Abstract
Breeding new wheat varieties with salt resistance is one of the best ways to solve a constraint on the sustainability and expansion of wheat cultivation. Therefore, understanding the molecular components or genes related to salt tolerance must contribute to the cultivation of salt-tolerant varieties. The present study used a recombinant inbred line (RIL) population to genetically dissect the effects of different salt stress concentrations on wheat seed germination and seedling traits using two quantitative trait locus (QTL) mapping methods. A total of 31 unconditional and 11 conditional QTLs for salt tolerance were identified on 11 chromosomes explaining phenotypic variation (PVE) ranging from 2.01 to 65.76%. Of these, 15 major QTLs were found accounting for more than 10% PVE. QTL clusters were detected on chromosomes 2A and 3B in the marker intervals ‘wPt-8328 and wPt-2087’ and ‘wPt-666008 and wPt-3620’, respectively, involving more than one salt tolerance trait. QRdw3B and QSfw3B.2 were most consistent in two or more salt stress treatments. 16 candidate genes associated with salt tolerance were predicted in wheat. These results could be useful to improve salt tolerance by marker-assisted selection (MAS) and shed new light on understanding the genetic basis of salt tolerance in wheat.
Similar content being viewed by others
Introduction
The degree of land salinization has a significant upward trend in the world, causing serious harm to the quality and yield of crops1. To date, the arable land affected by salinity worldwide is greater than 800 million ha, which is approximately six percent of the global land area2,3. Moreover, the proportion of salt-affected soil seems to be increased because of irrigation every year, which could result in salt accumulation in agricultural soils4,5. Therefore, salinization has become a major limitation of grain yield2. Wheat (Triticum aestivum L.) is one of the most important food crops in the world, and it is a part of the daily diet of over 70% of the world population6. To meet the food needs of the growing world population, a significant increase in wheat yield is required7,8. However, most wheat cultivars hardly have salt tolerance in production because salinity stress influences seedling establishment at early growth stages of common wheat and severely reduces yield9. Therefore, breeding salt-tolerant wheat cultivars is a practical method to reduce the harm of salinization for food security10.
From genetic and physiological points of view, salt tolerance is complex. Tolerance often shows the characteristics of a multigenic trait controlled by polygenes11. Quantitative trait locus (QTL) mapping provides an effective approach to dissect complicated quantitative traits into component loci to study their relative effects on a specific trait12,13, thereby providing breeders with targets for marker-assisted selection (MAS)14. Previous researchers have identified some QTLs for salt tolerance traits in some crops, such as rice (Oryza sativa)15 and maize (Zea mays)16. Using common wheat, some results were also reported by QTL mapping17,18,19,20,21 and genome-wide association studies (GWAS) under salt stress conditions21,22,23.
Salinity stress negatively affects plant growth, development and overall productivity by inducing ion toxicity, osmotic stress, hormonal disturbance and oxidative stress24,25. At the initial stage of salt stress, the water absorption capacity of the root system is reduced, and water loss from the leaves is accelerated due to osmotic stress of high salt accumulation in soil and plants26. Over time, soil salinity causes toxic concentrations of Na+ to accumulate in leaves. This imposes an additional limitation to growth by reducing the longevity of photosynthetic tissues27.
There were two pathways that play a key role in salt tolerance in wheat, namely, HKT genes that mediate Na+ exclusion and the SRO gene that regulates ROS homeostasis2. TmHKT1;4-A2 decreased the leaf blade Na+ concentration by 50%, TmHKT1;5-A decreased it by 30%, and both genes together decreased it by 60%28. TaSRO can act as a transcription factor regulator or covalent binding factor to interact with different transcription factors, thereby extensively participating in plant responses to stress conditions29. In addition, many other genes have been investigated. For example: TaAQP8, NHX gene family, TaCYP81D5, TaCIPK29, etc30,31,32,33,34.
Generally, previous studies evaluated characteristics under salt stress in wheat, mainly including the germination rate (GR), germination potential (GP), mean root length (MRL), root fresh weight (RFW), root dry weight (RDW), shoot height (SH), shoot fresh weight (SFW) and shoot dry weight (SDW), at the seedling stage17,18,19,22,35. Few studies have been reported at the adult stage involving plant height, spike length, grain number, yield per plant, spike number, and thousand-kernel weight20,23. Because QTL mapping based on linkage and marker trait association can be effectively used for gene pyramiding, germplasm screening of diversified material for abiotic (salinity, cold, salt, drought) and biotic stresses (disease, pest), etc., the identification and location of specific genes mediating quantitative characters is of great importance in plant breeding54. The objectives of QTL mapping are to offer a direct means to investigate the number of genes influencing the trait, to determine the location of the gene that affects traits of interest, to know the effect of genes on variation of the trait and to carry out a study on linkage between genes of interest54. To date, more than 500 QTLs (excluding those involved in digenic epistatic interactions and QTL x treatment interactions) have been identified on all 21 wheat chromosomes, explaining 8.4% to 40.0% of the phenotypic variation for each QTL36. Only a dozen significant QTLs, mostly distributed on chromosomes 2D, 3B, 4B, 4D, 5A, and 7A25, were reported. For example, a locus Kna 1 for Na+ exclusion was mapped on chromosome 4DL37, and the QTL QNax.aww-7AS explaining 40% of Na+ variation was identified using two mapping populations38. For shoot dry weight, there was one major QTL, qSNAX.7A.3, with approximately 19% phenotypic variation19. Most 31 QTLs for salinity tolerance were detected on chromosomes 3B and 5B, while two QTLs for fresh weight and height of shoots were detected on chromosomes 1A and 3A, which explained 18% and 12.9% of the phenotypic variation, respectively17. There were some QTLs for the ionic traits across the three growth stages on 1BS, 2AL, 2BS and 3AL detected22. Two QTLs for seedling height and five QTLs for taproot length were detected using 168 doubled haploid lines39. 69 QTLs associated with seven seedling traits were found on 20 chromosomes, except for chromosome 1A40.
Although many QTLs for wheat salt tolerance traits have been identified, some of them are not well used in wheat breeding under salt stress, and they were mapped using the unconditional QTL mapping method, which could not dissect the genetic difference under salt stress with different salt concentrations. To dissect the interactions and molecular differences under different treatments or between related traits, the method of conditional QTL mapping was developed by Zhu41, which can be used to exclude the contribution of a treatment from the variation of the resultant treatment. Conditional variation or net variation is defined as the remaining variation of the resultant treatment, suggesting that the extra effects of these genes/QTLs are independent of the causal treatment. Therefore, the genetic effects of conditional variations of the resultant treatment can be dissected at the QTL/gene level. By comparing unconditional and conditional QTLs, the genetic interdependencies between them can be identified at the individual QTL level. This comparison might provide valuable information for marker-assisted selection to improve salt tolerance without negative effects on wheat.
To date, conditional QTL mapping has rarely been used in wheat salt stress for traits related to seed germination and seedling stage. Therefore, a recombinant inbred line (RIL) population derived from the cross between Nuomai 1 (NM 1) and Gaocheng 8901 (GC 8901) was used to genetically dissect the effects of different salt stress concentrations on wheat seed germination and seedling traits using unconditional and conditional quantitative trait locus (QTL) mapping. The objectives of this study were to (1) identify novel stable QTLs under various salt concentrations using unconditional and conditional QTL mapping and (2) find new candidate genes for significant QTLs that were predicted. These results will provide important information on breeding wheat cultivars with salt tolerance.
Results
Phenotypic variation and correlation analysis
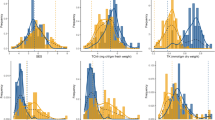
All of the evaluated traits showed approximately continuous variation in each of the treatments (Tables 1 and 2). The salt-tolerant parent NM1 showed significantly higher values (14.46–50% greater) than the salt-sensitive parent GC8901 for GR in the T1 and T2 treatments. There were no significant differences in GR in the two parents under the N treatment. However, for GP, significant differences were not found in the two parents under the N, T1 and T2 treatments (Table 1). With increasing salt concentration, the means of GR and GP decreased. In Table 2, significant differences were found between the two parents for seedling traits under different treatments, such as MRL under the N, T1 and T2 treatments, RFW under the T3 treatment, and SDW under the T1, T2 and T3 treatments. In the RIL population, transgressive segregation was observed on both the high and low sides for GR, MRL, RFW, RDW, SH, SFW and SDW, which indicated that the alleles with positive effects were contributed by both parents (Tables 1 and 2).
Significant positive correlations were observed between SH and MRL and SFW and RFW under all treatments, but they were negatively correlated with RDW (Table S1). The highest positive correlation coefficient was observed with 0.8 between SH and MRL, but the lowest negative correlation coefficient was − 0.59 between SH and RDW under the T2 treatment. MRL was negatively correlated with RDW under all treatments but positively correlated with SDW, SFW, RFW and GR under most treatments. SFW was significantly and negatively correlated with RDW but positively correlated with SH, MRL, SDW, SFW, RDW, RFW and GR. RDW was significantly and positively correlated with SDW under the N, T2 and T3 treatments but negatively correlated with all other traits except GP under most of the treatments. In most of the different salt stress treatments, GP was significantly and positively correlated with GR, and GR was significantly and positively correlated with SH, MRL, SFW and GP.
QTL analysis
A total of 31 unconditional QTLs and 11 conditional QTLs for 8 traits were detected on nine chromosomes (Tables 3, 4 and 5). Of 31 unconditional QTLs, 11 unconditional QTLs for GR and GP were identified, explaining from 2.1 to 35.34% of phenotypic variation (PVE) (Table 3), and twenty unconditional QTLs were found for SH, MRL, SDW, SFW, RDW and RFW, accounting for 4.58–65.76% of the phenotypic variation (Table 4). Fourteen major unconditional QTLs were identified. The conditional QTLs accounted for 1.07–20.09% of the phenotypic variation under the different treatments (Table 5).
Four unconditional QTLs were detected for GP in three different concentrations of salt stress (Table 3), which were distributed on chromosomes 2B, 3B, and 6A. The additive effect of QGp3B came from GC 8901 with a maximum PVE of 35.34%, which was co-located with the unconditional QTLs QRdw3B, QSfw3B.1 and QSdw3B (Table 4, Fig. 1).
Seven unconditional QTLs were detected for GR under salt stress at three different concentrations (Table 3), accounting for from 2.1 to 6.4% of PVE, which were distributed on seven chromosomes (2A, 3A, 3B, 3D, 5A, 6A, 6B). The additive effect of two QTLs, QGr6A and QGr3D, came from GC 8901, and the other came from NM1. However, they all were minor QTLs. The QTL QGr3B.1 was colocalized with the other QTLs, QMrl3B.1, QRdw3B, QSfw3B.2 and QSdw3B.
Five unconditional QTLs and three conditional QTLs for the MRL were detected, and the PVE of a single QTL ranged from 4.90 to 30.18% (Tables 4 and 5). The synergistic alleles of the QTLs QMrl2A, QMrl2B and QMrl4A were derived from NM 1. The synergistic alleles of the QTLs QMrl3B.1 and QMrl3B.2 were derived from GC 8901. The unconditional QTL QMrl2A was co-located with the unconditional QTL QSh2A in the same marker interval. Three newly identified conditional QTLs were induced by salt stress. Of these, the conditional QTL QMrl2A(T3|T2) was located near the unconditional QTL QMrl2A and the conditional QTL QSh2A(T2/T1) with the same marker wPt-2185, which indicated that a locus on chromosome 2A may play an important role in seedling growth under salt stress.
One unconditional QTL and one conditional QTL were identified for RFW. The unconditional QTL QRfw2A was detected under the T3 treatment, and the PVE was 14.89% (Table 4). The conditional QTL identified for RFW was QRfw2A (T3|T2), which was induced by higher salt stress and played a critical role in salt tolerance.
Six unconditional QTLs and one conditional QTL controlling RDW were detected. The QTL QRdw3B.1 was identified in the same marker interval under four treatments, accounting for 34.7%-65.76% of PVE (Table 4). The synergistic alleles were derived from GC 8901. QRdw3B.2 was found under the T3 treatment. The conditional QTL QRdw2B(T1|N) was identified for RDW, which was induced by salt stress and may play a role under salt stress.
Three unconditional QTLs and one conditional QTL affecting SH were detected, and the PVE of each QTL ranged from 4.84 to 9.27%. The synergistic genes were all from NM 1. There were no QTLs detected under the T2 treatment. The conditional QTL QSh2A(T2|T1) identified for SH was induced by salt stress, which may play a role in salt tolerance.
Four unconditional QTLs and four conditional QTLs for the SFW were identified, ranging from 1.07 to 23.63% of PVE for a single QTL. Most of them were co-located with QRdw3B, except QSfw2A. Of the eight QTLs, the seven loci on chromosome 3B were derived from GC 8901. The QTL QSfw2A was contributed by NM 1-derived alleles. Four conditional QTLs were identified in almost the same marker intervals on chromosome 3B, and the unconditional QTLs for SFW under the N and T2 treatments were also found in similar marker intervals. This indicated that these loci on chromosome 3B seemed to be important for salt tolerance.
One unconditional QTL and one conditional QTL for SDW were observed with 53.67% and 20.09% PVE, respectively. Two QTLs were located on chromosome 3B in marker intervals wPt-5870-wPt-3620 and wPt-666008-wPt-5870 (Fig. 1). The synergistic allele was derived from GC 8901.
For biomass-related traits, two conditional QTLs were identified for different traits. The conditional QTLs QMrl2A(T3|T2), QSh2A (T2|T1) and QRfw2A(T3|T2) were located in the nearby marker intervals and near QMrl2A (Fig. 1). The QTLs QSfw3B(T2|N), QSfw3B(T2|N), QSfw3B(T2|T1), QSfw3B(T3|T2) and QSdw3B(T2|T1), QGr3B.1, QMrl3B.1, QRdw3B, QSfw3B.1 and QSfw3B.2 were located in the nearby marker intervals, and the latter was also identified in unconditional analysis for GR, MRL, RDW and SFW. However, the QTL QRdw2B(T1|N) was only detected by conditional QTL mapping.
Prediction of candidate genes for important loci
Six markers with high PVE values were selected from loci significantly linked with the traits related to salt tolerance for prediction (Table S2). Some candidate genes were identified. The marker wPt-7187 on the chromosome had one candidate gene, TraesCSU02G009300.1, from wheat (Table S2). The functions of this gene are related to calcium ion binding and polysaccharide binding. Meanwhile, from Oryza brachyantha, one candidate gene, OB01G45130, was found to be related to the functions of peroxidase activity, calcium ion binding, and oxidoreductase activity, acting on NAD(P)H, with oxygen as an acceptor. It participates in the oxidation–reduction process. Four candidate genes were found for the marker wPt-2185 from wheat, but their functions are unknown. However, one candidate gene, OB09G21770, from Oryza brachyantha was found to be related to the functions calcium transmembrane transporter activity, phosphorylative mechanism, and calmodulin binding, which take part in the biological process of ion transport and calcium ion transport calcium ion transmembrane transport. This indicated that the four candidate genes perhaps had these functions but need to be further studied. For the marker wPt-2087, three candidate genes, TraesCS2A02G048300, TraesCS2A02G048400, and TraesCS2A02G048500, were found on chromosome 2A, but their functions were unknown in wheat. However, interestingly, each candidate gene was found in Oryza brachyantha, Oryza glumipatula, and Brachypodium distachyon. The functions of the OB06G15330 gene included ion channel activity and voltage-gated potassium channel activity, participating in the processes of ion transport, potassium ion transport, potassium ion transmembrane transport, and ion transmembrane transport. The OGLUM01G14020 gene participated in the response to salt stress. For the marker wPt-5870, three and one candidate genes were found on chromosomes 3B and 2D, respectively, but the functions on chromosome 3B are unknown in wheat. The other eight homologous genes from Oryza glumipatula, Aegilops tauschii, Oryza barthii, Triticum turgidum, Oryza brachyantha, Triticum dicoccoides, Oryza sativa Indica Group, and Oryza glaberrima are all involved in the biological processes of sodium ion transport, transmembrane transport and response to salt stress. This indicated that the candidate genes from common wheat may be related to the response to salt stress, which needs to be further identified. For the marker wPt-3620, three candidate genes were found on chromosome 3B, but the functions are unknown in common wheat. However, in Brachypodium distachyon, the candidate gene BRADI_4g16243v3 was related to the function of zinc ion binding and metal ion binding. For the marker wPt-666008, there were three candidate genes found on chromosomes 3B and 1D, but their functions are unknown. However, the candidate gene ORGLA04G0060900 had the functions of magnesium ion binding and metal ion binding in Oryza glaberrima.
In general, these candidate genes found in common wheat may be related to salt tolerance, and their functions will be explored in future research.
Discussion
Breeding salt-tolerant wheat varieties is an effective and environmentally sustainable way to utilize salt-affected soils and increase global food production21,22. Previous studies showed that most morphological and physiological indexes of wheat were differentially affected by salt stress, of which root dry weight, root fresh weight and the ratio of dry weight root to shoot were greatly affected and were also very sensitive to salt stress55. Similarly, the phenotypic performance of the present study also decreased with increasing salt concentration, but the germination rate and germination potential were greatly affected, which indicated that the salt tolerance of seed germination could also be important for wheat production. Although a number of physiological traits have been reported to correlate with salt tolerance in plants, the abilities of excluding Na+ and maintaining a high cytosolic K+/Na+ ratio seemed to be important for plant salt tolerance. In the present study, seedling biomass traits and germination traits were focused on salt stress. Shoot height was significantly positively correlated with shoot fresh weight under both non-salt stress and salt stress, which was almost consistent with previous studies55,56. In addition, Li found that the germination rate was significantly and positively correlated with shoot height and maximum root length, which was also found in our research. Meanwhile, the germination rate was also significantly and positively correlated with SFW and GP in the present study. These results indicated that germination traits were important for wheat growth under salt stress.
To speed up the breeding of salt-tolerant varieties by marker-assisted selection, knowledge of salt tolerance loci and genes is required. Previous researchers have studied many traits under salt stress with different concentrations at the seedling stage and adult stage using QTL mapping and GWAS17,18,19,20,21,22,36. Of these, some studies involved traits including Na+ exclusion/content, K+ content and K+/Na+ ratio, seedling shoot fresh weight, and plant height, which mainly mapped on chromosomes 2D, 3B, 4B, 4D, 7A and 5A for major QTLs (PVE > 20%)36. However, in the present study, the major QTLs were mainly involved in two chromosomes, 2A and 3B. On chromosome 3B, QTL clusters were found for shoot height, shoot fresh weight and shoot dry weight under salt stress in seedling stages17. Although the QTL loci were all found on chromosome 3B36,43, they were different loci by comparison with previous studies. Moreover, this new locus was involved in many traits, such as GP, MRL, RDW, SFW and SDW, under salt stress in this study, which was consistent with their significant phenotypic correlations. Therefore, it is important for salt tolerance and should be further studied in the future.
In addition, previous reports have identified some important QTLs or genes mainly affecting Na+ content, K+ content, and the K+/Na+ ratio on chromosome 2A. For example, the QTL_2AL.1 region (R2 > 16.93%) was associated with stress tolerance traits across three growth stages, germination, seedling and adult-field-grown plants, including the leaf K+/Na+ ratio, which is proximal to the codominant SSR marker gwm312, which is closely linked to the Nax1 gene22. By QTL mapping of NAX, there were two closely located QTLs, qSNAX.2A.1 and qSNAX.2A.1, and qRNAX.2A.1 coincided with the major NAX locus Nax1 or HKT1:4 in durum wheat and three NAX QTLs found on 2A in bread wheat19,44,45. In the present study, QTL clusters were also found on chromosome 2A in the regions between wPt-8328 and wPt-2087. These QTLs mainly involved salt tolerance traits, including MRL, SH and SFW, in the seedling stage, which was consistent with their significant positive phenotypic correlations. By comparing with the previous consensus map of different marker types, such as simple sequence repeat (SSR), diversity array technology (DArT) markers and SNP46, we found that this QTL cluster for seedling biomass should be different from the QTL_2AL.1 region; that is, this region perhaps involves a new salt tolerance gene, but which needs to be further studied in the future.
Although the conditional QTL mapping method has been used in previous studies42,47, it was not reported on the conditional QTL mapping of salt tolerance traits under different salt-stress conditions in common wheat. In fact, conditional QTL mapping is a good method to dissect QTLs/genes induced or not induced by salt stress. In this study, some QTLs were induced to be identified by salt stress, such as QRfw2A (T3|T2), QMrl2A(T2|N), QSh2A(T2|T1), and QMrl3B(T2|T1). Most interestingly, some of them were induced by low-salt stress conditions, and some were induced by high-stress conditions. Moreover, the QTLs controlling the SFW on chromosome 3B seemed to be unaffected by salt stress because they were all identified under different conditions. Therefore, its region is important for salt tolerance.
By screening the candidate genes for important loci on chromosomes 2A and 3B, there were a total of sixteen candidate genes, including TraesCSU02G009300.1, TraesCSU02G212400, TraesCS2A02G048300, TraesCS3B03G0601400LC.1, TraesCS3B03G0301100LC.1, and TraesCS3B03G0143800LC found in common wheat, but most of their molecular functions and biological processes are unknown, which are different from previous genes related to salt tolerance, such as TaMYB32, TaOPR1, TaSRO1 and TaHKT148,49,50. Of these, the predicted gene, TraesCSU02G009300.1, had the function of calcium ion binding and polysaccharide binding. However, most of the candidate genes from rice, Brachypodium distachyon, and Triticum dicoccoides in this study are related to the function of metal ion binding (calcium, zinc, sodium) and transmembrane transporter activity, mainly participating in sodium ion transport and response to salt stress. This indicates that these candidate genes predicted in common wheat are most likely related to salt tolerance, which should be further studied in the future.
Conclusions
In all, a total of 31 unconditional QTLs and 11 conditional QTLs for 8 salt tolerance traits were detected on 11 chromosomes under different salt stresses. There were 14 major unconditional QTLs and one major conditional QTL identified. On chromosomes 2A and 3B, QTL clusters were found for salt tolerance traits. A total of sixteen candidate genes were predicted. This information is very useful in marker-assisted breeding to enhance salt tolerance in wheat.
Materials and methods
Ethics statement
All samples analysed in our study adhered to all local, national or international guidelines and legislation, and no ethical approval was required.
Plant materials
The RIL population of 256 lines was developed by a single-seed descent method after crossing between Gaocheng 8901 (GC 8901) and Nuomai 1 (NM 1). The parent of NM 1 (Jiangsu Baihuomai/Guandong107) was bred by China Agricultural University and released in 2005 in Beijing. GC 8901 (77546-2/Linzhang) was bred by the Gaocheng Agricultural Science Research Institute and released in 1998 in Hebei Province. These materials were kept and provided by our team.
Experimental design
The experiment was conducted in a hydroponic culture under a greenhouse at Shandong Agricultural University with a 16/8 day/night photoperiod, 27/20 °C day/night temperature and a relative humidity of approximately 60% in 2020. One hundred seeds for each parent and each RIL were selected. Seeds of parents and 256 RILs were sterilized in 3% H2O2 for 20 min, rinsed with deionized water, and then allowed to germinate on two-double filter paper in petri dishes containing distilled water in the dark at 20 ± 2 °C. They were evaluated for salt tolerance under four salt treatments, 0, 50, 100 and 200 mM NaCl, designated the N, T1, T2 and T3 treatments, respectively. Each treatment was designed with three replications. When the hypocotyl elongation was 1 cm, the most uniform seedlings were selected and placed in an 84-hole tray of 5 cm × 5 cm. Then, the seedling pan was placed in a culture pot with four salt concentrations. For each treatment, the phenotypic data of ten seedlings of each of the 256 lines were used for QTL analysis.
Trait measurements
The germination rate (GR) and germination potential (GP) of wheat seeds were determined after treatment with four different salt concentrations for 4 and 7 days, respectively. Meanwhile, after 7 days of salt treatment, the ten most uniform seeding samples from each line and parents were collected and rinsed with distilled water. The roots and the shoots were separately harvested. Mean root length (MRL) and shoot height (SH) were recorded. The root fresh weight (RFW) and shoot fresh weight (SFW) were weighed, and then they were oven-dried at 103 °C for 1 h and then at 80 °C for 8 h. The root dry weight (RDW) and the shoot dry weight (SDW) were weighed.
Statistical and QTL analysis
Statistical analyses (e.g., normal distribution and correlation) were performed using SPSS 17.0 software (SPSS, Chicago, USA). Trait measurements were averaged over three replications prior to QTL analysis.
The linkage map of the RIL population was constructed in a previous study42. A total of 501 markers were mapped, including 479 DArT markers, 17 SSR markers, 2 HMW-GS markers, and 3 Wx protein markers. It covered 4213.2 cM with an average distance of 8.4 cM, producing 25 linkage maps. These markers were identified on 21 chromosomes.
Conditional genetic analysis was conducted according to Deng et al.’s described method42. It was based on the phenotypic values under treatment 2 conditioned on treatment 1, which were obtained by the mixed-model approach51. Conditional phenotypic values y(T2|T1) were obtained by the mixed model approach for the conditional analysis of quantitative traits, where T2| T1 means treatment 2 conditioned on treatment 1. QGAStation 1.051 software was used to calculate the conditional phenotypic values y(T1|T2), which were subsequently used as input data for conditional QTL mapping.
Unconditional and conditional QTL mapping were performed using the software QTL IciMapping V4.152. The LOD threshold was set to 2.5. To clarify the designations of the examined QTLs, the following rules were adopted: ‘Q’ is an abbreviation of its corresponding trait, whereas a numerical number followed by an upper case letter, ‘A’, ‘B’, or ‘D’, is an indication of the chromosome number present in a given wheat genome where the corresponding QTL was detected, and if there is more than one QTL on one chromosome, a serial number behind a hyphen is added (e.g., QMrl3B.2 stands for the second QTL for MRL was detected on chromosome 3B).
Forecasting candidate genes for salt tolerance at the seed germination and seedling stages
To identify the position of important QTL loci on a physical map and possible candidate genes, significant markers detected in this study were used to identify putative candidate genes according to the method of Ji53. A BLAST (Basic Local Alignment Search Tool) search was performed on the International Wheat Genome Sequencing Consortium database (wheat Chinese Spring IWGSC RefSeq v1.1 and v2.1 genome assembly; http://www.wheatgenome.org/) using the sequence of the significant DArT markers identified by QTL mapping. When a DArT marker sequence from the IWGSC was 100% identical to any wheat contig, the sequence was extended 2 Mb for each marker using the IWGSC BLAST results53. Then, the extended sequence was used to run a BLAST search at the National Center for Biotechnology Information (NCBI) database (http://www.ncbi.nlm.nih.gov) and Ensembl Plants (http://plants.ensembl.org/Triticum_aestivum/Tools/Blast) to confirm possible candidate genes and functions53.
Ethics approval
Wheat is a common crop extensively cultivated worldwide. This study does not contain any research requiring ethical consent or approval.
Data availability
All data used during the current study are included in this published article or are available from the corresponding author upon reasonable request.
References
Sajid, H. et al. Effects of salt stress on rice growth, development characteristics, and the regulating ways: A review. J. Integr. Agric. 11, 5–22 (2017).
Wang, M. & Xia, G. The landscape of molecular mechanisms for salt tolerance in wheat. Crop J. 6(1), 42–47 (2018).
Wang, D. et al. Molecular genetic and genomic analysis of wheat milling and end-use traits in China: Progress and perspectives. Crop J. 6(1), 68–81 (2018).
Munns, R. & James, R. A. Screening methods for salinity tolerance, a case study with tetraploid wheat. Plant Soil 253, 201–218 (2003).
Yamaguchi, T. & Blumwald, E. Developing salt-tolerant crop plants: Challenges and opportunities. Trends Plant Sci. 10(12), 615–620 (2005).
Tang, Y. L. et al. Identification of QTLs for yield-related traits in the recombinant inbred line population derived from the cross between a synthetic hexaploid wheat derived variety chuanmai 42 and a Chinese Elite Chuannong 16. Agric. Sci. (China) 10, 1665–1680 (2011).
Jia, M. et al. Wheat functional genomics in the era of next generation sequencing: An update. Crop J. 6(1), 7–14 (2018).
Rasheed, A. et al. Crop breeding chips and genotyping platforms: Progress, challenges, and perspectives. Mol. Plant 10(8), 1047–1064 (2017).
Bahrani, A. & Hagh-Joo, M. Response of some wheat (Triticum aestivum L.) genotypes to salinity at germination and early seedling growth stages. World Appl. Sci. J. 16, 599–609 (2012).
Meng, X. et al. Effect of salt stress on germination of wheat and screening of salt tolerance indices. Acta Agricult. Boreali-Sin. 29(4), 175–180 (2014).
Munns, R. & Tester, M. Mechanisms of salinity tolerance. Annu. Rev. Plant Biol. 59, 651–681 (2008).
Doerge, R. W. Mapping and analysis of quantitative trait loci in experimental populations. Nat. Rev. Genet. 3(1), 43–52 (2002).
Gao, S. et al. Construction of wheat genetic map and QTL analysis of main agronomic traits using SNP genotyping chips technology. Chin. J. Appl. Environ. Biol. 22(1), 0085–0094 (2016).
Quarrie, S. A. et al. A high-density genetic map of hexaploid wheat (Triticum aestivum L.) from the cross Chinese Spring 9 SQ1 and its use to compare QTLs for grain yield across a range of environments. Theor. Appl. Genet. 110, 865–880 (2005).
Takagi, H. et al. MutMap accelerates breeding of a salt-tolerant rice cultivar. Nat. Biotechnol. 33, 445–449 (2015).
Cui, D. et al. QTL mapping for salt tolerance based on snp markers at the seedling stage in maize (Zea mays L.). Euphytica 203(2), 273–283 (2015).
Ghaedrahmati, M. et al. Mapping QTLs associated with salt tolerance related traits in seedling stage of wheat (Triticum aestivum L.). J. Agr. Sci. Tech. 16, 1413–1428 (2014).
Xu, Y. et al. Mapping QTLs for salt tolerance with additive, epistatic and QTL 3 treatment interaction effects at seedling stage in wheat. Plant Breed. 132, 276–283 (2013).
Hussain, B., Lucas, S. J., Ozturk, L. & Budak, H. Mapping QTLs conferring salt tolerance and micronutrient concentrations at seedling stage in wheat. Sci. Rep. 7, 1566 (2017).
Devi, R. et al. QTL mapping for salt tolerance associated traits in wheat (Triticum aestivum L.). Euphytica 215, 210 (2019).
Li, L. et al. Genetic insights into natural variation underlying salt tolerance in wheat. J. Exp. Bot. 72, 1135–1150 (2021).
Oyiga, B. C., Sharma, R. C., Baum, M., Ogbonnaya, F. C. & Léon-Ballvora, J. A. Allelic variations and diferential expressions detected at quantitative trait loci for salt stress tolerance in wheat. Plant Cell Environ. 41, 919–935 (2018).
Hu, P. et al. Genome-wide association study of yield and related traits in common wheat under salt-stress conditions. BMC Plant Biol. 21, 27 (2021).
Rahnama, A., James, R. A., Poustini, K. & Munns, R. Stomatal conductance as a screen for osmotic stress tolerance in durum wheat growing in saline soil. Funct. Plant Biol. 37(3), 255–263 (2010).
Ahanger, M. A. et al. Plant responses to environmental stresses—from gene to biotechnology. AoB Plants 9, 4 (2017).
Munns, R. Genes and salt tolerance: Bringing them together. New Phytol. 167(3), 645–663 (2005).
Munns, R. Comparative physiology of salt and water stress. Plant Cell Environ. 25(2), 239–250 (2002).
James, R. A., Blake, C., Byrt, C. S. & Munns, R. Major genes for Na+ exclusion, Nax1 and Nax2 (wheat HKT1; 4 and HKT1; 5), decrease Na+ accumulation in bread wheat leaves under saline and waterlogged conditions. J. Exp. Bot. 62(8), 2939–2947 (2011).
Jiang, W. et al. Genome-wide identification and characterization of SRO gene family in wheat: Molecular evolution and expression profiles during different stresses. Plant Physiol. Biochem. 154, 590–611 (2020).
Deng, X. et al. TaCIPK29, a CBL-interacting protein kinase gene from wheat, confers salt stress tolerance in transgenic tobacco. PLoS ONE 8(7), e69881 (2013).
Wang, M. et al. Ta CYP 81D5, one member in a wheat cytochrome P450 gene cluster, confers salinity tolerance via reactive oxygen species scavenging. Plant Biotechnol. J. 18(3), 791–804 (2020).
Hu, W. et al. Overexpression of a wheat aquaporin gene, TaAQP8, enhances salt stress tolerance in transgenic tobacco. Plant Cell Physiol. 53(12), 2127–2141 (2012).
Yarra, R. The wheat NHX gene family: Potential role in improving salinity stress tolerance of plants. Plant Gene 18, 100178 (2019).
Yarra, R. et al. Overexpression of a wheat Na+/H+ antiporter gene (TaNHX2) enhances tolerance to salt stress in transgenic tomato plants (Solanum lycopersicum L.). Plant Cell Tissue Organ Culture 111(1), 49–57 (2012).
Ayed, S. et al. Effect of salt stress (sodium chloride) on germination and seedling growth of durum wheat (Triticum durum Desf.) genotypes. Int. J. Biodivers. Conserv. 6(3), 20–325 (2014).
Gupta, P. K., Balyan, H. S., Sharma, S. & Kumar, R. Genetic of yield, abiotic stress tolerance and biofortification in wheat (Triticum aestivum L.). Theor. Appl. Genet. 133, 1569–1602 (2020).
Dubcovsky, J., Sanata, M. G., Epstein, E., Luo, M. C. & Dvorak, J. Mapping of K+/Na+ discrimination locus Kna1 in wheat. Theor Appl Genet 2, 448–454 (1996).
Edwards, J., Shavrukov, Y., Ramsey, C., Tester, M. & Langridge, P. Identifcation of a QTL on chromosome 7ASfor sodium exclusion in bread wheat. In (eds Appels, R. et al.) Proceedings of 11th international wheat genetics symposium. Sydney University Press, Australia (2008)
Yang, C. et al. QTL mapping for seedling height and taproot length in wheat under different salt stresses. Acta Agric. Boreali-Sin. 27(2), 91–96 (2012).
Zhou, S., Wu, Q., Xie, J., Chen, J. & Liu, Z. Mapping QTLs for wheat seedling traits in RILs population of Yanda 1817×Beinong 6 under normal and salt-stress conditions. Acta Agronom. Sin. 42(12), 1764–1778 (2016).
Zhu, J. Analysis of conditional genetic effects and variance components in developmental genetics. Genetics 141, 1633–1639 (1995).
Deng, Z. Y. et al. Genetic dissection of flour whiteness by unconditional and conditional quantitative trait locus mapping in wheat. J. Agric. Sci. 155, 556–568 (2017).
Masoudi, B. et al. QTL mapping of salt tolerance traits with different effects at the seedling stage of bread wheat. Plant Mol. Biol Rep. 33, 1790–1803 (2015).
Genc, Y. et al. Sodium exclusion QTL associated with improved seedling growth in bread wheat under salinity stress. Theor. Appl. Genet. 121, 877–894 (2010).
Lindsay, M. P., Lagudah, E. S., Hare, R. A. & Munns, R. A locus for sodium exclusion (Nax1), a trait for salt tolerance, mapped in durum wheat. Funct. Plant Biol. 31, 1105–1114 (2004).
Maccaferri, M. et al. A high-density, SNP-based consensus map of tetraploid wheat as a bridge to integrate durumand bread wheat genomics and breeding. Plant Biotechnol. J. 12, 648–663 (2015).
Tian, B., Deng, Z. Y., Xie, Q. G. & Tian, J. C. Genetic dissection of the developmental behaviour of total starch content and its components in wheat grain. Crop Past. Sci. 66, 445–455 (2015).
Dong, W. et al. Wheat oxophytodienoate reductase gene TaOPR1 confers salinity tolerance via enhancement of abscisic acid signaling andreactive oxygen species scavenging. Plant Physiol. 161, 1217–1228 (2013).
Zhang, L., Zhao, G., Jia, J., Liu, X. & Kong, X. Molecular characterization of 60 isolated wheat MYB genes and analysis of their expression during abiotic stress. J. Exp. Bot. 63, 203–214 (2012).
Zhao, Y. et al. A wheat allene oxide cyclase gene enhances salinity tolerance via jasmonate signaling. Plant Physiol. 164, 1068–1076 (2014).
Liu, G. F., Yang, J., Xu, H. M., Hayat, Y. & Zhu, J. Genetic analysis of grain yield conditioned on its component traits in rice (Oryza sativa L.). Austral. J. Agric. Res. 59, 189–195 (2008).
Meng, L., Li, H., Zhang, L. & Wang, J. QTL IciMapping: Integrated software for genetic linkage map construction and quantitative trait locus mapping in bi-parental populations. Crop J. 3, 269–283 (2015).
Ji, M. et al. Genome wide association study of the whiteness and colour related traits of four and dough sheets in common wheat. Sci. Rep. 11, 8790 (2021).
Khan, S. QTL mapping: A Tool for improvement in crop plants. Res. J. Recent Sci. 4, 7–12 (2015).
Li, X. K. et al. Screening and evaluation of salt-tolerant germplasm of synthetic hexaploid wheat. J. Triticeae Crops 41(12), 1487–1495 (2021).
Masoudi, B. et al. QTL mapping of salt tolerance traits with different effects at the seedling stage of bread wheat. Plant Mol. Biol. Rep. 33, 1790–1803 (2015).
Funding
This work was supported by the Shandong Provincial Agriculture Liangzhong Project Foundation of China (No. 2019LZGC01702), the Natural Science Foundation of China (No. 31871613), and the Shandong “Double Tops” Program.
Author information
Authors and Affiliations
Contributions
Z.D. designed and revised this paper, X.G. analysed the data and wrote the manuscript, C.W., D.W., G.W., and K.J. investigated and analysed the phenotypic data, and Y.Z. screened the candidate genes, and J.T. reviewed this paper. All authors have read and approved this manuscript. The authors consented for publication.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Guo, X., Wu, C., Wang, D. et al. Conditional QTL mapping for seed germination and seedling traits under salt stress and candidate gene prediction in wheat. Sci Rep 12, 21010 (2022). https://doi.org/10.1038/s41598-022-25703-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-25703-3
This article is cited by
-
A wheat WRKY transcription factor TaWRKY17 enhances tolerance to salt stress in transgenic Arabidopsis and wheat plant
Plant Molecular Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.