can still induced aldolase A secretion from cancer cells (Fig. 2c, Supplementary Fig. 6). Next, the interaction of the non-pathogenic E. coli laboratory strain JM109 (a K-12 derivative) transformed with the azurin-encoding gene with cancer cells was evaluated to determine whether the secretion of azurin and aldolase A is specific to the P. aeruginosa strain we used. E. coli expressing azu gene secreted a considerable amount of azurin in the presence of Mel-2 cells. Additionally, aldolase A secretion from Mel-2 cells was induced by azu-expressing E. coli (Supplementary Fig. 7), suggesting that (1) azurin is a major inducer of aldolase A secretion, but not a sole inducer from P. aeruginosa, and (2) E. coli has a secretion mechanism similar to that of P. aeruginosa.
a Induction of aldolase A secretion by host cells in the presence of P. aeruginosa. Human host cells (cancer and normal) and P. aeruginosa were co-incubated for 30 min at a concentration of 500,000 human cells/ml with P. aeruginosa at OD = 0.3. Aldolase secretion varied between the co-cultures with cancer and normal cells. Secretion of azurin and aldolase into the culture supernatant was assessed by western blot analysis; the graph shows the observed band intensities. b Correlation between the aldolase A and azurin levels in co-cultures of P. aeruginosa with cancer or normal cells. c Cancer cells (Mel-2 and MDA-MB-231) and P. aeruginosa (WT and azunull mutant) were co-incubated for 30 min at a concentration of 500,000 human cells/ml with P. aeruginosa at OD = 0.3. Secretion of aldolase into the culture supernatant was assessed by western blot analysis. d P. aeruginosa (OD = 0.3) was treated with purified aldolase A protein at concentrations of 1 nM, 10 nM, 100 nM, and 1 μM for 30 min. Treatment with 1 μM aldolase A stimulated azurin secretion from P. aeruginosa, suggesting that aldolase A is a stimulatory factor for azurin secretion. Secretion of azurin by P. aeruginosa into the culture supernatant was assessed by western blot analysis; the graph shows the observed band intensities. MDA-MB-231 (e) and Mel-2 (f) cells were treated with purified azurin protein at concentrations of 100 nM, 10 μM, and 1 mM for 30 min. Secretion of aldolase A by cancer cells into the culture supernatant was assessed by western blot analysis; the graph shows the observed band intensities. P. aeruginosa (Pa only) did not show any signal as it did not secret any proteins that cross-react with anti-aldolase A antibody. g Aldolase A secretion in the presence of P. aeruginosa is distance-dependent. Mel-2 cells and P. aeruginosa were separated with a permeable Transwell® insert. In this co-culture system, P. aeruginosa (OD = 0.3) and Mel-2 cells (3,000,000 cells/ml) were separated at distances ranging from 0 mm to 12 mm and incubated for 30 min at 37 °C. Aldolase A secretion decreased as the distance between the two cell populations increased. Secretion of aldolase by Mel-2 cells into the culture supernatant was assessed by western blot analysis; the graph shows the observed band intensities.
To further investigate the impact of human aldolase A on azurin secretion during the interaction of bacteria and human cancer cells, P. aeruginosa cells were cultured in the presence of purified human aldolase A (purity >95%), and azurin secretion was measured by western blotting. Exposure of P. aeruginosa to aldolase A induced azurin secretion (Fig. 2d). Conversely, when breast cancer and melanoma cells were cultured in the presence of purified azurin, these cancer cells secreted a considerable amount of human aldolase A into the culture medium (Fig. 2e, f) without increasing gene expression or intracellular levels of aldolase A, suggesting that aldolase A secretion induced by azurin occurs in a transcription-independent manner (Supplementary Fig. 8a, b). Similar to the pattern of azurin secretion shown in Fig. 1d, the levels of secreted aldolase A were highest at the shortest distance (2 mm) between P. aeruginosa and cancer cells (Fig. 2g), that secreted azurin has a major role to induce aldolase A secretion, and their interaction is bidirectional and concentration density-dependent inter-communication between P. aeruginosa and cancer cells via azurin and aldolase A (Fig. 2).
Aldolase A promotes P. aeruginosa localization on cancer cells
Although the cytosolic role of aldolase in glycolysis and gluconeogenesis has long been recognized, aldolase is reportedly also involved in host cell adhesion and biofilm formation in bacteria and parasites such as Streptococcus, Neisseria, Toxoplasma and Plasmodium38,39,40,41,42. This prompted us to investigate whether aldolase A secreted by human cancer cells plays a similar biological role in the adherence of P. aeruginosa to cancer cells38,43. First, we tested whether silencing aldolase A gene will alter the adherence of P. aeruginosa to cancer cells. The P. aeruginosa adhesion assay showed that siRNA-induced silencing aldolase A in MDA-MB-231 and Mel-2 cells (Supplementary Fig. 9a) significantly reduced the adherence of P. aeruginosa (Fig. 3a). Conversely, in the presence of purified human aldolase A, P. aeruginosa exhibited significantly increased adherence to cancer cells, and the increase was dose-dependent and saturable (>1 µM) (Fig. 3b). It has been reported that MUC1, an O-glycosylated membrane-tethered mucin on cancer cells, interacts with P. aeruginosa through flagellin44,45. Muc1‒/‒ animals displayed ~50% less adherence of P. aeruginosa in the lungs compared with Muc1+/+ mice46. Hence, we tested the effect of siRNA-induced MUC1 silencing in cancer cells (Supplementary Fig. 9b) by our P. aeruginosa adhesion assay. Silencing MUC1 in MDA-MB-231 and Mel-2 cells significantly reduced the adherence of P. aeruginosa (Fig. 3c). Moreover, when recombinant aldolase A was added to MUC1 silenced cancer cells, the adhesion rate of P. aeruginosa was similar to control values, suggesting that aldolase-mediated adherence of P. aeruginosa was, at least partly, independent of MUC1-mediated adhesion.
a Adherence assays were performed essentially as previously described38. Monolayer MDA-MB-231 (red) and Mel-2 (blue) cells were compared to siRNA-induced silencing of aldolase gene in the cancer cell lines (+) when they were co-incubated with P. aeruginosa for 2 h. To assess total cell association, monolayers were washed to remove unbound P. aeruginosa and were then disrupted and homogenized in 0.1% saponin/PBS. P. aeruginosa cells were counted by serial dilution of the homogenized suspensions and subsequent determination of colony-forming units (CFU) by plating on LB agar. Control (–) expressed as 100%. Mean+SE, *P < 0.05. b Monolayer MDA-MB-231 (red) and Mel-2 (blue) cells were co-incubated with P. aeruginosa in the presence or absence of exogenous aldolase A for 2 h. Similar to above, total P. aeruginosa association on cancer cells were counted by plating on LB agar. Control (0 µM aldolase A) expressed as 100%. c Monolayer MDA-MB-231 (red) and Mel-2 (blue) cells were compared to siRNA-induced silencing of MUC1 genes in the cancer cell lines (+) when they were co-incubated with P. aeruginosa in the presence or absence of exogenous aldolase A at 200 nM for 2 h. Total P. aeruginosa association on cancer cells were counted by plating on LB agar. *P < 0.05, NS: not significant. Azurin secretion is P. aeruginosa dose-dependent. MDA-MB-231 (d) and Mel-2 (e) cells (500,000 cells/ml) were co-incubated with P. aeruginosa at concentrations corresponding to an OD ranging from 0.0 to 0.6. Secretion of azurin by P. aeruginosa into the culture supernatant was assessed by western blot analysis; the graph shows the observed band intensities. The mean+SE values were calculated.
These results suggest that aldolase A induces P. aeruginosa adhesion and colonization in cancer cells. Although mucins have been suggested as the preferred sites for adherence and colonization of P. aeruginosa on host cells47, P. aeruginosa can also bind to several other host proteins. These proteins include aldolase, and this binding is mediated by hydrophobic interactions48.
Azurin secretion is P. aeruginosa density dependent
The complexity of P. aeruginosa genome reflects evolutionary adaptation permitting it to thrive in diverse environments, including eukaryotic hosts, with which it has coexisted for millions of years49. Pseudomonas spp. are renowned among prokaryotes for their complex quorum sensing (QS) systems that regulate biofilm formation50,51. In general, bacterial QS of stimuli and responses correlates to population density52. Based on our data, P. aeruginosa secretes low levels of azurin due to the presence of only a small population of either cancer cells (Fig. 1b, c) or P. aeruginosa (Fig. 3d, e). This finding suggests that a high concentration of aldolase A due to a large population of cancer cells enhances azurin secretion by increasing bacterial adherence and the density of the bacterial population. However, this mode of density sensing in P. aeruginosa is likely independent of the intercellular QS communication system since it has been previously shown that azurin is actively expressed in Pseudomonas mutants carrying mutations in the Gac/Rsm system that activates the QS machinery primarily by stimulating N-butanoyl-l-homoserine lactone (C4-HSL) production53. Additionally, azurin expression is not regulated as a virulence factor in P. aeruginosa, as GacA is a major positive regulator of virulence in P. aeruginosa53. Similar to the relationship between Burkholderia (previously considered members of the Pseudomonas genus) and plants54, this QS-independent mechanism appears to play important roles in the P. aeruginosa-cancer interaction. Additional studies need to be carried out to conclusively determine a mechanism of sensing in P. aeruginosa.
P. aeruginosa is abundant in primary tumors, and its product azurin inhibits tumor growth
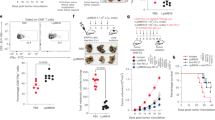
Our in vitro results indicated that P. aeruginosa and cancer cells communicate with each other through the secreted proteins azurin and aldolase A, leading to the localization of P. aeruginosa on cancer cells (Figs. 1–3). This effect might be applicable to clinical settings. To measure P. aeruginosa localization in human melanoma and breast tumors, PCR with azu-specific primers was carried out. We included both primary and metastatic tumors, as their different characteristics may affect P. aeruginosa localization55. Among tumors from patients with melanoma (primary: N = 29, age range = 21–84 years old; metastatic: N = 34, age range = 24–85 years old) and breast cancer (primary: N = 22, age range = 30–81 years old; metastatic: N = 14, age range = 28–79 years old) (Supplementary Fig. 10), the P. aeruginosa azu gene was detectable in 27.6% of primary melanomas and 5.9% of metastatic melanomas (Fig. 4a). Primary breast cancers showed a higher positive rate for the azu gene than metastatic breast tumors (22.7% vs. 14.3%, Fig. 4b). The presence of P. aeruginosa and its product azurin was further confirmed by transmission electron microscopy (TEM) within azu-positive melanoma cells stained by an anti-P. aeruginosa antibody or anti-azurin antibody (Fig. 4c, d, Supplementary Fig. 11a–c). P. aeruginosa cells were found in the cytoplasm, and azurin was localized in both the cytoplasm and nucleus. This finding is consistent with those of our preclinical studies showing that azurin localizes in the nucleus1,3. These results indicate that P. aeruginosa was detectable in human tumors, raising the possibility that, in some individuals, azurin-producing P. aeruginosa may affect the biological activity of tumors through bacterial-cancer interactions. To test this possibility, we compared survival between patients with azu-positive and those with azu-negative primary melanoma and breast cancer tumors. In both melanoma (Fig. 4e) and breast cancer (Fig. 4f), patients with azu-positive tumors had better overall survival times than patients with azu-negative tumors, suggesting that P. aeruginosa azurin positively affects the prognosis of patients, at least for those with melanoma or breast cancer. Based on these findings, we intend to design clinical studies, which are needed to examine whether the presence of azurin-expressing P. aeruginosa is an important factor for patient prognosis and can be used as a biomarker.
The azurin-encoding gene (azu) was detected in tumors from patients with breast cancer and melanoma. It was amplified by PCR with P. aeruginosa azu-specific primers and confirmed by DNA sequencing. The amplified PCR product, as a single band, was sequenced and showed 100% identity to P. aeruginosa azu. In melanoma (a), 27.6% (8 of 29) of primary tumors and 5.9% (2 of 34) of metastatic tumors were azu positive (P < 0.05). In breast cancer (b), 22.7% (5 of 22) of primary tumors and 14.3% (2 of 14) of metastatic tumors were azu positive (P > 0.05). In addition, frozen tumor samples were processed for immunogold transmission electron microscopy (TEM) by using anti-P. aeruginosa and anti-azurin antibodies. TEM images of uranyl acetate-stained sections (arrowheads) showed the intracellular localization of P. aeruginosa (c) and its product azurin (d) in human melanoma sections that were azu positive by PCR. Magnification: ×3000. e, f Patients with azu-positive tumors displayed increased survival. Survival analysis of patients with azu-positive vs. azu-negative primary melanoma (e, P < 0.01) and primary breast cancer (f, P < 0.05) tumors. g Hemizygous MMTV-PyMT transgenic mice were injected with either PBS control or 2.5 mg/kg azurin (intraperitoneally, 3× weekly) for 2 months. The mean+SE values of tumor volume were calculated. h Tumor-free survival curve was generated with the PBS control and 2.5 mg/kg azurin groups of MMTV-PyMT transgenic mice. Median tumor-free survival of the PBS control and azurin-treatment groups was 67 and 72 days, respectively. P = 0.015.
In this study, we used a transgenic mouse model that spontaneously develops mammary tumors to investigate the impact of P. aeruginosa azurin on cancer in vivo. Based on the critical role of azurin in P. aeruginosa–cancer interactions (Figs. 1–3), purified P. aeruginosa azurin was injected into transgenic mice that spontaneously develop mammary tumors56,57. An in vitro experiment indicated that P. aeruginosa azurin secretion was significantly higher in the presence of triple-negative MDA-MB-231 cells than in the presence of T-47D(ER+, PR+, Her2−) cells (Supplementary Fig. 12); thus, triple-negative, p53wt MMTV-PyMT transgenic mice were used. When these mice were exposed to azurin, NIR dye-conjugated azurin preferentially localized to spontaneously developed mammary tumors (Supplementary Fig. 13a). Tumor growth and tumor weight were significantly inhibited without affecting body weight (Fig. 4g, Supplementary Fig. 13b, c), and tumor-free survival was significantly extended in the azurin-treatment group (Fig. 4h). To the best of our knowledge, these findings suggest a new role for the bacterium P. aeruginosa in host defense via azurin production.
Discussion
In this study, we demonstrated the bidirectional regulation of bacterial-cancer communication in relation to the potential for P. aeruginosa-secreted azurin to inhibit tumor growth. Human cancer cells upregulated aldolase A secretion in response to increasing proximity to P. aeruginosa/azurin, which correlated with enhanced P. aeruginosa adherence to the cancer cells. Our results also show that cancer patients had detectable P. aeruginosa and azurin in their tumors and exhibited increased overall survival when they did. Finally, our results suggest host–bacterial symbiotic mutualism acting as a diverse adjunct to the host defense system on tumor homeostasis.
Increasing evidence suggests that this symbiotic host-microbiome relationship is dependent on the nature of microorganism-host interactions. For example, infectious agents cause ~20% of cancers worldwide58. Helicobacter pylori colonizes ~50% of people worldwide59 and cause ~90% of gastric cancers58. On the other hand, H. pylori colonization can benefit human health, because decolonization increases the risk of severe gastroesophageal reflux disease60. The relationship of Pseudomonas spp. with plants has been extensively studied. Plants develop in a microbe-rich environment and interact with microorganisms, such as Pseudomonas spp. (e.g., P. aeruginosa, P. fluorescens and P. syringae). Some bacteria, including Pseudomonas spp., can be either beneficial or pathogenic61,62. Collectively, this evidence suggests that some bacteria might be double-faced microbes in a symbiotic relationship that can both harm (parasitism) and benefit (mutualism) the host. It is still poorly understood exactly how host–microbe relationships were initiated; however, a previous report revealed that mutualism in proteobacteria, including Pseudomonas, has evolved independently between 34 and 39 times, implying that this arrangement is evolutionarily favored63. Those findings also indicate that in bacteria, mutualism has arisen most often in species that were originally parasites and pathogens63,64. Taken together, these results suggest that through evolution, P. aeruginosa has acquired abilities to both harm (parasitism) and benefit human health (mutualism) and adjusts its behavior depending on physiological/cellular conditions that can change depending on disease status61. Our results suggest that P. aeruginosa can have a symbiotic relationship with humans characterized by both mutualism and parasitism. In its parasitic relationship, P. aeruginosa, an opportunistic pathogen, increases morbidity and mortality in patients with CF, while in its mutualistic relationship, P. aeruginosa can benefit human health by suppressing65 and attacking1,3,4 malignant cells through secretion of the bacterial protein azurin in hosts harboring malignant cells (Supplementary Fig. 14).
P. aeruginosa, a common gram-negative, rod-shaped bacterium, can be found in numerous moist sites in soils and plants and in a wide variety of aquatic environments but also, as an opportunistic pathogen, causes serious infection under certain circumstances (e.g., in hospitalized patients with burns or CF adults with multi-resistant pseudomonas)66. Infection with P. aeruginosa continues to be a threat to cancer patients, particularly when patients have weakened immune activity due to cancer treatment. In contrast, P. aeruginosa, as a potential commensal bacterium, can be found in the skin of some healthy individuals and has been isolated from the throat (5%) and stool (3%) of non-hospitalized people67. In our study, P. aeruginosa expressing azu was found in tumors of primary melanoma and breast cancer patients who did not receive chemotherapy before specimen collection. Furthermore, our preliminary data on human tumors indicated that azu-positive patients had longer overall survival times than azu-negative patients, suggesting that P. aeruginosa localized in tumors may positively influence cancer prognosis. This possibility needs to be investigated in larger-scale clinical studies.
The bacteria-based strategies have great promise in the treatment of tumors68. Various bacterial species including P. aeruginosa showed potent anti-tumor efficacy with or without modulating immune activity69. A recent report suggested that factors secreted from P. aeruginosa altered the proliferation of breast cancer cells and enhanced the activity of doxorubicin, suggesting that the signaling stimulated by factors secreted from P. aeruginosais dependent on the abundance of P. aeruginosa70. This evidence was consistent with our data that azurin secretion by P. aeruginosa is population- and distance-dependent (Figs. 1, 3). Furthermore, several proteins from P. aeruginosa such as azurin, exotoxin A (ExoA), exoenzyme T (ExoT), Pacaspase recruitment domain (Pa-CARD) have been found to exert potent cytotoxicity against cancer69. Moreover, azurin can enhance the efficacy of standard chemotherapeutic agents such as 5-fluorouracil (5-FU)71. Since P. aeruginosa also produces numerous virulence factors49, it is logical to design potential anticancer agents based on the cupredoxin family of proteins30,31,32. As we previously reported, a peptide fragment identified in azurin (a.k.a. p28) enhances the activity of doxorubicin though activation of the p53-p21 axis29.
In conclusion, we attempted to understand the interaction of P. aeruginosa with human cancer cells and its role in tumor homeostasis. We demonstrated that molecular determinants of host–bacterial mutualism act as diverse adjuncts to the host defense system through inter-kingdom signaling by evolutionarily conserved proteins, the bacterial cupredoxin azurin and human aldolase A. To the best of our knowledge, these data on a host–microbe interaction provided novel insight into the symbiotic strategies based on the microbial cupredoxin azurin in modulating host defense mechanisms, which occurred independent of immune system stimulation. Our results highlight the importance of the bacteria-cancer interaction and will be helpful for guiding further studies to understand the differences between pathogens and beneficial microorganisms. Since evidence that microbes play an important role in establishing host homeostasis (e.g., tumor regression) is increasing, a better understanding of the symbiotic relationship of such bacteria with humans offers enormous potential to develop therapeutic strategies for human diseases such as cancer.
Methods
P. aeruginosa and human cell lines
P. aeruginosa strain 8822 isolated from the sputum of a CF patient is a generous gift from Dr. Ananda M. Chakrabarty72. Human cell lines of prostate cancer (DU-145), normal prostate (CRL-11611), ovarian cancer (SK-OV3), normal ovary (HOSE6-3); breast cancer (MCF-7, T-47D, MDA-MB-231), normal breast (MCF-10A) were purchased from the American Tissue Culture Collection (ATCC, VA). Human melanoma (UISO-Mel-2) and CMN cell lines were developed in our laboratory as described73,74.
ELISA
CF subjects were patients followed at the Adult CF Clinic at Emory University who had signed informed consent to provide blood samples in accordance with the Emory University IRB (Emory #00042577). Control sera from no-disease volunteers were obtained from Discovery Life Sciences (Los Osos, CA). Serum samples (N = 50 in each group) in carbonate bicarbonate buffer (pH 9.4) were used to coat 96-well plates (MaxiSorb, Thermo Fisher) in triplicate. Standard curves as internal controls were generated by using purified azurin75 to coat 96-well plates. Polyclonal rabbit anti-azurin antibody1,2 and alkaline phosphatase-conjugated secondary anti-rabbit antibody (SigmaMillipore) were used to detect azurin.
Growth media
All human cell lines were maintained in MEME with 10% FBS except for MCF-10A which was maintained in DMEM with 10% FBS. Luria-Bertani medium was used to grow P. aeruginosa before transfer to the experiment’s medium of 0.5% Minimal Glucose Medium (MGM) without any antibiotics.
P. aeruginosa quantitation
This was conducted using the turbidimetry method after the establishment of the standard growth curve of P. aeruginosa in 0.5% MGM. An optic density (OD) of 0.3 correlated with the mid-log phase of the bacterial growth and was chosen as the standard OD at 600 nm for the entire experiment.
Expression and purification of azurin: Cloning, expression of azu gene in E. coli and purification of recombinant azurin were described previously1.
P. aeruginosa and human cancer cells co-incubation
P. aeruginosa strain 8822 were grown overnight in LB medium at 37 °C then transferred at 1:100 v:v into 0.5% MGM without adding antibiotics. The OD of the medium was measured until the goal OD for the experiment was reached. Human cells were grown in the appropriate medium as reported above. Cells were trypsinized and counted using a Coulter Counter® Cell and Particle Analyzer. The needed number of cells was washed 2× with PBS and finally suspended in 0.5% MGM. For direct co-incubation assays, human cells and P. aeruginosa were co-incubated for 30 min at a concentration of 500,000 human cells/ml and P. aeruginosa at an optical density (OD) = 0.3. The needed number of cells was transferred based on the specifics of each experiment. For the indirect co-incubation assays, Corning Transwell® polyester membrane cell culture plates and inserts (TC-treated, sterile 24 mm Transwell with 0.4 µm pores, SigmaMillipore) were used. Human cells were immobilized in the lower compartment (well) whereas bacteria were incubated in the upper compartment (insert) at 37 °C for 30 min with a semi-permeable membrane of 0.4 µm pores separating the two compartments. Wild type (WT) P. aeruginosa PAO1 and its azunull mutant cells were used for aldolase secretion assay. Mutant P. aeruginosa was maintained on LB agar plates containing spectinomycin37. Other experimental conditions were the same as above.
Protein extraction
Comparative analyses of secreted proteins in media were performed as described previously76,77,78. The same volume of medium was collected from cell suspensions (direct co-incubation assays) or the lower compartment well (indirect co-incubation assays) after incubation and centrifuged at high speed (12,000 × g). The supernatant was then filtered using an Amicon® 0.22 µm pore filter. The equal volume of trichloroacetic acid (TCA; final concentration of 20%) was added to the supernatant and incubated on ice for 30 min. After spinning at 18,000 × g, TCA was decanted, and 100% acetone precipitation was applied twice with 5 min incubation. At the final step, acetone was decanted, and protein pellets were allowed to dry at room temperature for 15 min and resuspended in PBS. The equal volume of each sample was loaded per lane.
Western blotting
After running the NuPAGE, proteins were transferred to PVDF membranes (BioRad Laboratories Inc, Hercules, CA) which were blocked in SuperBlock (T20 TBS buffer, Thermo Fisher) for one hour. For secreted proteins, equal loading was confirmed by Ponceau staining of the membranes79. The PVDF membranes were incubated in a polyclonal rabbit anti-azurin antibody1 (1:5000) or a monoclonal mouse anti-aldolase A (1:200, Santa Cruz Biotechnology, TX) at 4 °C overnight. For cell lysates, GAPDH was used as a loading control (Novus Biologicals). The secondary antibody was applied (1:1000, polyclonal goat anti-rabbit IgG-HRP; Santa Cruz Biotechnology). The signal was detected using enhanced chemiluminescence (ECL, Thermo Fisher Scientific). Quantitative analysis of the bands was performed with a computer-assisted imaging densitometer (UN-SCAN-IT gel version 5.1).
Host cell adhesion assay
The assays were performed essentially as described before38. Cancer cells were co-incubated with P. aeruginosa 8822 in 96-well plates in the presence or absence of various concentrations of aldolase A for 2 h. Monolayers of cancer cells were extensively washed with PBS to remove unbound P. aeruginosa. Monolayers were homogenized in 0.1% saponin (SigmaMillipore) in PBS. P. aeruginosa were enumerated by serial dilution of the homogenized suspensions and subsequent determination of colony-forming units (CFU/ml) by plating on LB agar plates. Control values of CFU/ml expressed as 100%. siRNA-induced silencing of aldolase and MUC1 genes in MDA-MB-231 and Mel2 cells as conducted as described previously80,81. Briefly, SMARTpool human ALDOA, MUC1, and non-targeting siRNA pool (non-targeting siRNAs) (Dharmacon, PA) were used as siRNA targeting aldolase A, MUC1, and negative control, respectively. MDA-MB-231 and Mel2 cells were seeded in 96-well plates, cultured to 80–90% confluence, transfected with 120 nM siRNAs using FuGENE HD (Promega) according to the manufacturer’s protocol for 48 h, and used for host cell adhesion assays. Knockdown of ALDOA and MUC1 by siRNA transfection was examined by western blot analyses using monoclonal mouse anti-aldolase A and MUC1 antibodies (1:200, Santa Cruz Biotechnology).
Detection of azurin gene in human tumors
Tumor samples of breast cancer and melanoma were collected at University of Illinois at Chicago. All patients included in the analysis were diagnosed with either breast cancer or melanoma. Supplementary Fig. 10 contains relevant patient information with similar age range and median age. The detection of azu gene in the human tumors was performed by PCR with azu-specific primers; 5′-CAGTTCACCGTCAACCTGTCC-3′ and 5′-TGGTGTGGGCGATGACACG-3′. Human GAPDH was also amplified as a loading control; 5′-AACGGGAAG CTTGTCATCAA-3′ and 5′-TGGACTCCACGACGTACTCA-3′. We used ultraclean instruments, kits, and reagents to minimize and control for contamination. DNeasy Blood & Tissue Kits (Qiagen) were used to prepare template DNA. The amplified PCR products were confirmed as a single band on 2% agarose gels. Products were purified by using QIAquick PCR Purification kits (Qiagen) and sequenced with an ABI Prism 3700 DNA analyzer. The Fisher’s Exact Test was applied to determine if the positive rates are significantly different between the subjects with the primary and the metastatic tumors.
P. aeruginosa and azurin visualization by TEM
Frozen tumors were processed for immunogold transmission electron microscopy in accordance with a published method82,83 with minor modifications. Tumor samples along with their survival data were obtained from patients who had signed informed consent (IRB #H-96-772). The frozen samples were diced into 1 × 1 mm cubes on dry ice then fixed by incubating them in PBS-containing 4% paraformaldehyde and 0.75% glutaraldehyde for 72 h at 4 °C. The samples were thoroughly washed with PBS then were incubated with PBS-containing 0.1% saponin for 1 h at room temperature on a rotary mixer (Ted Pella, Inc, Redding, CA). After the samples were thoroughly washed with PBS, they were incubated with PBS-containing 5% bovine serum albumin (BSA), 0.1% cold water fish skin gelatin (CWFS), and 5% normal goat serum (NGS) (Goat Gold Conjugate Blocking Solution, Electron Microscopy Sciences, Hatfield, PA) for 1 h at 4 °C. The samples were thoroughly washed with PBS-containing 0.1% acetylated bovine serum albumin (BSA-c) (Aurion BSA-c, Electron Microscopy Sciences) then incubated with either mouse anti-Pseudomonas aeruginosa antibody (clone B11, 1:50 dilution, Thermo Fisher Scientific, Waltham, MA) or rabbit anti-azurin antibody (1:100 dilution)1 for 48 h at 4 °C. After extensive washing, the samples were incubated with 0.1% BSA-c/PBS-containing either goat anti-mouse or goat anti-rabbit secondary antibody for 24 h at 4 °C. To visualize each specific primary antibody, these antibodies were labeled with 10 nm gold particles (1:50 dilution, Electron Microscopy Sciences). The samples were washed with PBS-0.1% BSA-c and deionized water then incubated with deionized water containing 2.5% glutaraldehyde (Electron Microscopy Sciences) for 1 h at room temperature. After extensive washing with deionized water, the sections were fixed with deionized water containing 0.5% osmium tetroxide and 1.5% potassium ferricyanide (Electron Microscopy Sciences) for 15 min in the dark. Next, the samples were dehydrated by incubation in an ascending series of ethanol (25, 50, 75, 95, 100%, Electron Microscopy Sciences) followed by incubation in a 1 to 1 ratio of 100% ethanol to epoxy resin (comprised of a mixture of EMbed 812, nadic methyl anhydride, dodecenyl succinic anhydride, and 2,4,6-Tris(dimethylaminomethyl)phenol, Electron Microscopy Sciences) for 12 h at room temperature on a rotary mixture (Ted Pella, Inc). The samples were incubated with 100% epoxy resin for 12 h at room temperature on a rotary mixer (Ted Pella, Inc). The epoxy resin was changed and the samples were incubated for 2 h at room temperature on a rotary mixer. The samples were placed into flat embedding molds filled with epoxy resin then allowed to polymerize at 70 °C for 24 h. Ultrathin sections (70 nm) were cut with an ultramicrotome (EM UC7, Leica Microsystems, Buffalo Grove, IL), mounted on formvar- and carbon-coated 200 mesh copper grids (Electron Microscopy Sciences) then stained with filtered 1% uranyl acetate prior to imaging. Samples were imaged with a Philips CM 120 transmission electron microscope (TSS Microscopy, Hillsboro, OR) equipped with a BioSprint sixteen-megapixel digital camera (Advanced Microscopy Techniques, Woburn, MA).
Patients’ survival
Survival of each patient of primary melanoma and breast cancer was measured from the date of diagnosis until death from any cause or last follow-up. The experiments were based on randomized trials and the investigators were blinded to the assignments in the experiments and outcome evaluations. Two independent sample tests were performed for comparing the survival days of azu gene positive and negative groups. The Shapiro–Wilk’s test showed that normality was severely violated in data of melanoma patients. The Wilcoxon’s rank sum test for the medians in the melanoma survival data was chosen due to its robustness to the deviation from normality. For breast cancer patients, Shapiro–Wilk’s test showed that the survival days in both azu gene positive and negative groups can be well-modeled by a normal distribution. As equal variances between the two groups were reasonable and was confirmed by a F test, a pooled t test on means was performed to compare the survival days between azu gene positive and negative groups. Data were analyzed using R (v.4.0.2) and RStudio (v.1.3.1093).
Effect of azurin on transgenic mice
Hemizygous MMTV-PyMT (mouse mammary tumor virus-polyoma middle tumor-antigen) female mice were obtained from The Jackson Laboratory. Since MMTV-PyMT mice develop spontaneous mammary tumors that closely resemble the progression and morphology of human breast cancers, the mouse model is widely used and well characterized84. Mice at 4-weeks old were randomized into control (N = 7) and azurin-treatment (N = 5) groups. Control animals received PBS and azurin-treatment animals received 2.5 mg/kg of azurin in sterile PBS i.p. 3× week for nearly two months (80 days old). Based on azurin levels in CF patients’ sera, the highest level of azurin was 32 µg/ml. In order to reach this level of azurin in mouse (25 g-body weight, 2 ml total blood volume), azurin does at 2.5 mg/kg was determined. Tumor volume and body weights were determined three times weekly. All tumors were excised and weighted at the end of the study. All experiments were approved by University of Illinois at Chicago (UIC) Institutional Animal Care and Use Committee (IACUC) and conformed to the guidelines set by United States Animal Welfare Act and the National Institutes of Health.
Statistics and reproducibility
The paired student’s t test the one-way analysis of variance were used for comparisons. P value < 0.05 was considered significant. Number of replicates (N) can be found in the figure legends. Data were analyzed using Graphpad Prism software (ver. 8), R (ver.4.0.2), and RStudio (ver.1.3.1093).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data that support the findings of this study are available in the supplementary information. Also, the datasets presented in the main figures are deposited in the repository Figshare; https://doi.org/10.6084/m9.figshare.21644321, https://doi.org/10.6084/m9.figshare.21644333, https://doi.org/10.6084/m9.figshare.21644336, https://doi.org/10.6084/m9.figshare.21644342, https://doi.org/10.6084/m9.figshare.21644345, https://doi.org/10.6084/m9.figshare.21644348, and https://doi.org/10.6084/m9.figshare.21644351. No computer code is involved in this study. Uncropped images for blots are available in the supplementary information.
References
Yamada, T. et al. Bacterial redox protein azurin, tumor suppressor protein p53, and regression of cancer. Proc. Natl Acad. Sci. USA 99, 14098–14103 (2002).
Yamada, T. et al. The bacterial redox protein azurin induces apoptosis in J774 macrophages through complex formation and stabilization of the tumor suppressor protein p53. Infect. Immun. 70, 7054–7062 (2002).
Punj, V. et al. Bacterial cupredoxin azurin as an inducer of apoptosis and regression in human breast cancer. Oncogene 23, 2367–2378 (2004).
Pseudomonas Gives Cancer Cells the Blues. Science’s STKE 2002, tw416-tw416, https://doi.org/10.1126/stke.2002.158.tw416 (2002).
Coley, W. B. The treatment of malignant tumors by repeated inoculations of erysipelas. With a report of ten original cases. 1893. Clin. Orthop. Relat. Res. 3–11 (1991).
Coley, W. B. II Contribution to the Knowledge of Sarcoma. Ann. Surg. 14, 199–220 (1891).
McCarthy, E. F. The toxins of William B. Coley and the treatment of bone and soft-tissue sarcomas. IOWA Orthop. J. 26, 154–158 (2006).
Dolgin, E. Fighting cancer with microbes. Nature 577, S16–S18 (2020).
Guglielmi, G. How gut microbes are joining the fight against cancer. Nature 557, 482–484 (2018).
Zhou, S., Gravekamp, C., Bermudes, D. & Liu, K. Tumour-targeting bacteria engineered to fight cancer. Nat. Rev. Cancer 18, 727–743 (2018).
Roy, S. & Trinchieri, G. Microbiota: a key orchestrator of cancer therapy. Nat. Rev. Cancer 17, 271–285 (2017).
Xavier, J. B. et al. The cancer microbiome: distinguishing direct and indirect effects requires a systemic view. Trends Cancer 6, 192–204 (2020).
Urbaniak, C. et al. The microbiota of breast tissue and its association with breast cancer. Appl. Environ. Microbiol. 82, 5039–5048 (2016).
Yamada, T. et al. Apoptosis or growth arrest: modulation of tumor suppressor p53′s specificity by bacterial redox protein azurin. Proc. Natl Acad. Sci. USA 101, 4770–4775 (2004).
Kwan, J. M. et al. Bacterial proteins as potential drugs in the treatment of leukemia. Leuk. Res. 33, 1392–1399 (2009).
Smith, W. D. et al. Current and future therapies for Pseudomonas aeruginosa infection in patients with cystic fibrosis. FEMS Microbiol. Lett. 364, https://doi.org/10.1093/femsle/fnx121 (2017).
Abraham, E. H. et al. Cystic fibrosis hetero- and homozygosity is associated with inhibition of breast cancer growth. Nat. Med. 2, 593–596 (1996).
Maisonneuve, P., Marshall, B. C., Knapp, E. A. & Lowenfels, A. B. Cancer risk in cystic fibrosis: a 20-year nationwide study from the United States. J. Natl Cancer Inst. 105, 122–129 (2013).
Zaborina, O. et al. Secreted products of a nonmucoid Pseudomonas aeruginosa strain induce two modes of macrophage killing: external-ATP-dependent, P2Z-receptor-mediated necrosis and ATP-independent, caspase-mediated apoptosis. Microbiology 146, 2521–2530 (2000).
Hiraoka, Y., Yamada, T., Goto, M., Das Gupta, T. K. & Chakrabarty, A. M. Modulation of mammalian cell growth and death by prokaryotic and eukaryotic cytochrome c. Proc. Natl Acad. Sci. USA 101, 6427–6432 (2004).
Ryden, L. & Lundgren, J. Homology relationships among the small blue proteins. Nature 261, 344–346 (1976).
Nar, H., Messerschmidt, A., Huber, R., van de Kamp, M. & Canters, G. W. Crystal structure analysis of oxidized Pseudomonas aeruginosa azurin at pH 5.5 and pH 9.0. A pH-induced conformational transition involves a peptide bond flip. J. Mol. Biol. 221, 765–772 (1991).
Marshall, N. M. et al. Rationally tuning the reduction potential of a single cupredoxin beyond the natural range. Nature 462, 113–116 (2009).
Yamada, T. et al. Internalization of bacterial redox protein azurin in mammalian cells: entry domain and specificity. Cell. Microbiol. 7, 1418–1431 (2005).
Harimoto, T. et al. Rapid screening of engineered microbial therapies in a 3D multicellular model. Proc. Natl Acad. Sci. 116, 9002–9007 (2019).
Huang, F. et al. Anticancer actions of azurin and its derived peptide p28. Protein J. 39, 182–189 (2020).
Bizzarri, A. R., Di Agostino, S., Andolfi, L. & Cannistraro, S. A combined atomic force microscopy imaging and docking study to investigate the complex between p53 DNA binding domain and Azurin. J. Mol. Recognit. 22, 506–515 (2009).
Gao, M., Zhou, J., Su, Z. & Huang, Y. Bacterial cupredoxin azurin hijacks cellular signaling networks: protein-protein interactions and cancer therapy. Protein Sci. 26, 2334–2341 (2017).
Yamada, T., Das Gupta, T. K. & Beattie, C. W. p28-mediated activation of p53 in G2-M phase of the cell cycle enhances the efficacy of DNA damaging and antimitotic chemotherapy. Cancer Res. 76, 2354–2365 (2016).
Razzak, M. Targeted therapies: one step closer to drugging p53. Nat. Rev. Clin. Oncol. 10, 246 (2013).
Warso, M. A. et al. A first-in-class, first-in-human, phase I trial of p28, a non-HDM2-mediated peptide inhibitor of p53 ubiquitination in patients with advanced solid tumours. Br. J. Cancer 108, 1061–1070 (2013).
Lulla, R. R. et al. Phase I trial of p28 (NSC745104), a non-HDM2-mediated peptide inhibitor of p53 ubiquitination in pediatric patients with recurrent or progressive central nervous system tumors: a pediatric Brain Tumor Consortium Study. Neuro-Oncol. 18, 1319–1325 (2016).
Mehta, N. et al. Bacterial carriers for glioblastoma therapy. Mol. Ther. Oncolytics 4, 1–17 (2017).
Zhang, Y. et al. Escherichia coli Nissle 1917 targets and restrains mouse B16 melanoma and 4T1 breast tumors through expression of azurin protein. Appl. Environ. Microbiol. 78, 7603–7610 (2012).
Bizzarri, A. R. et al. Interaction of an anticancer peptide fragment of azurin with p53 and its isolated domains studied by atomic force spectroscopy. Int. J. Nanomed. 6, 3011–3019 (2011).
Apiyo, D. & Wittung-Stafshede, P. Unique complex between bacterial azurin and tumor-suppressor protein p53. Biochem. Biophys. Res. Commun. 332, 965–968 (2005).
Vijgenboom, E., Busch, J. E. & Canters, G. W. In vivo studies disprove an obligatory role of azurin in denitrification in Pseudomonas aeruginosa and show that azu expression is under control of rpoS and ANR. Microbiol.143, 2853–2863 (1997).
Tunio, S. A. et al. The moonlighting protein fructose-1, 6-bisphosphate aldolase of Neisseria meningitidis: surface localization and role in host cell adhesion. Mol. Microbiol. 76, 605–615 (2010).
Welin, J., Wilkins, J. C., Beighton, D. & Svensater, G. Protein expression by Streptococcus mutans during initial stage of biofilm formation. Appl. Environ. Microbiol. 70, 3736–3741 (2004).
Jeffery Marano, R. et al. Secreted biofilm factors adversely affect cellular wound healing responses in vitro. Sci. Rep. 5, 13296 (2015).
Jewett, T. J. & Sibley, L. D. Aldolase forms a bridge between cell surface adhesins and the actin cytoskeleton in apicomplexan parasites. Mol. Cell 11, 885–894 (2003).
Diaz, S. A. et al. The binding of plasmodium falciparum adhesins and erythrocyte invasion proteins to aldolase is enhanced by phosphorylation. PloS One 11, e0161850 (2016).
Oldfield, N. J. et al. T-cell stimulating protein A (TspA) of Neisseria meningitidis is required for optimal adhesion to human cells. Cell. Microbiol. 9, 463–478 (2007).
Lillehoj, E. P. et al. Muc1 mucins on the cell surface are adhesion sites for Pseudomonas aeruginosa. Am. J. Physiol. Lung Cell. Mol. Physiol. 280, L181–L187 (2001).
Verceles, A. C. et al. MUC1 ectodomain is a flagellin-targeting decoy receptor and biomarker operative during Pseudomonas aeruginosa lung infection. Sci. Rep. 11, 22725 (2021).
Lu, W. et al. Cutting edge: enhanced pulmonary clearance of Pseudomonas aeruginosa by Muc1 knockout mice. J. Immunol. 176, 3890–3894 (2006).
Alrahman, M. A. & Yoon, S. S. Identification of essential genes of Pseudomonas aeruginosa for its growth in airway mucus. J. Microbiol. 55, 68–74 (2017).
Sajjan, U. et al. Binding of nonmucoid Pseudomonas aeruginosa to normal human intestinal mucin and respiratory mucin from patients with cystic fibrosis. J. Clin. Invest. 89, 657–665 (1992).
Stover, C. K. et al. Complete genome sequence of Pseudomonas aeruginosa PAO1, an opportunistic pathogen. Nature 406, 959–964 (2000).
Bodelon, G. et al. Detection and imaging of quorum sensing in Pseudomonas aeruginosa biofilm communities by surface-enhanced resonance Raman scattering. Nat. Mater. 15, 1203–1211 (2016).
Gokalsin, B., Aksoydan, B., Erman, B. & Sesal, N. C. Reducing virulence and biofilm of pseudomonas aeruginosa by potential quorum sensing inhibitor carotenoid: Zeaxanthin. Microb. Ecol. https://doi.org/10.1007/s00248-017-0949-3 (2017).
Whiteley, M., Diggle, S. P. & Greenberg, E. P. Progress in and promise of bacterial quorum sensing research. Nature 551, 313–320 (2017).
Rahme, L. G. et al. Plants and animals share functionally common bacterial virulence factors. Proc. Natl Acad. Sci. 97, 8815–8821 (2000).
Kwak, G.-Y. et al. Quorum sensing-independent cellulase-sensitive pellicle formation is critical for colonization of burkholderia glumae in rice plants. Front. Microbiol. 10, https://doi.org/10.3389/fmicb.2019.03090 (2020).
Hieken, T. J. et al. The microbiome of aseptically collected human breast tissue in benign and malignant disease. Sci. Rep. 6, 30751 (2016).
Leal, A. S. et al. Retinoid X receptor agonist LG100268 modulates the immune microenvironment in preclinical breast cancer models. npj Breast Cancer 5, 39 (2019).
Christenson, J. L. et al. MMTV-PyMT and derived Met-1 mouse mammary tumor cells as models for studying the role of the androgen receptor in triple-negative breast cancer progression. Horm. Cancer 8, 69–77 (2017).
de Martel, C. et al. Global burden of cancers attributable to infections in 2008: a review and synthetic analysis. Lancet Oncol. 13, 607–615 (2012).
Amieva, M. & Peek, R. M. Jr. Pathobiology of Helicobacter pylori-induced gastric cancer. Gastroenterology 150, 64–78 (2016).
Blaser, M. J. & Atherton, J. C. Helicobacter pylori persistence: biology and disease. J. Clin. Invest. 113, 321–333 (2004).
Passera, A. et al. Not just a pathogen? Description of a plant-beneficial pseudomonas syringae Strain. Front. Microbiol. 10, https://doi.org/10.3389/fmicb.2019.01409 (2019).
Singh, A., Jain, A., Sarma, B. K., Upadhyay, R. S. & Singh, H. B. Rhizosphere competent microbial consortium mediates rapid changes in phenolic profiles in chickpea during Sclerotium rolfsii infection. Micobiol. Res. 169, 353–360 (2014).
Sachs, J. L., Skophammer, R. G., Bansal, N. & Stajich, J. E. Evolutionary origins and diversification of proteobacterial mutualists. Proc. Biol. Sci. 281, 20132146–20132146 (2013).
Sachs, J. L., Skophammer, R. G. & Regus, J. U. Evolutionary transitions in bacterial symbiosis. Proc. Natl Acad. Sci. 108, 10800–10807 (2011).
Mehta, R. R. et al. A 28-amino-acid peptide fragment of the cupredoxin azurin prevents carcinogen-induced mouse mammary lesions. Cancer Prev. Res. 3, 1351–1360 (2010).
Bergmans, D. & Bonten, M. (ed. Jean-Louis Vincent) 131–140 (Springer,Berlin Heidelberg).
Iglewski, B. H. Pseudomonas. Medical Microbiology. 4th edition. Chapter 27 (1996).
Sieow, B. F., Wun, K. S., Yong, W. P., Hwang, I. Y. & Chang, M. W. Tweak to treat: reprograming bacteria for cancer treatment. Trends Cancer 7, 447–464 (2021).
Pang, Z., Gu, M. D. & Tang, T. Pseudomonas aeruginosa in cancer therapy: current knowledge, challenges and future perspectives. Front. Oncol. 12, 891187 (2022).
Chiba, A. et al. Neoadjuvant chemotherapy shifts breast tumor microbiota populations to regulate drug responsiveness and the development of metastasis. Mol. Cancer Res. https://doi.org/10.1158/1541-7786.mcr-19-0451 (2019).
Cho, J. H. et al. The bacterial protein azurin enhances sensitivity of oral squamous carcinoma cells to anticancer drugs. Yonsei Med. J. 52, 773–778 (2011).
Darzins, A. & Chakrabarty, A. M. Cloning of genes controlling alginate biosynthesis from a mucoid cystic fibrosis isolate of Pseudomonas aeruginosa. J. Bacteriol. 159, 9–18 (1984).
Mehta, R. R. et al. In vitro transformation of human congenital naevus to malignant melanoma. Melanoma Res. 12, 27–33 (2002).
Rauth, S., Kichina, J., Green, A., Bratescu, L. & Das Gupta, T. K. Establishment of a human melanoma cell line lacking p53 expression and spontaneously metastasizing in nude mice. Anticancer Res. 14, 2457–2463 (1994).
Goto, M. et al. Induction of apoptosis in macrophages by Pseudomonas aeruginosa azurin: tumour-suppressor protein p53 and reactive oxygen species, but not redox activity, as critical elements in cytotoxicity. Mol. Microbiol. 47, 549–559 (2003).
Shen, F. S. & Loh, Y. P. Intracellular misrouting and abnormal secretion of adrenocorticotropin and growth hormone in cpefat mice associated with a carboxypeptidase E mutation. Proc. Natl Acad. Sci. USA 94, 5314–5319 (1997).
Yang, Y. et al. Conformational determinants necessary for secretion of Paecilomyces thermophila β-1,4-xylosidase that lacks a signal peptide. AMB Express 8, 11 (2018).
Chevallet, M., Diemer, H., Van Dorssealer, A., Villiers, C. & Rabilloud, T. Toward a better analysis of secreted proteins: the example of the myeloid cells secretome. Proteomics 7, 1757–1770 (2007).
Lopes, M. A., Oliveira Franco, F., Henriques, F., Peres, S. B. & Batista, M. L. Jr. LLC tumor cells-derivated factors reduces adipogenesis in co-culture system. Heliyon 4, e00708 (2018).
Bozkaya, G. et al. Cooperative interaction of MUC1 with the HGF/c-Met pathway during hepatocarcinogenesis. Mol. Cancer 11, 64 (2012).
Grandjean, G. et al. Definition of a novel feed-forward mechanism for glycolysis-HIF1α signaling in hypoxic tumors highlights aldolase A as a therapeutic target. Cancer Res. 76, 4259–4269 (2016).
Figge, D. A., Rahman, I., Dougherty, P. J. & Rademacher, D. J. Retrieval of contextual memories increases activity-regulated cytoskeleton-associated protein in the amygdala and hippocampus. Brain Struct. Funct. 218, 1177–1196 (2013).
Gavini, C. K., Cook, T. M., Rademacher, D. J. & Mansuy-Aubert, V. Hypothalamic C2-domain protein involved in MC4R trafficking and control of energy balance. Metab. Clin. Exp. 102, 153990 (2020).
Rossdeutscher, L. et al. Chemoprevention activity of 25-hydroxyvitamin D in the MMTV-PyMT mouse model of breast cancer. Cancer Prev. Res. 8, 120–128 (2015).
Acknowledgements
The mass spectrometry data was obtained and analyzed by Protein lab at Research Resources Center, UIC. This work made use of a facility in the Biological Resources Laboratory, UIC. P. aeruginosa azurin null mutant was kindly gifted from Drs. Erik Vijgenboom and Gerard W. Canters (Leiden Institute of Physics, Huygens-Kamerlingh Onnes Laboratory, Netherlands). We thank Drs. R.R. Mehta, R.G. Mehta, and C.W. Beattie for their suggestions in the early phase of the project. Parts of this work were Dr. J. Choi’s Ph.D. thesis in Anatomy & Cell Biology, University of Illinois College of Medicine, and Dr. T.K. Das Gupta was her thesis advisor. Human CF patient samples were provided by the CF Biospecimen Registry at the Children’s Healthcare of Atlanta and Emory University CF Discovery Core. This research was supported in part by the NIH/National Institute of Biomedical Imaging and Bioengineering (R01EB023924) and NIH/National Cancer Institute (R21CA252370) to TY. Dr. Ananda M. Chakrabarty participated in intellectually developing this article before he passed away on 10 July 2020.
Author information
Authors and Affiliations
Contributions
J.K.C., S.A.N., M.G., J.W., K.C., A.M.C., T.K.D., and T.Y. conceived, conceptualized, and designed the study. A.A.S. provided clinical expertise and samples for azurin analysis in sera obtained from CF patients. J.K.C., S.A.N., M.G., K.C., A.G., and T.Y. performed biochemical experiments. D.R. collected and analyzed electron microscopy data. J.W. performed statistical analyses.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Tomasz Karpiński and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Eve Rogers.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Choi, J.K., Naffouje, S.A., Goto, M. et al. Cross-talk between cancer and Pseudomonas aeruginosa mediates tumor suppression. Commun Biol 6, 16 (2023). https://doi.org/10.1038/s42003-022-04395-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-022-04395-5
This article is cited by
-
Pseudomonas against cancer
Nature Reviews Microbiology (2023)
-
Environmental insults and compensative responses: when microbiome meets cancer
Discover Oncology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.