Abstract
Artemisia argyi (AA) has been proven to be effective in the adjuvant treatment of rheumatism (RA), but the mechanism of its action in RA is not clear. This study aims to clarify the molecular mechanism of AA as a potential therapy for RA by using network pharmacology. The TCM systems pharmacology (TCMSP) was used to screen the active components of AA, and identification of the potential target genes of active compounds and rheumatism was performed with PharmMapper and GeneCards, respectively. Construction of complex target networks and protein–protein interaction networks was based on the Cytoscape software. The biological functions and pathway analysis of targets and effective targets were analyzed using DAVID. Our study demonstrated that 105 target genes were associated with these active compounds and RA. ALB, AKT1, and MAPK1 were the first three hub genes, and the metabolic and signaling pathways related to these hub genes were remarkably abundant. Results showed that AA might play a role in RA by affecting multiple targets and multiple ways, reflecting that TCM was characterized by multicomponents and multitargets. AA has the potential to be a promising new candidate for the treatment of RA and has value for further research and development.
Similar content being viewed by others
Introduction
Rheumatism is a kind of disease that predominantly affects bone, joint, synovial bursa, tendon, fascia, muscle, nerve and other surrounding soft tissue. The prevalence of rheumatic diseases ranges from 11.6% to 46.4%, depending on the region, study protocol, and age of respondents. The prevalence of symptomatic osteoarthritis ranges from 5.1% to 20.8%, with the lumbar spine, knees, and cervical spine as the most affecte1. The disease is complex, lingering, and difficult to recover and has a high disability rate, thereby seriously threatening the physical and mental health of human beings. The main drugs used to treat rheumatoid arthritis (RA) usually have significant side effects2,3. Moxibustion is considered a safe and effective way to treat RA4.
Moxibustion, a traditional Chinese therapy, has been used for thousands of years in China and its neighbor5. In clinical practice, moxibustion therapy is usually used to treat RA and includes traditional (pure moxa stick) or indirect moxibustion methods, which are performed by implanting insulation materials between the acupoints and moxa stick6,7,8. Moxibustion can improve the immune function of the body through anti-inflammatory and immune effects, inhibit the secretion of synovial cytokines, and play a therapeutic role in rheumatoid arthritis9. Referring to the theory of TCM, moxibustion can help to warm and activate meridians and promote blood circulation and qi5. The results of randomized clinical trials by meta-analysis showed that the combination of moxibustion and drug therapy has a favorable effect in terms of effectiveness compared with conventional drug therapy alone10. Researchers analyzed the efficacy and safety of moxibustion in the treatment of knee osteoarthritis and showed that moxibustion has significant curative effect compared with Western medicine11. The clinical and experimental studies have confirmed that moxibustion may relieve the symptoms of osteoporosis, which can be explained that regulation of the bone metabolism index and endocrine and protein expression levels will help to reach the steady state of bone metabolism and inhibit osteoclast activity, thereby reducing bone resorption12,13. All the above studies supported that moxibustion is effective in the treatment of RA.
The leaves of Artemisia argyi (AA) are a common Chinese herbal medicine, which can be used as medicine or food, and have dual value of medicine and food. They are widely used to produce moxa stick as the main raw material in China5,14. In the moxibustion treatment, the antioxidants in AA combustion products adhere to the skin at acupoints and penetrate the body through the heat of moxibustion. Therefore, AA, as the raw material of moxibustion, plays a key role in the treatment of RA. However, the pharmacological mechanism of AA is still unclear, and no relevant research is available, which affects its clinical application and the research and development of related new drugs15. Network pharmacology based on the theory of system biology, whose systematic and multi-dimensional analysis characteristics are consistent with the holistic view of TCM, provides a powerful tool for revealing the complex mechanism of TCM. The research shows that the trend of combining calculation, experiment and clinic is a promising development direction of TCM network pharmacology16. In our study, network pharmacology method was used to predict and analyze the mechanism of AA in the treatment of RA from the molecular level, so as to provide scientific basis for the clinical application of AA and the development of new drugs.
Materials and methods
Screening for active components of AA
The principal components of AA were referred to the TCM systems pharmacology (TCMSP) database and analysis platfor17.
Oral bioavailability (OB) is a major index that reflects the degree to which oral drugs reach the targets by passing through intestinal wall barriers. Compounds with low OB may exhibit minimal effect because few active components appear in the blood. The drug-likeness (DL) index is applied in the identification of drugs and nondrugs, and compounds with low DL values may not be drugs18. Brain capillary wall and glial cells formed a barrier between plasma and brain cells, and so do choroid plexus formed between plasma and cerebrospinal fluid, which are called the blood–brain barrier (BBB), preventing certain substances (mainly harmful) from entering brain tissue through blood. The permeability of BBB is an indicator of drug permeability. In our research, 30%, 0.18, and 0.3 were selected as the best cutoff values of OB, DL, and BBB, respectively19,20. The components of AA with values higher than these reference standards were regarded as functional constituents.
Prediction of target genes
To investigate the relation between each selected active compound and its target genes, the main active ingredients of AA were organized in SDF and MOL2 formats, and then uploaded to the PharmMapper server (http://59.78.98.102/pharmmapper/) and analyze21,22,23. Using “human protein targets are only used to select target sets” as the search term, the rest were set by default. The selected protein target was imported into the UniProt database (https://www.uniprot.org/) in PDB ID format. The prediction indexes of the effective components of AA were achieved by searching and converting.
Screening of rheumatic disease targets
The keyword “rheumatic diseases” was used to search GeneCards (https://www.genecards.org/) to obtain the reported genes associated with rheumatic diseases.
Network construction and analysis
The target information of effective components in AA and disease target genes was processed with the Cytoscape 3.8 software (http://cytoscape.org/), and used to construct the model of the medicine-ingredients-disease-targets (M-I-D-T) network. In this network model, nodes represented molecules or target proteins, whereas edges represented relationships among components, disease, and targets.
Establishment of the protein–protein interaction network
The Search Tool for the Retrieval of Interacting Genes (STRING) database is an effective tool for exploring and constructing protein–protein interaction (PPI) networks24. By using the online STRING tool (version 11.0), the PPI of target genes was picked out while a required interaction score was above 0.4, and the target genes of PPI network having high connectivity degrees were selected as hub genes.
GO and KEGG pathway analysis
The biological functions of these selected genes were analyzed using the Database for Annotation, Visualization and Integrated Discovery (DAVID), so do pathway analysis of target biological functions and effective targets. Data were exported from David and screened for count > 5, p-value < 0.05, and FDR < 0.05. By using Cytoscape, a compound–target–pathway network was set up to understand the relationship among RA, targets, and compounds systematically.
Molecular docking
The 3D structure of key compounds was made by Chem Office software, then the 3D structure of core protein gene was downloaded from PDB database, the protein was dehydrated and dephosphorized by PyMOL software, the pdb format of compound and core protein gene was converted into pdbqt format and active pocket was found by AutoDock 1.5.6 software, and finally, Vina was run for docking. The binding activity of the two is evaluated according to the binding energy, and the binding energy ≤ − 5.0 kJ/mol is selected as the screening basis to evaluate the reliability of network analysis and prediction.
Results
Screening of active compounds
By considering OB ≥ 30%, DL ≥ 0.18, and BBB ≥ 0.3, TCMSP results showed 135 traditional components in AA, among which seven active compounds were retrieved from TCMSP (Table 1). The PharmMapper integrated pharmacophore matching platform was used for reverse prediction of these active compounds.
Target gene prediction
We identified 1134 target genes corresponding to RA from GeneCards and 431 target genes for active compounds from the Uniprot identifiers map after elimination of duplicates. On the basis of the above results, 105 overlapping genes between RA and active components were obtained (Fig. 1A).
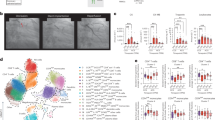
Construction of the compound-disease network. (A) Venn diagram of target genes for RA and active compounds. RA has 1134 target genes, whereas active components have 431 target genes. A total of 105 overlapping target genes are found between the two sets, (B) Medicine-ingredient-target-disease network with four parts (Red: Artemisia argyi (AA), green: RA, yellow: active ingredients of AA, blue: 105 potential common targets) was generated by Cytoscape (Version, 3.8) (http://cytoscape.org/).
M-I-D-T network
Through the treatment of the software of Cytoscape, a M-I-D-T network was produced and showed that the four components were closely related to one another (Fig. 1B).
PPI network
On the basis of the STRING database, we generated a PPI network containing 105 overlapping target genes (Fig. 2). Each protein was represented by a node, and the interaction was represented by lines between nodes. The number of lines connected to a specified node was considered as the connectivity degree. In the present research, hub genes were defined as genes with connectivity degree > 5. As shown in Fig. 3, among 20 hub genes, ALB, AKT1, MAPK1, MAPK8, MMP9, EGFR, CASP3, and IGF1 were the top 8 hub genes.
Protein–protein interaction (PPI) network analysis of 105 potential targets was generated by String (Version, 11.5) (https://cn.string-db.org/).
GO and KEGG pathway enrichment analyses
Based on the GO enrichment analysis, major targets could be separated into various functional modules, and the GO enrichment analysis was performed using the DAVID ver 6.8 gene annotation platform. Data were exported from the DAVID database, and the data of biological process (BP), molecular function (MF), and cellular component (CC) were filtered by considering p-value < 0.05 and FDR < 0.05.
As shown in Fig. 4A, BP was dominated by the proteolysis, extracellular matrix disassembly, response to hypoxia, negative regulation of apoptotic process and collagen catabolic process. CC was dominated by extracellular region, extracellular space, ficolin-1-rich granule lumen, extracellular exosome and cytosol. MF was dominated by zinc ion binding, the same protein binding, serine-type endopeptidase activity, drug binding and receptor binding. All the above GO entries played important roles in RA.
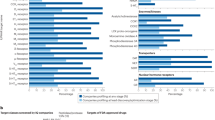
Functional enrichment analysis. (A) Enrichment analysis of potential targets of the active components of AA in rheumatism. The analysis of the Gene Ontology (GO) terms for biological process (BP), cellular component (CC), and molecular function (MF) is shown. LogP is the log-value of the P-value. P < 0.05 is considered significant. The results of the enrichment analysis are shown by LogP to present results intuitively. The count represents the number of genes, (B) Bubble diagram of KEGG enrichment analysis. The y-axis represents pathway and the x-axis represents gene ratio (amount of genes enriched in the pathway/amount of all genes in the background gene set). The size and color of the bubble represent the number of genes enriched in pathway and enrichment significance, respectively.
Thirteen pathways were selected in accordance with p < 0.05 and FDR < 0.05 (Fig. 4B) to clarify the key pathways of 105 potential targets for rheumatism treatment. These pathways included those involved in cancer (hsa05200), pancreatic cancer (hsa05212), estrogen signaling pathway (hsa04915), osteoclast differentiation (hsa04380), progesterone-mediated oocyte maturation (hsa04914), hepatitis B (hsa05161), Chagas disease (American trypanosomiasis, hsa05142), FoxO signaling pathway (hsa04068), protein-polysaccharide in cancer (hsa05205), toxoplasmosis (hsa05145), colorectal cancer (hsa05210), Ras signal pathway (hsa04014), and prolactin signal pathway (hsa04917).
GO was annotated, and the response pathway was analyzed. By importing the effective target proteins, chemical components, and pathways into Cytoscape software, the compound-target-pathway network diagrams were set up (Fig. 5).
The compound target network associated with AA in rheumatism was produced by Cytoscape (Version, 3.8) (http://cytoscape.org/). The rectangle, triangle, and circle represent the component, target, and rheumatism-related pathway.
Molecular docking results
Taking the binding energy ≤ − 5.0 kJ/mol as the standard, the molecular docking verification of key active components and key targets was carried out. The results show that the affinity of the two docking sites is less than that of − 5.0 kJ/mol, which indicates that there is a good binding activity between the key active components and key targets, proving that the prediction of this study is more reliable, as shown in Table 2. The molecular docking display is illustrated in Fig. 6.
Discussion
RA is a chronic inflammatory and autoimmune disease that is distinguished by the excessive proliferation of synovium and inflammatory cell infiltration, causing the chronic inflammation of joints and secondary erosion of cartilage and bone. The main causes of the disease are immune response, infection, and endocrine disorders. In the present study, we propose a network pharmacology method that combines the strategy of component prediction and target path analysis. This method is used to explore the potential regulatory effects of AA on inflammation, immune system, and angiogenesis, providing a new perspective for the treatment of RA. The potential cooperation mechanism of EEAA is predicted and analyzed from the point of view of biological network, and preliminary experimental validations are performed.
In this study, 135 active components of AA and 105 potential targets of AA in the treatment of RA were screened out. PPI network identified ALB, AKT1, MAPK1, MAPK8, MMP9, EGFR and CASP3 as key potential targets. The molecular docking results showed that the active ingredients had good binding activity with potential targets. KEGG results showed the top five pathways (cancer, pancreatic cancer, et al.) are directly or indirectly involved in the formation of RF. Based on the experimental results and a number of articles on the therapy of RA by AA, the authors concluded that AA plays an important role in the treatment of RA by regulating the proliferation and differentiation of bone cells and osteoclasts and immune-inflammatory response.
The destruction of articular cartilage and bone is the main cause of disability. In recent years, a large number of related research demonstrated that osteoclasts could be the key to the pathological process of bone destruction in RA. The main active ingredient beta-sitosterol in AA can regulate the proliferation and differentiation of osteocytes and osteoclasts, so as to achieve the effect of treating RA25. AKT1 and MAPK8, as hub genes, play a major role in osteoclast differentiation. AKT1, a unique signaling intermediate in osteoblasts, can regulate the differentiation of osteoblast and osteoclast at the same time. The targeted inhibition of AKT1 can result in effective bone acquisition, and related AKT1 and AKT2 have significant influence on the cell differentiation pathway26. The MAPK8 signal body is the target to prevent joint destruction27. MAPK is a collection of evolutionarily conserved serine/threonine protein kinases, which are activated by certain extracellular stimuli and may startup the signal transmition from cell membrane to nuclear. The activation of the MAPK pathway can induce the expression of heat shock protein (which has the function of regulating cell proliferation) in osteoblasts, thus promoting the formation of osteoblasts. KEGG pathway analysis demonstrates that osteoclast differentiation and Ras signal pathway are involved in the formation of RA. Therefore, AA may serve a purpose in the treatment of RA by regulating the proliferation and differentiation of osteoclasts and osteoclasts.
RA is a chronic inflammatory and autoimmune disease characterized by excessive synovial proliferation and inflammatory cell infiltration, resulting in chronic inflammation of joints and resultant erosion of cartilage and bone. Therefore, AA plays an important role in the treatment of RA by regulating immune-inflammatory response. The main active ingredient stigmasterol in AA can regulate the expression of inflammatory factors, thus achieving anti-inflammatory effect28. In addition, MAPK8 is also called JNK, and its activation in RA synovium can mediate articular damage in rats with adjuvant-induced arthritis29. KEGG results showed that FOXO signaling pathway is directly associated with the formation of RA. FOXO3a can mediate TNF-α through PI3K/AKT signaling pathway to inhibit the proliferation and invasion of trophoblast cells and promote their apoptosis, thus being expected to result in inflammation in the body30. In vitro experiment, AA could significantly down regulate the level of IL-17, IL-1βand TNF-α in RA rats31. Therefore, the authors suggest that AA achieves its therapeutic effect on rheumatoid arthritis mainly by regulating the immune-inflammatory response. Therefore, MAPKs and Akt signal pathway are promising targets for RA therapy, and the research and development of corresponding inhibitors for both pathways are hot spots in the research and development of RA therapeutic drugs. These results demonstrated the complicated relationship among compounds, targets, and pathways of AA in rheumatism32.
However, this research has the following three limitations: (1) all the prediction results were achieved by analysis based on limited online databases, which are kept updating. (2) The composition of AA is so complex that some compounds not identified are likely not involved in the analysis and discussion. (3) The future study should focus on verification experiments which help to explain the prediction results of this study.
Conclusion
TCM usually plays an important role to prevent and treat various diseases. Therefore, in this paper, the network pharmacology method was used to screen RA-related targets and AA signaling pathways. Thus, the genomic space might be connected to the pharmacological space. In conclusion, we predicted the mechanism of action of EEAA in the treatment of RA by analyzing key targets and KEGG pathway. The methods applied to TCM should correspond to the synergistic mechanism.
Data availability
Data inquiries can be directed to the corresponding author.
References
Zeng, Q. Y., Chen, R., Darmawan, J., Xiao, Z. Y. & Chen, S. B. Rheumatic diseases in China. Arthritis. Res. Ther. 10, 1–11 (2008).
Guo, J. C. Rationality and safety evaluation of 180 cases of rheumatoid arthritis non-steroidal anti-inflammatory drugs. Capital Food Med. 27, 83 (2020) (Chinese).
Wang, D., Liu, J., Chen, Y. & Feng, D. Association between MTHFRC677T polymorphism and adverse drug reactions of methotrexate in rheumatoid arthritis patients. Shandong Med. J. 61, 11–14 (2021) (Chinese).
Ma, W., Gou, B. H., Xie, C., Chen, Y. & Cao, L. Effects of moxibustion combined with tripterygium wilfordii hook in patients with rheumatoid arthritis. World J. Tradit. Chin. Med. 15, 473–476 (2020) (Chinese).
Huang, C. et al. Moxibustion in early Chinese medicine and its relation to the origin of meridians: a study on the unearthed literatures. Evid. Based Complement Altern. Med. 2017, 1–9 (2017).
Xu, D. M., Xu, H., Jing, L., Tong, W. & Cao, Y. Effect of thunder-fire moxibustion on pain, quality of life, and tension of multifidus in patients with primary osteoporosis: A randomized controlled trial. Med. Sci. Monit. 24, 2937–2945 (2018).
Zhang, M., Zhao, C., Jiang, L. & Zhu, Y. Clinical effect and mechanism of moxibustion combined with western medication for rheumatoid arthritis of liver-kidney deficiency. Chin. Acup. Moxib. 41, 489–492 (2021).
Hong, Y. et al. Clinical efficacy of moxibustion as supplement on rheumatoid arthritis and the exploration on its mechanism. Chin. Acup. Moxib. 36, 1–17 (2016).
Zhang, C. Y. & Tang, Z. L. Progress of mechanism study on rheumatoid arthritis treated by moxibustion. J. Acupunct. Tuina. Sci. 7, 65–70 (2009).
Choi, T. Y., Kim, T. H., Kang, J. W., Lee, M. S. & Ernst, E. Moxibustion for rheumatic conditions: A systematic review and meta-analysis. Clin. Rheumatol. 30, 937–945 (2011).
Yuan, T. et al. The effectiveness and safety of moxibustion for treating knee osteoarthritis: A PRISMA compliant systematic review and meta-analysis of randomized controlled trials. Pain Res. Manag. 2019, 2653792 (2019).
Zhang, H. S., Li, T., Liu, X. S., Wang, F. C. & Liang, W. J. Experimental study of the effect on bone metabolism and bone histomorphometry of osteoporosis rats with birdpecking and revolving moxibustion on twelve back-shu points. Cell Biochem. Biophys. 71, 173–178 (2015).
Pang, W. P. The impact on estrogen and bone mineral density through the use of moxibustion on postmenopual osteoporosis rats. J. Guangzhou Univ. Tradit. Med. China 1–27 (2009) Chinese.
Lim, M. Y., Huang, J., Zhao, B. X., Zou, H. Q. & Yan, Y. H. Influence of storage duration and processing on chromatic attributes and flavonoid content of moxa floss. J. Integr. Med. 14, 69–76 (2016).
Hopkins, A. L. Network pharmacology: The next paradigm in drug discovery. Nat. Chem. Biol. 4, 682–90 (2008).
Wang, X., Wang, Z. Y., Zheng, J. H. & Li, S. TCM network pharmacology: A new trend towards combining computational, experimental and clinical approaches. Chin. J. Nat. Med. 19(1), 1–11 (2021).
Ru, J. et al. TCMSP: A database of systems pharmacology for drug discovery from herbal medicines. J. Cheminform. 6, 13 (2014).
Brustle, M. et al. Physical properties, and drug-likeness. J. Med. Chem. 45, 3345–3355 (2002).
Wang, T. et al. Network pharmacology-based prediction of the active ingredients and potential targets of Chinese herbal Radix Curcumae formula for application to cardiovascular disease. J. Ethnopharmacol. 145, 1–10 (2013).
Tattersall, M. H. et al. Pharmacokinetics of actinoymcin D in patients with malignant melanoma. Clin. Pharmacol. Ther. 17, 701–708 (1975).
Liu, X. et al. PharmMapper server: A web server for potential drug target identification using pharmacophore mapping approach. Nucleic Acids Res. 38, w609–w614 (2010).
Wang, X., Pan, C. X., Gong, J. Y., Liu, X. F. & Li, H. L. Enhancing the enrichment of pharmacophore-based target prediction for the polypharmacological profiles of drugs. J. Chem. Inf. Model. 56, 1175–1183 (2016).
Wang, X. et al. PharmMapper2017 update: A web server for potential drug target identification with a comprehensive target phar-macophore database. Nucleic Acids Res. 45, W356–W360 (2017).
Szklarczyk, D. et al. STRING v11: Protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47, D607–D613 (2019).
Zeng, L. P. et al. Regulation of β-sitosterol in duzhongye on homeostasis of bone metabolism. Lishizhen Med. Mater. Med. Res. 23, 1051–1053 (2012) (Chinese).
Mukherjee, A. & Rotwein, P. Selective signaling by Akt1 controls osteoblast differentiation and osteoblast-mediated osteoclast development. Mol. Cell Biol. 32, 490–500 (2012).
Sundarrajan, M., Boyle, D. L., Chabaud-Riou, M., Hammaker, D. & Firestein, G. S. Firestein, Expression of the MAPK kinases MKK-4 and MKK-7 in rheumatoid arthritis and their role as key regulators of JNK. Arthritis. Rheum. 48, 2450–2460 (2003).
Wu, L. C., Li, J. F. & Zhang, T. T. Study on anti-inflammatory effect of stigmasterol based on network pharmacology and cell experiment. Chin. Tradit. Patent Med. 44, 609–615 (2022) (Chinese).
Schett, G. et al. Activation, differential localization, and regulation of the stress-activated protein kinases, extracellular signal-regulated kinase, c-JUN N-terminal kinase, and p38 mitogen-activated protein kinase, in synovial tissue and cells in rheumatoid arthritis. Arthritis. Rheum. 43, 2501–2512 (2000).
Zhang, X. et al. FOXO3a mediates inflammatory factor TNF-alpha through PI3K/AKT signaling pathway to inhibit the proliferation, invasion and apoptosis of trophoblasts. Immunol. J. 36, 482–489 (2020).
Wan, Y. & He, F. The effect of Cataplasma of Artemisia Argyi by supercritical CO2 fluid extraction in the treatment of rheumatoid arthritis rats. J. Zhejiang Chin. Med. Univ. 6, 839–844 (2013).
Li, S. Network pharmacology evaluation method guidance-Draft. World J. Tradit. Chin. Med. 7, 146–154 (2021).
Acknowledgements
We appreciate funding support from Doctoral Foundation Project of Huanggang normal University (No. 2019057); Hubei Key Laboratory of Economic Forest Germplasm Improvement and Resources Comprehensive Utilization, Hubei Collaborative Innovation Center for the Characteristic Resources Exploitation of Dabie Mountains Open Fund Project (No. 202141104).
Author information
Authors and Affiliations
Contributions
W.W. was responsible for the conception of experimental scheme; H.Y.Z., J.X.P. and L.J.Y. performed program implementation; W.H.P. contributed to data analysis and paper materials preparation; W.W. was in charge of writing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Huang, Y., Jin, X., Liu, J. et al. Systems pharmacology approach to investigate the mechanism of Artemisia argyi in treating rheumatic diseases. Sci Rep 12, 18786 (2022). https://doi.org/10.1038/s41598-022-23635-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-23635-6
This article is cited by
-
A review of the research progress on Artemisia argyi Folium: botany, phytochemistry, pharmacological activities, and clinical application
Naunyn-Schmiedeberg's Archives of Pharmacology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.