Abstract
Rice quality is one of the main targets of rice breeding and is a complex trait that involves grain appearance, milling, cooking, eating and nutritional quality. For many years, rice breeding has contended with imbalances in rice yield, quality, and disease and lodging resistance. Here, the milling and appearance quality, cooking quality, starch rapid viscosity analyzer (RVA) profile, and nutritional quality of grains of Yuenongsimiao (YNSM), an indica rice variety with high yield, high quality and disease resistance, were determined. YNSM had excellent appearance and quality, with low amylose contents and high gel consistency, and these characteristics exhibited significant correlations with the RVA profile such as hot paste viscosity, cool paste viscosity, setback viscosity, and consistency. Moreover, 5 genes related to length-to-width ratio (LWR) as well as the Wx gene were used to detect the main quality genotype of YNSM. The results showed that YNSM is a semilong-grain rice with a relatively high brown rice rate, milled rice rate and head rice yield and low chalkiness. The results indicated that the LWR and food quality of YNSM might be related to gs3, gw7 and Wxb. This study also reports the quality characteristics of hybrid rice developed using YNSM as a restorer line. The quality characteristics and the genotype for grain quality determined through gene analysis in YNSM may facilitate the breeding of new rice varieties that achieve a balance of grain yield, resistance and quality.
Similar content being viewed by others
Introduction
Rice (Oryza sativa L.) grain quality is one of the main targets of rice breeding. Rice grain quality involves several components, such as the grain appearance, milling, cooking, eating and nutritional quality1,2,3. In particular, cooking and eating quality traits, such as the amylose content (AC), grain thickness (GT), gel consistency (GC), pasting viscosity and aroma are important factors in determining the quality of cooked rice4.
Grain size is an important factor affecting rice yield and is also important for rice appearance quality. As a typical complex quantitative trait, grain size is closely associated with grain weight and usually measured by grain length (GL), grain width (GW), GT and length-to-width ratio (LWR)5,6. GS3, a major quantitative trait locus (QTL) for grain size, functions as a negative regulator of LWR and organ size7,8. GS9 negatively regulates glume division by the GS9-OFP8/14-GSK2 pathway6,9. GW5 negatively regulated rice GW10. GW7/GL7 is a main QTL for grain length and width. The overexpression of GW7 increases longitudinal cell division in the grain and reduces transverse cell division, resulting in more narrower grains and simultaneously improving rice yield and grain quality11,12. Additionally, GW7 expression is directly regulated by OsSPL16/GW8, a transcription factor encoded by the grain-width locus OsSPL16/GW813. However, LWR is closely related to milling quality parameters, such as the head rice yield.
Starch is the main chemical component of rice grains and is made up of amylose and amylopectin. The characteristics of starch directly affect the cooking and eating quality of rice14. AC has long been considered the most important factor in determining the quality of rice15,16. Waxy, the main gene that controls amylose synthesis, encodes granule-bound starch synthase (GBSS) and affects the AC in rice endosperm and pollen directly. In nonglutinous rice varieties, the Wx gene differentiates into Wxa and Wxb alleles; of these, wild rice and most indica rice had the Wxa genotype, with high amylose content, the vast majority of japonica rice has the Wxb genotype, with a relatively low amylose content17,18. Additionally, Wx can also control the gel consistency of rice19.
The starch viscosity profile, known as the RVA profile, refers to the curve of starch viscosity that varies with temperature during heating, a period of high temperature and cooling; the profile reveals the peak viscosity (PKV), hot paste viscosity (HPV), cool paste viscosity (CPV), peak time (PeT), paste temperature (PaT), breakdown viscosity (BDV, PKV minus HPV), consistency (CS, CPV minus HPV), and setback viscosity (SBV, PV minus PKV)20,21. It has been shown that the characteristic RVA profile values of rice starch are closely related to the rice cooking quality and food quality, especially BDV, CS and SBV22. Generally, the higher the amylose content is, the lower the BDV is, and the greater the SBV and CS are. The characteristic values of the RVA profile of rice starch can be used to distinguish differences in cooking and eating quality among rice varieties with similar apparent amylose contents.
Rice breeding in China is facing an imbalance among high yield quality, disease resistance and lodging. YNSM is a new rice variety that exhibits high yields, high quality and disease resistance that was bred by the Rice Research Institute of Guangdong Academy of Agricultural Sciences (GDAAS)23,24,25. To further reveal its quality characteristics, the milling quality, appearance quality, cooking and eating quality, nutritional quality, and RVA profile of rice starch were determined for YNSM. These results will provide a theoretical basis for the breeding of new rice varieties with high eating quality.
Results
Development of YNSM
YNSM was bred using Yuetai13, a high-yielding, disease-resistant variety, as the male parent and the good-quality variety Huanghuazhan as the female parent26. We used phenotypic selection (yield and plant height) for pedigree breeding from F3 generation23. In 2009, a line from the F8 generation with high-yielding and high resistance for blast was selected and named Yuenongsimiao, The blast resistance were detected in Conghua, Guangzhou. For its relative balance of high yield and blast resistance (Fig. 1), YNSM has been certified in Guangdong Province in 2011 (Guangdong authorized variety no.2011023), Hainan Province (2013), Jiangxi Province (2017) and Hubei Province (2017), China (http://www.ricedata.cn/variety/varis/610872.htm).
The milling and appearance quality of YNSM
The grain size of YNSM is between those of Yuexiangzhan (YXZ) and Xiangyaxiangzhan (XYXZ). The grain length is 9.28 mm, the grain width is 2.3 mm, and the LWR is 3.2–3.4; these values are different from those of the small-grain variety YXZ and the thin long-grain variety XYXZ (Fig. 2a–d). Grain size is controlled mainly by genetic factors rather than environmental factors. To reveal the regulatory factors for YNSM LWR, we used known molecular markers. The results showed that LWR in YNSM, YXZ and XYXZ was controlled by GS9, GW5, and GW8. For the GS3 gene, the long grain varieties YNSM and XYXZ had the genotype gs3, and the small variety YXZ had the genotype GS3. Moreover, unlike in YNSM and YXZ, LWR in XYXZ was found to be controlled by GW7 (Figs. 2e, S1, S2).
Grain appearance. (a), Rice grain appearance (n = 150 grains); (b), 10-grain length of 3 varieties; (c), 10-grain width of 3 varieties; (d), Statistical analysis of grain length, grain width and length/width ratio (LWR), ES, early season, LS, late season; (e), genotype of LWR genes. ES, early season; LS, late season; M, marker; Bar = 1 cm.
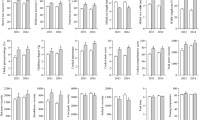
The milling quality of the rice was different in different seasons. In the early season, the brown rice rate, milled rice rate and head rice yield of YNSM were similar to those of YXZ and XYXZ, at approximately 70%, 60% and 55%, respectively. However, in the late season, all of the milling quality parameters of YNSM were much better than those of YXZ and XYXZ (Fig. 3a,b).
Generally, the white grain rate and chalkiness rate were greatly affected by the season, and the rice quality was better in the late season than in the early season. The transparency level of YNSM was 3, which was better than those of YXZ and XYXZ. The chalkiness rate and chalkiness degree of YNSM in the early season were 11.84% and 1.61%, respectively. In the late season, the chalkiness rate and chalkiness degree were 6.71% and 0.84%, respectively, which were significantly lower than those of YXZ but equal to or better than those of XYXZ (Fig. 3c).
Cooking quality characters of YNSM
The AC values of YNSM in the early and late seasons were 17.05% and 17.24%, respectively, which were lower than those of YXZ (25.05% and 26.47%) and similar to those of XYXZ (15.9% and 18.2%). The GC values of YNSM in the early and late seasons were 72.0 mm and 75.0 mm, respectively, which were similar to the values of XYXZ but significantly higher than those of YXZ (44 mm and 46 mm). These results showed that YNSM has equally low AC and high GC as other high-quality rice varieties. The eating score of YNSM (85.6–87.1/100) was similar to that of XYXZ (85.7–88.1/100) and significantly higher than that of XYZ (57.0–63.8/100) (Table 1).
Because Wx is the main gene that controls amylose synthesis, the genotype of Wx was determined in YNSM17. The results showed that YNSM and XYXZ had the Wxb genotype, while YXZ had the Wxa genotype, which was in line with the results for AC (Figs. 4, S1).
The RVA profile of YNSM
In both the early and late seasons, the starch paste viscosity of the tested rice varieties displayed an increasing–decreasing-increasing trend. In the early season, the PV of YNSM was 248.9 RVU, which was half that of YXZ and similar to that of XYXZ (Table 2). The HV of YNSM was 88.1 RVU, which was significantly less than that of YXZ (312.1 RVU) but slightly higher than that of XYXZ (52.4 RVU). The CPV of YNSM was 163.9 RVU, which was substantially lower than that of YXZ (446.3 RVU) and higher than that of XYXZ (99.2 RVU). In the early season, the BD, CS and SB of YNSM were 160.8 RVU, -85.0 RVU and 75.8 RVU, respectively. These values were similar to those of XYXZ, but the BD was higher than that of YXZ, and the BD and CS were lower than those of YXZ. In the late season, the HPV, CPV and SB of YNSM were similar to those of XYXZ, but the PV, HPV and PeT were not significantly different from those of XYXZ. These results indicated that CPV, SB and CS were significantly correlated with taste quality.
Although the RVA profiles of the rice varieties were different between the early and late seasons, the correlations between profiles for the same variety were higher than 0.96, which means the RVA profiles of each variety in different seasons is positive correlation (Table 3). In the early season, the correlation between the RVA profiles of YNSM and XYXZ was 0.977, while it was lower than 0.77 between the profile of YXZ and that of YNSM or XYXZ. In the late season, the correlation between the RVA profiles of YNSM and XYXZ was 0.99, which was stronger than the correlation between the profile of YXZ and that of YNSM (0.887) or XYXZ (0.816). In general, rice quality tends to be higher in the late season than in the early season in South China; however, the relationship of the RVA profile of YNSM in the early season and the RVA profile of XYXZ in the late season was 0.994, further suggesting that the quality of YNSM is related to the high quality of XYXZ rather than YXZ.
Protein and fatty acid content of YNSM
Rice quality includes the nutritional quality of rice, such as its storage protein, fatty acid, anthocyanin and mineral contents27,28. Rice grains contain a certain proportion of protein and fat, which are related to its nutrition, luster and palatability. In the early season, YNSM had the highest protein and fat contents of the tested varieties; nonetheless, in the late season, there were no significant differences in protein and fat content among the rice varieties (Fig. 5). The protein contents of YNSM were 7.68% in the early season and 7.52% in the late season, which were similar to those of XYZ and higher than those of XYXZ. Interestingly, the content of fatty acids decreased greatly in the late season (Fig. 5).
Quality-yield breeding applications of YNSM
YNSM not only produces excellent-quality rice but also exhibits outstanding comprehensive agronomic traits with highly resistant to blast and bacterial blight. YNSM is also an excellent restorer line (R1212) for two-line and three-line hybrid rice. At present, more than 20 hybrid rice combinations have been prepared with YNSM as a restorer line, and the rice quality of most of these hybrids is above grade 3, the national standard (Table 4, http://www.ricedata.cn/variety/). Among them, Guangtaiyou-YNSM, Jitianyou-YNSM, Huiliangyou-YNSM, Cliangyou-YNSM, Xinrongyou-YNSM, and Zaoyou-YNSM exhibited whole polished rice rates above 65%, GC values of 61–80 mm, AC values of 14.6–19.2%, and rice quality scores of to grade 2 . These results indicate that YNSM can improve the grain quality of hybrid rice.
Discussion
The influence of grain size on rice quality
Grain size is one of the most important factors influencing rice appearance quality and rice yield, and several genes related to grain size have been cloned7,8,11,12,13. GS3 and GW7 coordinately function in controlling grain size7,13,29. In this research, the genotype of the small-grain variety YXZ was GS3/gw7, that of YNSM was gs3/gw7, and that of XYXZ was gs3/GW7. Although GW7 dramatically improved the appearance quality trait, it was also associated with lower head rice yield. Slender rice, such as XYXZ, is easily broken during milling, leading to secondary food losses. Therefore, rice yield and head rice yield should be considered in rice improvement with YNSM.
Effect of starch content on taste
Amylose is the main component of grain starch. Usually, rice with more AC is not good for eating21. It has been clearly shown that Wx is the main gene that controls amylose synthesis17. For most rice varieties with good flavor, the genotype of Wx is Wxb, which causes low AC17,18. In this research, we revealed that the low AC is controlled by Wxb in the YNSM genome, is useful information for the molecular breeding of high-quality rice (Fig. 4).
Effects of RVA profile on taste
The characteristics of the rice starch RVA profile are an important factor affecting rice cooking and eating quality. The RVA profile can reflect not only the taste differences caused by differences in amylose content among different varieties but also the differences in taste and palatability that occur in rice varieties with similar amylose contents. In the RVA pasting viscosity parameters only BD, SB and CS seemed to be controlled by Wx gene, which is closed linked with good taste quality21. The RVA profile is also related to the gel consistency. Various rice varieties with low amylose contents usually have a high gel consistency, with higher BD and lower SB and CS20. In this research, YNSM and XYXZ were of similar quality, and their RVA profiles were markedly different from that of the high-AC rice variety YXZ. RVA profiles require few samples and have good repeatability; thus, they should be effectively applied in early generation selection. In this research, we found that the PaT was not significantly different among varieties but was significantly different in the different seasons, implying that the ripening temperature is important for rice quality30. Although YNSM and XYXZ both have low starch contents, there are still distinct differences in their taste quality, such as their nutrient quality content and RVA profile. Therefore, YNSM/XYXZ recombinant inbred lines (RILs) have been constructed to further reveal the genetic network of food quality.
Improvements in rice varieties with high food and taste quality
The coordination of yield resistance and quality is an important goal of breeding. At present, given the large amount of genome-wide data, the continuous advancement of molecular marker-assisted selection and molecular design breeding, and CRISPR/Cas9 technology, it is possible to improve the yield, quality and resistance of rice in a precise and efficient manner. Here, we recommend YNSM, which not only produces high-quality grains but is also high yielding and disease resistance (Figs. 1,2; Tables 4; S1). It is known that Pi2 is a broad-spectrum blast resistance gene and have been widely used in rice breeding31. In the previous research have revealed YNSM have Pi232. In this research, we uncovered that Wx, gs3/gs9/GW7 are important for the yield and quality. Hence, this study provided a new approach for rice breeding.
Methods
Materials
In this study, the indica rice variety YNSM (No: Yueshendao2011023) was tested, and the varieties Yuexiangzhan (YXZ, No: Yueshendao1998001), which has high ACC, and Xiangyaxiangzhan (XYXZ, No: Yueshendao2006044), which has low ACC and excellent flavor, were used as controls. All varieties were collected from Rice Research Institute of GDAAS, and planted at the experimental base of it in Guangzhou city (N 22°, E 112°), Guangdong Province, in the early season (planted in March) and late season (planted in July) of 2019. Before sowing, the seeds were accelerated for germination at 37 ℃, and planted at 3–4 leaf stage in the moderate fertility in the moderate-fertility soil. All varieties were planted for 3 times. Fertilization and weeding of plants in the present study complies with international, national and/or institutional guidelines.
Rice quality determination
Seeds were harvested after maturity, and 100 g of air-dried rice that was intact and free from pests and diseases was stored in a dry, ventilated indoor environment (25 ℃) for 3 months until the physicochemical properties of the seeds stabilized33. Before testing, the samples to be tested were placed in a dry, ventilated place or an air-conditioned room for approximately 7 days to keep the moisture content of the samples below 14%.
Processing quality determination
An amount of 30 g of rice grain was taken from each of the samples. Using a rice huller (JLG-III), the sample was processed into brown rice, placed into a brown rice polishing machine (JNM-III) and processed. Then, the rice was polished to a diameter of 1.5 mm and passed through round holes in a screen to obtain milled rice. Finally, the samples were sieved and isolated to obtain whole white rice. Brown rice rate = brown rice weight (g)/30 (g) × 100%, milled rice rate = white rice weight (g)/30 (g) × 100%, head rice yield = whole white rice weight (g)/30 (g) × 100%.
Appearance quality determination
An SC-E rice appearance quality detector (Wanshen Testing Technology Co. LTD, Hangzhou, China) was used to measure the length/width ratio of whole brown rice and the chalky grain rate, chalkiness degree and transparency of milled rice. Finally, all the refined rice was ground with a miniature universal pulverizer and screened with a 100-mesh sieve. The rice flour was stored at 4 °C for physical and chemical property analysis.
Evaluation of rice cooking and eating quality
The AC and GC were measured using standard procedures as described previously. The AC was determined using a colorimetric method with KI-I2 and measured by a continuous flow analyzer (Futura-II) with the manufacturer’s protocol; GC was measured as the gel length, with longer gels being considered softer than shorter gels34. The paste viscosity of rice starch was measured using a RVA profile by RVA-TecMaster (Perten, Sweden). Briefly, 3 g of rice flour was mixed with 25 mL of water in the RVA sample can. An RVA-Super 3 Viscometer instrument operated using Thermocline Windows control and analysis software version 1.2 was used (Newport Scientific, Sydney, Australia). The RVA profile is generally composed of five primary and two secondary parameters of the pasting curve: PKV, HPV, CPV, PeT, PaT, BDV, CS, and SBV.
The grain protein content in rice flour was determined by the Kjeldahl method using a Kjeltec 2300 Autoanalyser (Foss AB, Sweden). A nitrogen conversion factor of 6.25 was used to calculate the grain protein content35.
The taste value of the rice was measured by a SATAKE rice taste analyzer36. Each rice sample was washed with water 3 times in a cup with filtered water. Then, water was added to the stewed cup at a rice:water ratio of 1:1.4. After the rice was cooked and cooled, 7 ± 1 g of milled rice was weighed out, pressed with a molding tool into a microcake shape, and then put into a rice taste tester to determine the rice taste value.
The protein content of the brown rice was determined according to NY/T3-1982, and the fatty acid content of the brown rice was determined according to GB/T5510-2017. The fatty acid content was expressed as the potassium hydroxide mass (mg) needed to neutralize the free fatty acids in a 100 g sample.
PCR-based genotyping
Plant genomic DNA was extracted from the fresh leaves of heading-stage plants using a previously described CTAB protocol23. The molecular markers for genotyping Wx, GS3, GS9, GW5, GW7 and GW8 are listed in Table 1. PCR amplification was performed on a Bio-Rad C1000 Touch Thermal Cycler (USA), and the protocol for PCR amplification with the appropriate parameters was performed as described previously. The PCR products were resolved on a 3.0% agarose gel in 1 × TBE buffer23.
Statistical analysis
Statistical analysis was performed with independent samples using Student’s t-test and analysis of variance (ANOVA) and report the least Significant differences (LSD). The data are represented as means ± standard deviations (means ± SDs).
Conclusion
Rice quality is one of the main targets of rice breeding and is a complex trait involving grain appearance, milling, cooking, eating and nutritional quality. For a long time, rice breeding has experienced imbalances among yield, quality, and disease and lodging resistance. This study showed that the indica rice variety YNSM has excellent quality, with a relatively high brown rice rate, milled rice rate and head rice yield, low chalkiness and amylose content, and long gel consistency and that these properties were significantly correlated with the HPU, CPV, SB, and CS. This study also revealed that the grain size and food quality of YNSM may be related to gs3, gw7 and Wxb. This study also reported the quality characteristics of hybrid rice developed using YNSM as a restorer line. These results will facilitate the improvement of new rice varieties that achieve a balance of grain yield, resistance and quality.
Abbreviations
- AC:
-
Amylose content
- GL:
-
Grain length
- GW:
-
Grain width
- LWR:
-
Length-to-width ratio
- RVA:
-
Rapid viscosity analyzer
- GC:
-
Gel consistency
- HPV:
-
Hot paste viscosity
- CPV:
-
Cool paste viscosity
- PKV:
-
Peak viscosity
- SBV, PV minus PKV:
-
Setback viscosity
- CS, CPV minus HPV:
-
Consistency
- BDV, PKV minus HPV:
-
Breakdown viscosity
- PeT:
-
Peak time
- PaT:
-
Paste temperature
References
Butardo, V. M. Jr., Sreenivasulu, N. & Juliano, B. O. Improving rice grain quality: State-of-the-art and future prospects. Methods Mol. Biol. 19–55, 2019. https://doi.org/10.1007/978-1-4939-8914-0_2 (1892).
Birla, D. S. et al. Progress and challenges in improving the nutritional quality of rice (Oryza sativa L.). Crit. Rev. Food Sci. Nutr. 57, 2455–2481. https://doi.org/10.1080/10408398.2015.1084992 (2017).
Peng, B. et al. OsAAP6 functions as an important regulator of grain protein content and nutritional quality in rice. Nat. Commun. 5, 4847. https://doi.org/10.1038/ncomms5847 (2014).
Hori, K., Suzuki, K., Iijima, K. & Ebana, K. Variation in cooking and eating quality traits in Japanese rice germplasm accessions. Breed. Sci. 66, 309–318. https://doi.org/10.1270/jsbbs.66.309 (2016).
Zhang, L. et al. Grain size selection using novel functional markers targeting 14 genes in rice. Rice 13, 63. https://doi.org/10.1186/s12284-020-00427-y (2020).
Li, N., Xu, R., Duan, P. & Li, Y. Control of grain size in rice. Plant Reprod. 31, 237–251. https://doi.org/10.1007/s00497-018-0333-6 (2018).
Fan, C., Yu, S., Wang, C. & Xing, Y. A causal C-A mutation in the second exon of GS3 highly associated with rice grain length and validated as a functional marker. Theor. Appl. Genet. 118, 465–472. https://doi.org/10.1007/s00122-008-0913-1 (2009).
Mao, H. et al. Linking differential domain functions of the GS3 protein to natural variation of grain size in rice. Proc. Natl. Acad. Sci. U.S.A. 107, 19579–19584. https://doi.org/10.1073/pnas.1014419107 (2010).
Zhao, D. S. et al. GS9 acts as a transcriptional activator to regulate rice grain shape and appearance quality. Nat. Commun. 9, 1240. https://doi.org/10.1038/s41467-018-03616-y (2018).
Weng, J. et al. Isolation and initial characterization of GW5, a major QTL associated with rice grain width and weight. Cell Res. 18, 1199–1209. https://doi.org/10.1038/cr.2008.307 (2008).
Wang, Y. et al. Copy number variation at the GL7 locus contributes to grain size diversity in rice. Nat. Genet. 47, 944–948. https://doi.org/10.1038/ng.3346 (2015).
Wang, S. et al. Control of grain size, shape and quality by OsSPL16 in rice. Nat Genetics 44, 950–954. https://doi.org/10.1038/ng.2327 (2012).
Wang, S. et al. The OsSPL16-GW7 regulatory module determines grain shape and simultaneously improves rice yield and grain quality. Nat. Genet. 47, 949–954. https://doi.org/10.1038/ng.3352 (2015).
Li, N. et al. Amylose content and molecular-order stability synergistically affect the digestion rate of indica rice starches. Int. J. Biol. Macromol. 144, 373–379. https://doi.org/10.1016/j.ijbiomac.2019.12.095 (2020).
Fasahat, P., Rahman, S. & Ratnam, W. Genetic controls on starch amylose content in wheat and rice grains. J. Genet. 93, 279–292. https://doi.org/10.1007/s12041-014-0325-8 (2014).
Li, Q. F. et al. Down-Regulation of SSSII-2 Gene Expression Results in Novel Low-Amylose Rice with Soft, Transparent Grains. J. Agric. Food Chem. 66, 9750–9760. https://doi.org/10.1021/acs.jafc.8b02913 (2018).
Zhou, H. et al. Critical roles of soluble starch synthase SSIIIa and granule-bound starch synthase Waxy in synthesizing resistant starch in rice. Proc. Natl. Acad. Sci. U.S.A. 113, 12844–12849 (2016).
Teng, B. et al. Detection of allelic variation at the Wx locus with single-segment substitutionlines in rice (Oryza sativa L.). Mol. Breed. 30, 583–595 (2012).
Su, Y. et al. Map-based cloning proves qGC-6, a major QTL for gel consistency of japonica/indica cross, responds by Waxy in rice (Oryza sativa L). Theor. Appl. Genet. 123, 859–867. https://doi.org/10.1007/s00122-011-1632-6 (2011).
Nakamura, S., Katsura, J., Kato, K. & Ohtsubo, K. Development of formulae for estimating amylose content and resistant starch content based on the pasting properties measured by RVA of Japonica polished rice and starch. Biosci. Biotechnol. Biochem. 80, 329–340. https://doi.org/10.1080/09168451.2015.1088373 (2016).
Tong, C. et al. Genetic diversity of amylose content and RVA pasting parameters in 20 rice accessions grown in Hainan, China. Food Chem. 161, 239–245. https://doi.org/10.1016/j.foodchem.2014.04.011 (2014).
Yan, C. J. et al. Genetic analysis of starch paste viscosity parameters in glutinous rice (Oryza sativa L.). Theor. Appl. Genet. 122, 63–76. https://doi.org/10.1007/s00122-010-1423-5 (2011).
Wang, S. et al. Distribution of bacterial blight resistance genes in the main cultivars and application of Xa23 in rice breeding. Front. Plant Sci. 11, 555228. https://doi.org/10.3389/fpls.2020.555228 (2020).
Lu, Z. et al. Genetic analysis and gene identification of high-quality rice‘Yuenongsimiao’with broad spectrum resistance against rice blast. J. Plant Genet. Resour. 21, 827–833 (2020).
Lu, Z. et al. Influencing factors and genetic analysis of lodging resistance of high-quality rice ‘Yuenongsimiao’. J. Plant Genet. Resour. https://doi.org/10.13430/j.cnki.jpgr.20200918003 (2020).
Zhou, D. et al. Pedigree-based analysis of derivation of genome segments of an elite rice reveals key regions during its breeding. Plant Biotechnol. J. 14, 638–648. https://doi.org/10.1111/pbi.12409 (2016).
Wang, X. et al. A lipid transfer protein, OsLTPL36, is essential for seed development and seed quality in rice. Plant Sci. 239, 200–208. https://doi.org/10.1016/j.plantsci.2015.07.016 (2015).
Dyer, J. M. & Mullen, R. T. Engineering plant oils as high-value industrial feedstocks for biorefining: The need for underpinning cell biology research. Physiol. Plant. 132, 11–22. https://doi.org/10.1111/j.1399-3054.2007.01021.x (2008).
Fan, C. et al. GS3, a major QTL for grain length and weight and minor QTL for grain width and thickness in rice, encodes a putative transmembrane protein. Theor. Appl. Genet. 112, 1164–1171. https://doi.org/10.1007/s00122-006-0218-1 (2006).
Chun, A., Lee, H. J., Hamaker, B. R. & Janaswamy, S. Effects of ripening temperature on starch structure and gelatinization, pasting, and cooking properties in rice (Oryza sativa). J. Agric. Food Chem. 63, 3085–3093. https://doi.org/10.1021/jf504870p (2015).
Zhou, B., Dolan, M., Sakai, H. & Wang, G. L. The genomic dynamics and evolutionary mechanism of the Pi2/9 locus in rice. Mol. Plant-Microbe Interact. 20, 63–71. https://doi.org/10.1094/MPMI-20-0063 (2007).
Lu, Z. et al. Genetic analysis and gene identification of high-quality rice ‘Yuenong Simiao’with broad spectrum resistance against rice blast. J. Plant Genet. Resour. 21, 827–833 (2020).
Zhang, Y. D. et al. Haplotypes of qGL3 and their roles in grain size regulation with GS3 alleles in rice. Genet. Mol. Res. https://doi.org/10.4238/gmr.15017587 (2016).
Yang, Y. et al. Genetic improvement of panicle-erectness japonica rice toward both yield and eating and cooking quality. Mol. Breed. 40, 51. https://doi.org/10.1007/s11032-020-01127-7 (2020).
Yang, Y. et al. Identification of quantitative trait loci responsible for rice grain protein content using chromosome segment substitution lines and fine mapping of qPC-1 in rice (Oryza sativa L.). Mol. Breed. 35, 130. https://doi.org/10.1007/s11032-015-0328-z (2015).
Tang, Z. Q. et al. Study on Relationship between Palatability Value and Other Quality Traits in Rice. North Rice (2010).
Li, Z. et al. Development and utilization of SSR primer in Wx gene of rice (Oryza sativa L.), Southwest China. J. Agric. Sci. 22, 231–235 (2009).
Liu, L., He, Y., Yan, H., Li, J. & Zhao, Q. Development of functional markers and multiple PCR detection of ricegrain width genes GW5 and GW8. Mol. Plant Breed. 17, 4280–4288 (2019).
Acknowledgements
This work was supported by the following funders: the Rice Innovation Team Project of Modern Agricultural Industrial Technology System in Guangdong Province (2020KJ105); the Quality Rice and Seed Major Scientific Research Joint Project of the Department of Agriculture and Rural Affairs, Guangdong Province ([2019]73); the Natural Science Foundation of Guangdong Province (2018A0303130172); the Discipline Team Building Project of the 13th Five-Year Plan of GDAAS (201635TD); Guangzhou science and technology plan project (201804010467); and the National Natural Science Foundation of China (31471175).
Author information
Authors and Affiliations
Contributions
Z.L. wrote the paper; X.H. and Z.L. designed the study; Z.L. and Z.F. performed most of the experiments and analyzed the data; W.L., D.L., X.W., S.W. and J.X. participated in the data preparation and analysis; and X.H. oversaw and improved the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lu, Z., Fang, Z., Liu, W. et al. Grain quality characteristics analysis and application on breeding of Yuenongsimiao, a high-yielding and disease-resistant rice variety. Sci Rep 13, 6335 (2023). https://doi.org/10.1038/s41598-022-21030-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-21030-9
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.