Abstract
Previous studies examining the role of the histone deacetylase Hdac3 within myeloid cells demonstrated that Hdac3 promotes M2 activation and tissue healing in inflammatory conditions. Since myeloid lineage cells are required for proper bone formation and regeneration, in this study we examined the functions of Hdac3 during bone healing. Conditional deletion of Hdac3 within myeloid progenitors accelerates healing of cortical bone defects. Moreover, reduced osteoclast numbers within the defect site are correlated with Hdac3 suppression. Ex vivo osteoclastogenesis assays further demonstrate that Hdac3 deficiency limits osteoclastogenesis, the number of nuclei per cell and bone resorption, suggesting a defect in cell fusion. High throughput RNA sequencing identified the transmembrane protein Pmepa1 as a differentially expressed gene within osteoclast progenitor cells. Knockdown of Pmepa1 partially restores defects in osteoclastogenesis induced by Hdac3 deficiency. These results show that Hdac3 is required for optimal bone healing and osteoclast fusion, potentially via its regulation of Pmepa1 expression.
Similar content being viewed by others
Introduction
An estimated 5–10% of bone fractures will exhibit healing complications including delayed healing or non-union1,2. While many factors can limit healing, enhanced inflammation and altered immune responses are significant contributors3,4. Macrophages participate in inflammation and are recruited to sites of infection and damage where they play a central role in the tissue regeneration process. Macrophages contribute to both the initial injury response as well as to subsequent tissue healing. Although a spectrum of polarization exists, macrophages can be classified into two primary polarization states. Inflammatory macrophages (e.g., M1) are primarily involved during the inflammatory phase, whereas alternative macrophages (e.g., M2) are effectors of the tissue healing phase5. M1 respond to inflammatory mediators such as IFNγ, LPS and TNFα, but M2 are stimulated by IL4 and IL136. These immune cells contribute to bone healing following injury7,8,9,10. Both circulating macrophages and tissue resident macrophages within bone (e.g., osteomacs) contribute to bone healing11,12, and osteomacs aid in bone growth during development and promote bone formation8,9,11,13,14,15,16,17,18,19,20,21,22.
Osteoclasts are large, multinucleated cells that resorb bone. They arise from the fusion of myeloid progenitor cells, a process which is facilitated by two cytokines, M-CSF and RANKL. Although polarized macrophages do not give rise to osteoclasts following in vitro exposure to RANKL23,24, polarized macrophages can influence the differentiation osteoclast lineage cells. M2 are reported to suppress osteoclastogenesis via production of IL4 and IL10 and consequent repression of NFATc125,26,27,28,29,30. Conversely, inflammatory mediators produced by M1, including IFNγ, TNFα and IL6, promote either the proliferation of pre-osteoclast cells or their differentiation to the osteoclast lineage31,32,33,34,35,36,37,38. As these cells aid in healing and repair, macrophage polarization state may influence bone healing responses.
Histone deacetylases (Hdacs) are enzymes that remove acetyl groups from lysine side chains of histones and other proteins. Hdacs regulate numerous cellular and mitochondrial processes including gene transcription, DNA repair, protein stability, cytoskeletal dynamics, and signaling pathways to affect both development and aging39. Hdac3 is expressed by a wide variety of cell types and is predominantly localized to the nucleus. Germline deletion of Hdac3 results in embryonic lethality40, but conditional deletion revealed a central role for Hdac3 in facilitating endochondral and intramembranous ossification and promoting long bone growth41,42. Mesenchymal lineage Hdac3 is essential to maintain bone mass during aging and functions to suppress bone marrow adiposity43,44,45,46. Conditional deletion of Hdac3 within the monocyte/macrophage lineage (LysM-Cre) promotes alternative macrophage marker gene expression and hyper-responsiveness to IL447. Moreover, Hdac3 induces half of the inflammatory gene program of macrophages following stimulation with LPS, and Hdac3 deficiency enhances collagen deposition within atherosclerotic lesions and stabilizes plaques48,49. Importantly, Hdac inhibitors act as anti-inflammatory agents in mice and humans50,51,52; thus, Hdac3 inhibition may temper inflammatory responses of M1 within the periphery. Given the importance of macrophages during tissue healing, we examined the requirement of myeloid lineage-specific Hdac3 to bone healing and osteoclastogenesis. We find that 12-week-old female Hdac3-cKOLysM mice demonstrate more bone volume within cortical bone defects and limited osteoclast fusion.
Results
Deletion of Hdac3 in macrophage lineage cells confers alternate macrophage activation
Prior work demonstrated that conditional deletion of Hdac3 within bone marrow cells expressing LysM-Cre enhanced alternative macrophage activation47. We isolated bone marrow macrophages from male or female Hdac3 cKOLysM mice or their sex-matched WT control Cre+ littermates and confirmed elevated levels of the alternative macrophage marker Arg1 (Fig. 1A,B). BMMs exposed to the pan Hdac inhibitor SAHA for 24 h also exhibited increased expression of alternative macrophage activation markers and hyper-responsiveness to IL4 (Fig. 1C–E), known hallmarks of alternative macrophage activation. These data confirm prior observations made by Mullican et al.47.
Hdac3 deficiency or Hdac inhibition promotes expression of alternative macrophage activation markers. (A–D) Bone marrow macrophages were isolated and cultured for 24 h. Expression of Hdac3 (A) and Arg1 (B) was evaluated by qPCR. *p < 0.05. (C–E) Bone marrow macrophages were isolated in the presence of the Hdac inhibitor SAHA or vehicle control. Cells were then exposed to IL4 for 24 h in the presence of SAHA. Expression of Arg1 (C), Retnla (D) and Chi313 (E) was evaluated by qPCR. *p < 0.05.
Conditional deletion of Hdac3 enhances cortical bone healing
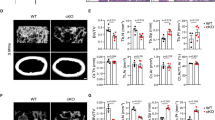
Macrophage activation can influence how tissues heal and regenerate. Hdac3-deficiency within the macrophage lineage limits atherosclerotic plaque lesions, but has no effect on spinal cord injury healing49,53. To determine if macrophage lineage Hdac3 expression affected bone healing, we induced single cortex defects in male and female Hdac3 cKOLysM 12-week-old mice or their Cre+ control littermates and assessed bone healing 14 days after defect generation. Radiography confirmed generation and uniform size of each induced defect (Fig. 2E). Blinded analyses revealed that Hdac3 cKOLysM female mice showed an average 140% increase in bone volume and BV/TV within the defect site, but no changes were observed in males (Fig. 2A,B). Increased BV/TV within the defect site of females was confirmed via histomorphometry, with a greater amount of bone volume along the periosteal surface within the defect of Hdac3 cKOLysM mice (Fig. 2C). More bone volume within the defect was accompanied by a significant reduction in osteoclasts per defect volume (Fig. 2D,E,F), but no change in osteoblast number was observed (data not shown). We also assessed the baseline cortical and trabecular parameters of the distal femur of intact 12-week-old male and female Hdac3 cKOLysM mice compared to their control Cre+ littermates, but did not find a change in these parameters at this age (Table 1). We confirmed these results via histomorphometry, but observed diminished osteoclast numbers within the distal femur (Fig. 3A–C).
Monocyte/Macrophage lineage Hdac3 deficiency enhances bone healing. (A–D) Hdac3 cKOLysM mice and their Control Cre+ littermates were aged to 12 weeks. Single cortex bone defects were generated in the left femur and bone healing was assessed after 14 days. Micro computed-tomography was performed by blinded study staff and used to determine (A) bone volume and (B) BV/TV within the defect. Femurs were sectioned and stained with Goldner’s trichrome. Blinded study staff determined (C) BV/TV or (D) TRAP to define the number of osteoclasts within the defect site. *p < 0.05 (E) Radiographs showing cortical bone defects at day 0 and day 14. (F) TRAP staining at the defect site.
Hdac3 suppression limits osteoclast numbers. Hdac3 cKOLysM mice and their Control Cre+ littermates were aged to 12 weeks. Femurs were sectioned and stained for TRAP and counterstained with fast green (A) and blinded study staff determined BV/TV (B) and defined the number of osteoclasts (C) within the distal femur. *p < 0.05.
Hdac3 is required for optimal osteoclast fusion
We observed decreased osteoclast numbers within female Hdac3 cKOLysM mice in vivo and Hdac3 knockdown in bone marrow macrophages was shown to reduce osteoclastogenesis54, but the mechanism behind this observation was not explored. To address the effects of Hdac3 conditional deletion on commitment of progenitor cells to the osteoclast lineage, we performed ex vivo osteoclastogenesis assays. Deletion of Hdac3 resulted in decreased numbers of multinucleated osteoclasts on day 4 (Fig. 4A). We performed an osteoclastogenesis time course and found that while the number of mono- or multinucleated cells was not altered early, at the onset of osteoclast fusion (day 3) the percentage of Hdac3 cKOLysM multinucleated cells was suppressed by 14.7 percent (Fig. 4B,C), whereas no change in the total number or TUNEL positive cells was observed (data not shown). This suggested a potential defect in osteoclast fusion. We therefore determined the number of nuclei per cell and found that osteoclasts derived from Hdac3 cKOLysM mice had reduced numbers of nuclei per multinucleated cell (Fig. 4D). We also delineated the number of cells with a defined number of nuclei. Hdac3 deficiency enhanced the number of mononuclear cells (Fig. 4E). There was no difference in cells with 4–8 nuclei, but Hdac3 deficiency decreased the occurrence of cells with eight or more nuclei (Fig. 4E). Expression of early osteoclastogenic genes including Ctsk and RANK also decreased within Hdac3-depleted cells (Fig. 4F). Moreover, activation of signaling downstream of RANKL, including phosphorylation of p65 NF-κB and Mek/Erk, was diminished within Hdac3 cKOLysM osteoclasts (Fig. 4G). These results demonstrate that Hdac3 deficiency suppresses osteoclast fusion.
Monocyte/Macrophage lineage Hdac3 deficiency decreases ex vivo osteoclastogenesis. Bone marrow macrophages were collected from Hdac3 cKOLysM 6-week-old male or female mice or their control Cre+ littermates and used to generate osteoclasts ex vivo. Shown are cultures from female mice. TRAP staining was performed (A). An osteoclastogenesis time course was performed (B) and the percent multi-nucleated cells (C) number of nuclei per cell (D) and occurrence of cells with a defined number of nuclei was determined (E). *p < 0.05. (F) Hdac3 levels and those of early osteoclastogenic genes Ctsk and RANK were assessed by qPCR. *p < 0.05. (G) Western blotting was performed as shown.
To perform an unbiased assessment of the effects of Hdac3 deficiency during osteoclastogenesis, we performed high-throughput RNA sequencing. BMMs were collected from Hdac3 cKOLysM 6-week-old male mice or their control littermates and cultured in osteoclastogenesis assays with RANKL and M-CSF. On day 3, when the defect in cell fusion was first observed, we collected mRNA for high-throughput sequencing. Using a false discovery rate of p < 0.05, we identified 71 differentially expressed genes versus sex-matched control littermates that had a fold change of two or greater (Table 2). Functional annotation using DAVID showed that the top categories affected by Hdac3 deficiency included Membrane, Extracellular Region and Cell Adhesion, all potentially involved in osteoclast progenitor fusion (Table 3).
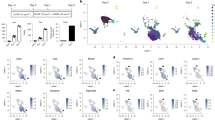
We next knocked down each of the 71 differentially expressed genes in wildtype BMMs using a siRNA-based approach to identify Hdac3 effector genes involved in osteoclast fusion (Fig. 5). Wildtype BMMs were collected from male 6 to 8-week old mice. On day 1, cells were transfected with an siRNA smart pool targeting one of the 71 differentially expressed genes or a non-targeting control siRNA. On day 4, cultures were TRAP and DAPI stained and the number of mono- and multinucleated cells was determined. Using this approach, siRNAs that targeted genes up regulated by Hdac3 deficiency and increasing the percentage of multinucleated TRAP+ cells were considered positive; likewise, siRNAs targeting genes that were down regulated by Hdac3 deficiency that also resulted in reduced numbers of TRAP+ multinucleated cells were also considered (Fig. 5A). Knockdown of 11 out of 71 differentially expressed genes changed the percentage of multinucleated cells (Fig. 5B). Of these 11 genes, two (Pmepa1 and Gdf15) fit our criteria (Fig. 5C). Knockdown of Gdf15 did not rescue the effects of Hdac3 deficiency (data not shown).
SiRNA-based screen to identify Hdac3 effector genes. A siRNA-based screen was performed as depicted in (A). Points where differentially expressed genes were excluded from the screen are noted with an X. Bone marrow macrophages were collected from wild type 6-week-old female mice. Cells were transfected with a siRNA targeting a differentially expressed gene or control siRNA on day 1 of osteoclast differentiation. Cells were then TRAP stained on day 4 and the targets that affected osteoclast multinucleation were identified (B). Genes that meet screening criteria outlined in (A) are highlighted (C).
Pmepa1 knock-down rescues Hdac3-dependent defects in osteoclastogenesis
To further explore the role of Pmepa1 as a downstream target of Hdac3, we first assessed expression of Pmepa1 during different phases of osteoclast differentiation in WT cells. BMMs were isolated from 6 to 8 week-old WT mice and placed in ex vivo osteoclastogenesis assays for the indicated days (Fig. 6). Pmepa1 transcripts increased along with those of Ctsk and RANK (Fig. 6A–C). Protein levels of Pmepa1 were increased during phases of osteoclast fusion on days 3 and 4 (Fig. 6D).
Pmepa1 expression increases during the fusion stage of osteoclastogenesis. Bone marrow macrophages were collected from WT 6-week-old male or female mice and cultured in the presence of RANKL and M-CSF for the indicated days. Shown are cultures from female mice. Expression levels of (A) Rank, (B) Ctsk and (C) Pmepa1 were evaluated by qPCR. *p < 0.05 (D) Western blotting was performed.
BMMs were then isolated from 6–8 week-old male or female Hdac3 cKOLysM mice and their sex-matched control littermates and placed in ex vivo culture with RANKL and M-CSF. On day 1, cells were transfected with siRNA smart pools targeting Pmepa1 or a non-targeting siRNA control. On day 4, cells were collected for analyses. Whereas deletion of Hdac3 decreased the number of multinucleated cells, knockdown of Pmepa1 in Hdac3 deficient cells significantly restored the size and number of multinucleated TRAP+ cells (Fig. 7A,B,C). Knockdown of Pmpea1 in control BMMs also enhanced the area and number of TRAP+ multinucleated cells (Fig. 7A,B,C). Real-time quantitative PCR verified suppression of Hdac3 levels in Hdac3 cKOLysM cultures and knockdown of Pmepa1 (Fig. 7D,E). Western blotting also confirmed enhanced Pmepa1 levels in Hdac3 suppressed cells (Fig. 7H). Pmepa1 attenuates phosphorylation of Smad3; as expected, knockdown of Pmepa1 enhanced phospho-Smad3 levels (Fig. 7H). Bone marrow macrophages derived from Hdac3 cKOLysM or their littermate controls were also seeded onto bone slices in osteoclastogenic conditions. On day 1, cells were transfected with Pmepa1 siRNAs or non-targeting siRNAs. Cultures were maintained for two weeks and the amount of resorbed bone was visualized. Less resorption occurred in cultures derived from Hdac3 cKOLysM bone marrow macrophages, which was further enhanced by knockdown of Pmepa1 (Fig. 7F,G). Together these data demonstrate that Pmepa1 suppresses osteoclast fusion and is a downstream target of Hdac3-mediated repression.
Pmepa1 knock-down rescues Hdac3 deficiency. Bone marrow macrophages were collected from Hdac3 cKOLysM 6-week-old male or female mice or their control Cre+ littermates and used to generate osteoclasts ex vivo. Shown are cultures from female mice. On day 4 cells were TRAP stained (A) and (B) osteoclast area and (C) percent multinucleated cells was determined. *p < 0.05. Cells were also seeded onto bone slices. On day 14, resorption pits were stained with Toluidine blue (F) and quantified (G). *p < 0.05 Expression levels of (D) Hdac3 and (E) Pmepa1 were evaluated by qPCR. *p < 0.05 Western blotting confirmed knockdown of Pmepa1 (F). Pit formation assays were also performed (H).
Discussion
Hdac inhibitors reduce inflammatory responses and alter macrophage polarization, but they also negatively impact the survival of highly proliferative cells such as mesenchymal progenitors55,56,57,58. In this study, we determined the effects of depleting Hdac3 within LysM-Cre-expressing cells on basal bone mass, cortical bone healing and osteoclastogenesis. We found that 12-week-old Hdac3 cKOLysM mice had diminished osteoclast numbers, but no basal bone phenotype. In contrast, bone healing of Hdac3 cKOLysM females was enhanced in a cortical bone injury model. Hdac3 is important for maintenance of bone mass during aging and its expression declines with age57. Although 12-week-old Hdac3 cKOLysM do not have changes in basal bone mass, this does not preclude development of a phenotype with increased age. Deletion of Hdac3 enhanced cortical bone defect healing in female mice two weeks after injury, but not in male mice. This sex-specific effect may be due to loss of Hdac3/NCoR-dependent transcriptional repression of ERα target genes59,60,61,62. Since Hdac3 deficiency enhanced bone healing in a cortical defect model and Hdac inhibitors are protective against osteoarthritis63, future studies will be aimed at determining if deletion of Hdac3 within myeloid lineage cells likewise promotes healing of other skeletal tissues.
We also found that Hdac3 deficiency within myeloid progenitor cells diminished osteoclastogenesis, most likely due to decreased cell fusion. This is in line with previous reports demonstrating that knockdown of Hdac3 in bone marrow macrophages suppressed osteoclast numbers54. We used an siRNA-based screen to identify downstream targets of Hdac3 during osteoclastogenesis. This screen was performed using WT bone marrow macrophages and also identified other regulators of osteoclastogenesis; however, since we did not verify effective knock-down of all targets within this large siRNA-based screen, false-negative results are an inherent limitation. In general, Hdac3 functions to repress gene expression, but also has demonstrated coactivator functions64. For this reason we chose to target all genes that were differentially regulated by Hdac3 deficiency using an siRNA-based approach in WT bone marrow macrophages. Using this tactic, we identified Pmepa1, a transmembrane protein and inhibitor of Tgfβ signaling65.
Pmepa1 is induced by p38 MAPK signaling in response to RANKL, but expression is downregulated after 72 h in osteoclastogenesis assays66. Pmepa1 localizes to intracellular organelle membranes, and to cathepsin-K positive vesicle membranes in osteoclast progenitor cells66. Xu et al. showed that knockdown of Pmepa1 suppressed osteoclast activity and that Pmepa1 is associated with the actin ring of osteoclasts during adjuvant-induced bone resorption67. In our study, we find that knockdown of Pmepa1 after 24 h of RANKL exposure enhances osteoclast fusion, but represses osteoclast activity. In future studies, the mechanism by which Pmepa1 controls osteoclast fusion will be explored. Determining if suppression of Pmepa1 also aids in coupling to bone formation, similar to other models where osteoclastogenesis is enhanced but bone resorption is suppressed, is also of interest68,69.
Another limitation of this study is that we cannot separate the functions of Hdac3 in monocytes/macrophages from those required for osteoclastogenesis and osteoclast activity. Thus, the functions of Hdac3 within the committed osteoclasts will need to be explored with alternative Cre-drivers (e.g., TRAP-Cre, Ctsk-Cre) in future studies. Although our analyses suggests that osteoclast fusion is suppressed by Hdac3 deficiency, the exact mechanism by which Hdac3 limits bone healing is unclear in this model. Hdac3 depletion within the myeloid lineage may likewise have different effects on stages of bone healing (e.g., initial inflammatory response, later bone remodeling). Although osteoblast number was unaffected, this does not preclude an effect on osteoblast activity.
Together our data demonstrate that Hdac3 deficiency in LysM-Cre-expressing cells enhances bone healing and suppresses osteoclast fusion. Moreover, Pmepa1 may be a downstream target of Hdac3 controlling osteoclast fusion and activity.
Methods
Generation of Hdac3-LysM mice
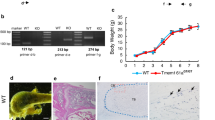
Hdac3fl/fl mice40 were crossed with mice expressing Cre-recombinase under the control of the LysM promoter70 to delete Hdac3 within LysM-expressing cells. Mice were genotyped for Cre as previously described46, or the Hdac3 floxed allele using the following primers: A: 5′-CCACTGGCTTCTCCTAAGTTC -3′, B: 5′- CCCAGGTTAGCTTTGAACTCT-3′ and C: 5′-TTTCCGTATTTGTGGAAGGA-3′. Conditional knockout animals from these crossings are referred to as Hdac3 cKOLysM mice and are on the C57Bl/6 background. Cre+ control littermates from crossings were used as controls as appropriate. Animals were housed in an accredited facility under a 12-h light/dark cycle and provided water and food ad libitum. All animal research was conducted according to guidelines provided by the National Institute of Health and the Institute of Laboratory Animal Resources, National Research Council. The Mayo Clinic Institutional Animal Care and Use Committee approved all animal studies.
Cortical bone defect generation
Single-cortex, fully stabilized defects were made in the mid-diaphysis of femurs as previously described71. Briefly, Cre+ male (n = 6) and female (n = 6) or Hdac3 cKOLysM male (n = 5) and female (n = 7) mice were given 0.09 mg/kg buprenorphine perioperatively. Mice were then anesthetized with isoflurane and prepared for aseptic surgery. A small incision was made in the skin overlying the left hamstring and the femoral bone shaft was exposed by blunt dissection of the underlying muscle. A 0.7 mm diameter steel burr drill bit (#19007-07, Fine Science Tools, Foster City, CA) and an electric drill was used to induce a single-cortex defect in the mid-diaphysis of the femur. Defects were immediately irrigated with 1 ml sterile saline followed by incision closure. Defect healing was monitored via radiography 14 days after surgery. Mice were sacrificed by carbon dioxide inhalation at postoperative day 14.
Micro-computed tomography
Bone architecture and mineralization were evaluated by ex vivo micro-CT. Femurs from 12-week-old male or female Hdac3 cKOLysM mice (n = 7 males, n = 5 females) and their sex-matched Cre+ littermates (n = 9 males, n = 10 females) were isolated and fixed in 10% neutral buffered formalin for 24 h. Femurs were then stored in 70% ethanol prior to scanning at 70 kV, 221 ms with a 10.5 μm voxel size using a Scanco Viva40 micro-CT. For cortical bone analyses, a region of interest was defined at 10% of total femur length beginning at the femoral midpoint; defining the outer cortical shell and running a midshaft analysis with 260-threshold air filling correction analyzed samples. For trabecular measurements, a region of interest was defined at 10% of total femur length starting immediately proximal to the growth plate. Bone volume within cortical bone defects was performed as previously described71. Briefly, a region of interest at the defect site was identified and a defined total volume of 0.5 mm3 was assessed. Samples were analyzed using a 220-threshold air filling correction. Bone volume, BV/TV, bone mineral density, trabecular number (mm−1), trabecular thickness (mm), and trabecular separation (mm) were computed using the manufacturer’s software.
Osteoclast cell culture, transfection, and treatment
Hind limbs were dissected from male or female 6 to 8-week-old WT, Cre+, Hdac3 cKOLysM or Hdac3fl/fl mice. Bone marrow macrophages were isolated as previously described72. Briefly, cells were flushed from the marrow cavity, pelleted and suspended in RBC Lysis Buffer (#00-4333-57, Invitrogen, Carlsbad, CA). Cells were pelleted and suspended in culture medium (phenol red-free alpha MEM, 10% FBS and 1% antibiotic/antimycotic) supplemented with 35 ng/mL M-CSF (#416-ML, R&D, Minneapolis, MN) and cultured overnight. Non-adherent cells were isolated and seeded at a density of 0.8 × 106 cells/mL in 24-well plates (day 0) in culture medium supplemented with 35 ng/mL M-CSF, and 60 ng/mL RANKL (#315-11, Prepro Tech, Rocky Hill, NJ). Cultures were fed on day 3 with culture medium plus 35 ng/ml M-CSF, and 60 ng/mL RANKL. Cells were treated as described within the text and figures. For knockdown experiments, a custom siRNA ON-TARGET plus plate was designed (Dharmacon, Lafayette, CO) to screen differentially regulated genes of interest. SiRNA smart pools targeting Pmepa1 and Gdf15 (L-040445-00-0005 or L-043512-01-0005) or a control siRNA (D-001810-10-05) were also purchased from Dharmacon (Lafayette, CO,). Cells were transfected (day 1) with each siRNA using Lipofectamine RNAiMax at a 1:1 ratio (#13778075, Invitrogen) and fed with culture medium supplemented with 35 ng/ml M-CSF, and 60 ng/mL RANKL after five hours. Each value shown is determined in triplicate and repeated three times. Shown is the average.
Bone resorption assays
Osteoclast precursors were plated on bone slices (NC1309388, Fisher Scientific) at a seeding density of 130,000 cells per well in culture medium supplemented with 35 ng/mL M-CSF and 60 ng/mL RANKL. Medium was replaced every 3–4 days. On Day 14, cells were lysed in 10% bleach washed 3 times with water. Pits on bone slices were stained with toluidine blue as previously described73. Resorption assays were performed in triplicate and repeated three times. Images were analyzed using ImageJ.
TRAP staining
TRAP staining was performed as previously described73. Briefly, cells were fixed on cover glass with 10% neutral buffered formalin for 10 min and then washed 3 times with phosphate-buffered saline (PBS)73. Fixed cells were TRAP stained using the Acid Phosphatase, Leukocyte (TRAP) Kit (#387A-1KT, Sigma-Aldrich) and mounted to slides using Vectashield with DAPI (#H-1200, Vector Laboratories, Burlingame, CA)73. For osteoclastogenesis experiments, three cover glasses were used per experimental condition. For each cover glass, three fields were imaged using a 10× objective73. Images were digitally photographed using the Zeiss AxioVert A1 and Zen software package. Osteoclasts were defined as TRAP+ cells with 3 or more nuclei73. The number of mononuclear cells and osteoclasts, as well as osteoclast area and nuclei per osteoclast within each image were quantified using ImageJ or Photoshop software73. The percent multinucleated cells is the number of osteoclasts divided by the number of mononuclear cells within each field. The average number of nuclei per cell was evaluated by counting the number of DAPI stained nuclei per TRAP+ cell and dividing by the total number of cells. The number of TRAP+ cells with defined numbers of nuclei was also determined. Each experiment was repeated using cells derived from male or female mice independently, each repeated three times. Shown is the average.
TUNEL and DAPI staining
TUNEL staining was performed using the Roche, In Situ Cell Death Detection Kit (Fluorescein) according to the manufacturer’s specifications (#11684795910, Sigma-Aldrich). DAPI staining was used as previously described to measure cell numbers73. To determine the percentage of TUNEL positive cells, the number of TUNEL-positive cells was divided by the number of DAPI positive cells in multiple regions.
RNA extraction and qPCR
Total RNA was extracted from primary osteoclasts using TRIzol (#15596026, Invitrogen) and chloroform, and 1 μg of RNA was reverse transcribed using the iScript Reverse Transcription Supermix for RT-qPCR kit (#1708840, Bio-Rad, Hercules, CA). The resulting cDNAs were used to assay gene expression via real-time PCR using the following gene-specific primers: Chi3l3, (5′-GTACCCTGGGTCTCGAGGAA-3′, 5′-CCTTGGAATGTCTTTCTCCACAG-3′), Ctsk (5′- TCCGAAAAGAGCCTAGCGAA-3′, 5′- AGAGATTTCATCCACCTTGCTGT-3′), Hdac3 (5′-CCCGCATCGAGAATCAGAAC-3′, 5′-TCAAAGATTGTCTGGCGGATCT-3′), Pmepa1 (5′-ATGGAGATCACGGAGCTGGAGT-3′, 5′-GGCTGACAGCTTGTAGTGGC-3′), Rank (5′- TAAAGTCTGTGATGCAGGCAAG-3′, 5′-CCGTATCCTTGTTGAGCTGC-3′), Retnla (5′-AACTGCCTGTGCTTACTCGT-3′, 5′-CAAGAAGCAGGGTAAATGGGC-3′) Tubulin (5′-TGCTCATCAGCAAGATCAGAG-3′, 5′-GCATTATAGGGCTCCACCACAG-3′), and Ywhaz (5′-GCCCTAAATGGTCTGTCACC-3′, 5′-GCTTTGGGTGTGACTTAGCC-3′). Fold changes in gene expression for each sample were calculated using the 2−ΔΔCq method relative to control following normalization of gene-specific Cq values to Tubulin or Ywhaz Cq values as noted74. Each value shown is determined in triplicate and repeated three times. Shown is the average.
Western blotting
Western blotting was performed as previously described41. Briefly, cell lysates were collected in a buffered SDS solution (0.1% glycerol, 0.01% SDS, 0.1 m Tris, pH 6.8) on ice. Total protein concentrations were obtained with the Bio-Rad DC assay (#5000112, Bio-Rad)41. Proteins (30 μg) were then resolved by SDS-PAGE and transferred to a polyvinylidene difluoride membrane41. Western blotting was performed with antibodies (1:2000 dilution) for anti-phospho-p65 NF-κB (#ab86299, Abcam, Cambridge, MA), anti-phospho-Mek1/2 (# 9121, Cell Signaling Technologies), anti-phospho-Erk1/2 (# 9101, Cell Signaling Technologies), anti-Mek1/2 (# 4694 Cells Signaling Technologies), anti-Erk1/2 (# 4370, Cell Signaling Technologies), anti-phospho-Ser423/Ser425-Smad3 (#ab52903, AbCam,), anti-Smad3 (#ab28379, AbCam), anti-Pmepa1 (#PA5-75980, Thermo Fisher Scientific, Waltham, MA), anti-Actin (#A2228, Sigma-Aldrich) and anti-Tubulin (E7, Developmental Studies Hybridoma Bank, Iowa City, IA) and corresponding secondary antibodies conjugated to horseradish peroxidase (HRP) (#7074S or #7076S, Cell Signaling Technology,)41. Antibody binding was detected with the Supersignal West Femto Chemiluminescent Substrate (#34096, Pierce Biotechnology, Rockford, IL)41. Each experiment was repeated using cells derived from male and female mice at least three times, and data from a representative experiment are shown.
Unbiased high throughput RNA sequencing
Bone marrow macrophages were isolated from Hdac3 cKOLysM male (n = 3) mice or their control littermates (n = 3) and cultured overnight with M-CSF. Non-adherent cells were then placed in osteoclastogenic conditions with RANKL and M-CSF as described above. On day 3, prior to cell fusion, cells were harvested for total RNA. Whole-transcriptome sequencing and bioinformatics analyses were performed as described previously42. The TruSeq RNA sample Prep Kit v2 (Illumina, San Diego, CA) was used to prepare RNA samples, and then samples were analyzed using the Illumina HiSeq 2000 with the TruSeq SBS sequencing kit version 3 and HCS v2.0.12 data collection software42. Following data collection, sequence data were processed using MAPRSeq (v.1.2.1) and the bioinformatics workflow (TopHat 2.0.6, HTSeq, and edgeR 2.6.2), where expression data were normalized using the reads per kilobase per million (RPKM) method42. Paired sample analysis was used to identify differentially expressed genes (n = 71) that had a two-fold or greater change with a false discover rate (FDR) less than 0.0542. One set of paired samples was excluded from the analysis due to insufficient reduction in Hdac3 RPKM values of the Hdac3 cKOLysM sample as compared to control. Gene functional annotation analysis was performed using DAVID Bioinformatics Resources 6.7 (http://david.abcc.ncifcrf.gov).
Bone histomorphometry
Femurs containing cortical bone defects (n = 5 WT, n = 6 Hdac3 cKOLysM) were decalcified, paraffin embedded and sectioned to a thickness of 7 µm. Sections were TRAP stained using the Acid Phosphatase, Leukocyte (TRAP) Kit (#387A-1KT, Sigma-Aldrich) or stained with Goldner’s trichrome stain (#1004850001, Sigma-Aldrich) according to the manufacture’s specifications73. The number of TRAP positive cells within the defect site or distal femur was determined, and standard bone histomorphometric measurements were collected as previously described73.
Statistical analysis
Data obtained are the mean ± standard deviation. p values were determined with the Student's t test when only one experimental comparison was made. For assessment of significance with greater than two conditions, a one-way analysis of variance was performed. p < 0.05 was considered statistically significant unless otherwise noted75. Statistical analyses were performed using Graphpad Prism 7 software.
Abbreviations
- IL:
-
Interleukin
- Hdac3:
-
Histone deacetylase 3
- cKO:
-
Conditional knockout
- Pmepa1:
-
Prostrate transmembrane protein, androgen induced 1
- Tgfβ:
-
Transforming growth factor beta
- IFNγ:
-
Interferon gamma
- LPS:
-
l Lipopolysaccharides
- TNFα:
-
Tumor necrosis factor alpha
- M-CSF:
-
Macrophage-colony stimulating factor
- RANKL:
-
Receptor activator of nuclear factor kappa-B ligand
- NFATc1:
-
Nuclear factor of activated T cells
- Arg1:
-
Arginase 1
- BMM:
-
Bone marrow macrophage
- SAHA:
-
Suberoylanilide hydroxamic acid
- BV/TV:
-
Bone volume per total volume
- TUNEL:
-
Terminal deoxynucleotidyl transferase dUTP nick end labeling
- DAVID:
-
Database for Annotation, Visualization and Integrated Discovery
- TRAP:
-
Tartrate-resistant acid phosphatase
- DAPI:
-
4′,6-Diamidino-2-phenylindole
- Prss46:
-
Serine protease 46
- Gdf15:
-
Growth and differentiation factor 15
- PCR:
-
Polymerase chain reaction
- Smad:
-
Sma and mothers against decapentaplegic
- MAPK:
-
Mitogen activated protein kinase
- Ctsk:
-
Cathepsin-K
- CT:
-
Computed tomography
- RBC:
-
Red blood cell
- MEM:
-
Minimal essential medium
- FBS:
-
Fetal bovine serum
- PBS:
-
Phosphate buffered saline
- RNA:
-
Ribonucleic acid
- DNA:
-
Deoxynucleic acid
- Chi3l3:
-
Chitinase-like 3
- Retnla:
-
Resistin like alpah
- Ywhaz:
-
Tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, zeta polypeptide
- SDS–PAGE:
-
Sodium dodecyl sulfate–polyacrylamide gel electrophoresis
- HRP:
-
Horseradish peroxidase
- RPKM:
-
Reads per kilobase per million
- FDR:
-
False discovery rate
- RANK:
-
Receptor activator of NF-Κb
- Ctsk:
-
Cathepsin K
- Mek1/2:
-
Mitogen-activated protein kinase kinases 1/2
- Erk1/2:
-
Extracellular regulated kinases 1/2
References
Einhorn, T. A. & Gerstenfeld, L. C. Fracture healing: Mechanisms and interventions. Nat. Rev. Rheumatol. 11, 45–54. https://doi.org/10.1038/nrrheum.2014.164 (2015).
Tzioupis, C. & Giannoudis, P. V. Prevalence of long-bone non-unions. Injury 38(Suppl 2), S3-9. https://doi.org/10.1016/s0020-1383(07)80003-9 (2007).
Franceschi, C. et al. Inflamm-aging. An evolutionary perspective on immunosenescence. Ann. N. Y. Acad. Sci. 908, 244–254. https://doi.org/10.1111/j.1749-6632.2000.tb06651.x (2000).
Clark, D., Nakamura, M., Miclau, T. & Marcucio, R. Effects of aging on fracture healing. Curr. Osteoporos. Rep. 15, 601–608. https://doi.org/10.1007/s11914-017-0413-9 (2017).
Lampiasi, N., Russo, R. & Zito, F. The alternative faces of macrophage generate osteoclasts. Biomed. Res. Int. 2016, 9089610. https://doi.org/10.1155/2016/9089610 (2016).
Martinez, F. O. & Gordon, S. The M1 and M2 paradigm of macrophage activation: Time for reassessment. F1000prime reports 6, 13. https://doi.org/10.12703/P6-13 (2014).
Baht, G. S., Vi, L. & Alman, B. A. The role of the immune cells in fracture healing. Curr. Osteoporos. Rep. 16, 138–145. https://doi.org/10.1007/s11914-018-0423-2 (2018).
Vi, L. et al. Macrophage cells secrete factors including LRP1 that orchestrate the rejuvenation of bone repair in mice. Nat. Commun. 9, 5191. https://doi.org/10.1038/s41467-018-07666-0 (2018).
Vi, L. et al. Macrophages promote osteoblastic differentiation in-vivo: Implications in fracture repair and bone homeostasis. J. Bone Miner. Res. 30, 1090–1102. https://doi.org/10.1002/jbmr.2422 (2015).
Wynn, T. A. & Vannella, K. M. Macrophages in tissue repair, regeneration, and fibrosis. Immunity 44, 450–462. https://doi.org/10.1016/j.immuni.2016.02.015 (2016).
Alexander, K. A. et al. Osteal macrophages promote in vivo intramembranous bone healing in a mouse tibial injury model. J. Bone Miner. Res. 26, 1517–1532. https://doi.org/10.1002/jbmr.354 (2011).
Ono, T. & Takayanagi, H. Osteoimmunology in bone fracture healing. Curr. Osteoporos. Rep. 15, 367–375. https://doi.org/10.1007/s11914-017-0381-0 (2017).
Chang, M. K. et al. Osteal tissue macrophages are intercalated throughout human and mouse bone lining tissues and regulate osteoblast function in vitro and in vivo. J. Immunol. 181, 1232–1244. https://doi.org/10.4049/jimmunol.181.2.1232 (2008).
Cho, S. W. et al. Osteal macrophages support physiologic skeletal remodeling and anabolic actions of parathyroid hormone in bone. Proc. Natl. Acad. Sci. U. S. A. 111, 1545–1550. https://doi.org/10.1073/pnas.1315153111 (2014).
Fernandes, T. J. et al. Cord blood-derived macrophage-lineage cells rapidly stimulate osteoblastic maturation in mesenchymal stem cells in a glycoprotein-130 dependent manner. PLoS ONE 8, e73266. https://doi.org/10.1371/journal.pone.0073266 (2013).
Guihard, P. et al. Oncostatin m, an inflammatory cytokine produced by macrophages, supports intramembranous bone healing in a mouse model of tibia injury. Am. J. Pathol. 185, 765–775. https://doi.org/10.1016/j.ajpath.2014.11.008 (2015).
Guihard, P. et al. Induction of osteogenesis in mesenchymal stem cells by activated monocytes/macrophages depends on oncostatin M signaling. Stem Cells 30, 762–772. https://doi.org/10.1002/stem.1040 (2012).
Nicolaidou, V. et al. Monocytes induce STAT3 activation in human mesenchymal stem cells to promote osteoblast formation. PLoS ONE 7, e39871. https://doi.org/10.1371/journal.pone.0039871 (2012).
Pirraco, R. P., Reis, R. L. & Marques, A. P. Effect of monocytes/macrophages on the early osteogenic differentiation of hBMSCs. J. Tissue Eng. Regen. Med. 7, 392–400. https://doi.org/10.1002/term.535 (2013).
Raggatt, L. J. et al. Fracture healing via periosteal callus formation requires macrophages for both initiation and progression of early endochondral ossification. Am. J. Pathol. 184, 3192–3204. https://doi.org/10.1016/j.ajpath.2014.08.017 (2014).
Pettit, A. R., Chang, M. K., Hume, D. A. & Raggatt, L. J. Osteal macrophages: A new twist on coupling during bone dynamics. Bone 43, 976–982. https://doi.org/10.1016/j.bone.2008.08.128 (2008).
Wu, A. C., Raggatt, L. J., Alexander, K. A. & Pettit, A. R. Unraveling macrophage contributions to bone repair. Bonekey Rep. 2, 373. https://doi.org/10.1038/bonekey.2013.107 (2013).
Jeganathan, S., Fiorino, C., Naik, U., Sun, H. S. & Harrison, R. E. Modulation of osteoclastogenesis with macrophage M1- and M2-inducing stimuli. PLoS ONE 9, e104498. https://doi.org/10.1371/journal.pone.0104498 (2014).
Nakanishi-Matsui, M., Yano, S., Matsumoto, N. & Futai, M. Lipopolysaccharide induces multinuclear cell from RAW264.7 line with increased phagocytosis activity. Biochem. Biophys. Res. Commun. 425, 144–149. https://doi.org/10.1016/j.bbrc.2012.07.050 (2012).
Evans, K. E. & Fox, S. W. Interleukin-10 inhibits osteoclastogenesis by reducing NFATc1 expression and preventing its translocation to the nucleus. BMC Cell Biol. 8, 4. https://doi.org/10.1186/1471-2121-8-4 (2007).
Liu, D., Yao, S. & Wise, G. E. Effect of interleukin-10 on gene expression of osteoclastogenic regulatory molecules in the rat dental follicle. Eur. J. Oral. Sci. 114, 42–49. https://doi.org/10.1111/j.1600-0722.2006.00283.x (2006).
Abu-Amer, Y. IL-4 abrogates osteoclastogenesis through STAT6-dependent inhibition of NF-kappaB. J. Clin. Investig. 107, 1375–1385. https://doi.org/10.1172/JCI10530 (2001).
Moreno, J. L., Kaczmarek, M., Keegan, A. D. & Tondravi, M. IL-4 suppresses osteoclast development and mature osteoclast function by a STAT6-dependent mechanism: Irreversible inhibition of the differentiation program activated by RANKL. Blood 102, 1078–1086. https://doi.org/10.1182/blood-2002-11-3437 (2003).
Stein, N. C. et al. Interleukin-4 and interleukin-13 stimulate the osteoclast inhibitor osteoprotegerin by human endothelial cells through the STAT6 pathway. J. Bone Miner. Res. 23, 750–758. https://doi.org/10.1359/jbmr.080203 (2008).
Wei, S., Wang, M. W., Teitelbaum, S. L. & Ross, F. P. Interleukin-4 reversibly inhibits osteoclastogenesis via inhibition of NF-kappa B and mitogen-activated protein kinase signaling. J. Biol. Chem. 277, 6622–6630. https://doi.org/10.1074/jbc.M104957200 (2002).
Dimitroulas, T., Nikas, S. N., Trontzas, P. & Kitas, G. D. Biologic therapies and systemic bone loss in rheumatoid arthritis. Autoimmun. Rev. 12, 958–966. https://doi.org/10.1016/j.autrev.2013.03.015 (2013).
Duplomb, L. et al. Interleukin-6 inhibits receptor activator of nuclear factor kappaB ligand-induced osteoclastogenesis by diverting cells into the macrophage lineage: Key role of Serine727 phosphorylation of signal transducer and activator of transcription 3. Endocrinology 149, 3688–3697. https://doi.org/10.1210/en.2007-1719 (2008).
Kim, K., Lee, S. H., Ha Kim, J., Choi, Y. & Kim, N. NFATc1 induces osteoclast fusion via up-regulation of Atp6v0d2 and the dendritic cell-specific transmembrane protein (DC-STAMP). Mol. Endocrinol. (Baltimore, Md) 22, 176–185. https://doi.org/10.1210/me.2007-0237 (2008).
Lam, J. et al. TNF-alpha induces osteoclastogenesis by direct stimulation of macrophages exposed to permissive levels of RANK ligand. J. Clin. Investig. 106, 1481–1488. https://doi.org/10.1172/JCI11176 (2000).
Tamura, T. et al. Soluble interleukin-6 receptor triggers osteoclast formation by interleukin 6. Proc. Natl. Acad. Sci. U.S.A. 90, 11924–11928. https://doi.org/10.1073/pnas.90.24.11924 (1993).
Vogel, D. Y. et al. Macrophages migrate in an activation-dependent manner to chemokines involved in neuroinflammation. J. Neuroinflamm. 11, 23. https://doi.org/10.1186/1742-2094-11-23 (2014).
Yao, Z. et al. Tumor necrosis factor-alpha increases circulating osteoclast precursor numbers by promoting their proliferation and differentiation in the bone marrow through up-regulation of c-Fms expression. J. Biol. Chem. 281, 11846–11855. https://doi.org/10.1074/jbc.M512624200 (2006).
Yokota, K. et al. Combination of tumor necrosis factor alpha and interleukin-6 induces mouse osteoclast-like cells with bone resorption activity both in vitro and in vivo. Arthritis Rheumatol. 66, 121–129. https://doi.org/10.1002/art.38218 (2014).
Bradley, E. W., Carpio, L. R., van Wijnen, A. J., McGee-Lawrence, M. E. & Westendorf, J. J. Histone deacetylases in bone development and skeletal disorders. Physiol. Rev. 95, 1359–1381. https://doi.org/10.1152/physrev.00004.2015 (2015).
Bhaskara, S. et al. Deletion of histone deacetylase 3 reveals critical roles in S phase progression and DNA damage control. Mol. Cell 30, 61–72. https://doi.org/10.1016/j.molcel.2008.02.030 (2008).
Bradley, E. W., Carpio, L. R. & Westendorf, J. J. Histone deacetylase 3 suppression increases PH domain and leucine-rich repeat phosphatase (Phlpp)1 expression in chondrocytes to suppress Akt signaling and matrix secretion. J. Biol. Chem. 288, 9572–9582. https://doi.org/10.1074/jbc.M112.423723 (2013).
Carpio, L. R. et al. Histone deacetylase 3 supports endochondral bone formation by controlling cytokine signaling and matrix remodeling. Sci. Signal 9, ra79. https://doi.org/10.1126/scisignal.aaf3273 (2016).
McGee-Lawrence, M. E. et al. Histone deacetylase 3 is required for maintenance of bone mass during aging. Bone 52, 296–307. https://doi.org/10.1016/j.bone.2012.10.015 (2012).
McGee-Lawrence, M. E. et al. Hdac3 deficiency increases marrow adiposity and induces lipid storage and glucocorticoid metabolism in osteochondroprogenitor cells. J. Bone Miner. Res. 31, 116–128. https://doi.org/10.1002/jbmr.2602 (2016).
McGee-Lawrence, M. E. et al. Loss of Hdac3 in osteoprogenitors increases bone expression of osteoprotegerin, improving systemic insulin sensitivity. J. Cell. Physiol. https://doi.org/10.1002/jcp.26148 (2017).
Razidlo, D. F. et al. Histone deacetylase 3 depletion in osteo/chondroprogenitor cells decreases bone density and increases marrow fat. PloS one 5, e11492 (2010).
Mullican, S. E. et al. Histone deacetylase 3 is an epigenomic brake in macrophage alternative activation. Genes Dev. 25, 2480–2488. https://doi.org/10.1101/gad.175950.111 (2011).
Chen, X. et al. Requirement for the histone deacetylase Hdac3 for the inflammatory gene expression program in macrophages. Proc. Natl. Acad. Sci. U.S.A. 109, E2865-2874. https://doi.org/10.1073/pnas.1121131109 (2012).
Hoeksema, M. A. et al. Targeting macrophage Histone deacetylase 3 stabilizes atherosclerotic lesions. EMBO Mol. Med. 6, 1124–1132. https://doi.org/10.15252/emmm.201404170 (2014).
Adcock, I. M. Histone deacetylase inhibitors as novel anti-inflammatory agents. Curr. Opin. Investig. Drugs 7, 966–973 (2006).
Adcock, I. M. HDAC inhibitors as anti-inflammatory agents. Br. J. Pharmacol. 150, 829–831. https://doi.org/10.1038/sj.bjp.0707166 (2007).
Halili, M. A., Andrews, M. R., Sweet, M. J. & Fairlie, D. P. Histone deacetylase inhibitors in inflammatory disease. Curr. Top. Med. Chem. 9, 309–319. https://doi.org/10.2174/156802609788085250 (2009).
Sanchez, S. et al. HDAC3 inhibition promotes alternative activation of macrophages but does not affect functional recovery after spinal cord injury. Exp. Neurobiol. 27, 437–452. https://doi.org/10.5607/en.2018.27.5.437 (2018).
Pham, L. et al. HDAC3 and HDAC7 have opposite effects on osteoclast differentiation. J. Biol. Chem. 286, 12056–12065. https://doi.org/10.1074/jbc.M110.216853 (2011).
Grabiec, A. M., Tak, P. P. & Reedquist, K. A. Function of histone deacetylase inhibitors in inflammation. Crit. Rev. Immunol. 31, 233–263 (2011).
Lee, S. et al. Histone deacetylase inhibitors decrease proliferation potential and multilineage differentiation capability of human mesenchymal stem cells. Cell Prolif. 42, 711–720. https://doi.org/10.1111/j.1365-2184.2009.00633.x (2009).
McGee-Lawrence, M. E. et al. Histone deacetylase 3 is required for maintenance of bone mass during aging. Bone 52, 296–307. https://doi.org/10.1016/j.bone.2012.10.015 (2013).
Ocker, M. & Schneider-Stock, R. Histone deacetylase inhibitors: Signalling towards p21cip1/waf1. Int. J. Biochem. Cell Biol. 39, 1367–1374. https://doi.org/10.1016/j.biocel.2007.03.001 (2007).
Guenther, M. G., Barak, O. & Lazar, M. A. The SMRT and N-CoR corepressors are activating cofactors for histone deacetylase 3. Mol. Cell. Biol. 21, 6091–6101. https://doi.org/10.1128/mcb.21.18.6091-6101.2001 (2001).
Heinzel, T. et al. A complex containing N-CoR, mSin3 and histone deacetylase mediates transcriptional repression. Nature 387, 43–48. https://doi.org/10.1038/387043a0 (1997).
Legare, S. & Basik, M. Minireview: The link between ERalpha corepressors and histone deacetylases in tamoxifen resistance in breast cancer. Mol. Endocrinol. (Baltimore, Md) 30, 965–976. https://doi.org/10.1210/me.2016-1072 (2016).
Li, J. et al. Both corepressor proteins SMRT and N-CoR exist in large protein complexes containing HDAC3. EMBO J. 19, 4342–4350. https://doi.org/10.1093/emboj/19.16.4342 (2000).
Carpio, L. R. & Westendorf, J. J. Histone deacetylases in cartilage homeostasis and osteoarthritis. Curr. Rheumatol. Rep. 18, 52. https://doi.org/10.1007/s11926-016-0602-z (2016).
Emmett, M. J. et al. Histone deacetylase 3 prepares brown adipose tissue for acute thermogenic challenge. Nature 546, 544–548. https://doi.org/10.1038/nature22819 (2017).
Itoh, S. & Itoh, F. TMEPAI family: Involvement in regulation of multiple signalling pathways. J. Biochem. 164, 195–204. https://doi.org/10.1093/jb/mvy059 (2018).
Funakubo, N. et al. Pmepa1 induced by RANKL-p38 MAPK pathway has a novel role in osteoclastogenesis. J. Cell Physiol. 233, 3105–3118. https://doi.org/10.1002/jcp.26147 (2018).
Xu, X. et al. Prostate transmembrane protein androgen induced 1 is induced by activation of osteoclasts and regulates bone resorption. FASEB J. 33, 4365–4375. https://doi.org/10.1096/fj.201801573R (2019).
Boyce, B. F., Yoneda, T., Lowe, C., Soriano, P. & Mundy, G. R. Requirement of pp60c-src expression for osteoclasts to form ruffled borders and resorb bone in mice. J. Clin. Investig. 90, 1622–1627. https://doi.org/10.1172/JCI116032 (1992).
Mattson, A. M. et al. Deficiency in the phosphatase PHLPP1 suppresses osteoclast-mediated bone resorption and enhances bone formation in mice. J. Biol. Chem. https://doi.org/10.1074/jbc.RA119.007660 (2019).
Clausen, B. E., Burkhardt, C., Reith, W., Renkawitz, R. & Forster, I. Conditional gene targeting in macrophages and granulocytes using LysMcre mice. Transgenic Res. 8, 265–277 (1999).
McGee-Lawrence, M. E. et al. Sclerostin deficient mice rapidly heal bone defects by activating beta-catenin and increasing intramembranous ossification. Biochem. Biophys. Res. Commun. 441, 886–890. https://doi.org/10.1016/j.bbrc.2013.10.155 (2013).
Bradley, E. W. & Oursler, M. J. Osteoclast culture and resorption assays. Methods Mol. Biol. 455, 19–35. https://doi.org/10.1007/978-1-59745-104-8_2 (2008).
Mattson, A. M. et al. Deficiency in the phosphatase PHLPP1 suppresses osteoclast-mediated bone resorption and enhances bone formation in mice. J. Biol. Chem. 294, 11772–11784. https://doi.org/10.1074/jbc.RA119.007660 (2019).
Castillejo Becerra, C. M. et al. DNA methylation and FoxO3a regulate PHLPP1 expression in chondrocytes. J. Cell Biochem. 119, 7470–7478. https://doi.org/10.1002/jcb.27056 (2018).
Gagnier, J. J. & Morgenstern, H. Misconceptions, misuses, and misinterpretations of P values and significance testing. J. Bone Joint Surg. Am. 99, 1598–1603. https://doi.org/10.2106/JBJS.16.01314 (2017).
Acknowledgements
We thank Xiaodong Li for technical assistance and the Mayo Clinic Bone Histomorphometry Core.
Funding
This work was made possible by research grants from the National Institutes of Health (AR065397, AR072634, AR068103, AR065402, and AR067129), the Mayo Clinic Center for Biomedical Discovery, the Mayo Clinic Foundation, and the University of Minnesota Stem Cell Institute and Board of Regents. These contents are solely the responsibility of the authors and do not necessarily represent the official views of the NIH.
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Molstad, D.H.H., Zars, E., Norton, A. et al. Hdac3 deletion in myeloid progenitor cells enhances bone healing in females and limits osteoclast fusion via Pmepa1. Sci Rep 10, 21804 (2020). https://doi.org/10.1038/s41598-020-78364-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-78364-5
This article is cited by
-
Histone deacetylase 3 (HDAC3) as an important epigenetic regulator of kidney diseases
Journal of Molecular Medicine (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.