Abstract
The frequency of errors upon decoding of messenger RNA by the bacterial ribosome is low, with one misreading event per 1 × 104 codons. In the universal genetic code, the AUN codon box specifies two amino acids, isoleucine and methionine. In bacteria and archaea, decoding specificity of the AUA and AUG codons relies on the wobble avoidance strategy that requires modification of C34 in the anticodon loop of isoleucine transfer RNAIleCAU (tRNAIleCAU). Bacterial tRNAIleCAU with 2-lysylcytidine (lysidine) at the wobble position deciphers AUA while avoiding AUG. Here we report cryo-electron microscopy structures of the Escherichia coli 70S ribosome complexed with elongation factor thermo unstable (EF-Tu) and isoleucine-tRNAIleLAU in the process of decoding AUA and AUG. Lysidine in tRNAIleLAU excludes AUG by promoting the formation of an unusual Hoogsteen purine–pyrimidine nucleobase geometry at the third position of the codon, weakening the interactions with the mRNA and destabilizing the EF-Tu ternary complex. Our findings elucidate the molecular mechanism by which tRNAIleLAU specifically decodes AUA over AUG.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The atomic coordinates were deposited in the RCSB Protein Data Bank (PDB) under accession codes 8G7P (structure I; 70S ribosome with EF-Tu•GDPCP•A/T-Ile-tRNAIleLAU bound to the cognate AUA codon), 8G7Q (structure II; 70S ribosome with EF-Tu•GDPCP•A/T-Ile-tRNAIleLAU bound to the near-cognate AUG codon), 8G7R (structure III; 70S ribosome with A-site tRNAIleLAU bound to the cognate AUA codon) and 8G7S (structure IV; 70S ribosome with the P-site tRNAIleLAU bound to the cognate AUA codon). The cryo-EM maps have been deposited in the Electron Microscopy Data Bank (EMDB) under accession codes EMD-29819 (structure I; 70S ribosome with EF-Tu•GDPCP•A/T-Ile-tRNAIleLAU bound to the cognate AUA codon), EMD-29820 (structure II; 70S ribosome with EF-Tu•GDPCP•A/T-Ile-tRNAIleLAU bound to the near-cognate AUG codon), EMD-29821 (structure III; 70S ribosome with A-site tRNAIleLAU bound to the cognate AUA codon) and EMD-29822 (structure IV; 70S ribosome with the P-site tRNAIleLAU bound to the cognate AUA codon). The unaligned multi-frame cryo-EM micrographs have been deposited in the Electron Microscopy Public Image Archive (EMPIAR)55 with the accession codes EMPIAR-11582 (structure I), EMPIAR-11583 (structure II), EMPIAR-11584 (structure III) and EMPIAR-11585 (structure IV).
References
Loveland, A. B., Demo, G., Grigorieff, N. & Korostelev, A. A. Ensemble cryo-EM elucidates the mechanism of translation fidelity. Nature 546, 113–117 (2017).
Voorhees, R. M., Schmeing, T. M., Kelley, A. C. & Ramakrishnan, V. The mechanism for activation of GTP hydrolysis on the ribosome. Science 330, 835–838 (2010).
Schmeing, T. M. et al. The crystal structure of the ribosome bound to EF-Tu and aminoacyl-tRNA. Science 326, 688–694 (2009).
Ogle, J. M. & Ramakrishnan, V. Structural insights into translational fidelity. Annu. Rev. Biochem. 74, 129–177 (2005).
Ogle, J. M., Murphy, F. V. IV, Tarry, M. J. & Ramakrishnan, V. Selection of tRNA by the ribosome requires a transition from an open to a closed form. Cell 111, 721–732 (2002).
Ogle, J. M. et al. Recognition of cognate transfer RNA by the 30S ribosomal subunit. Science 292, 897–902 (2001).
Loveland, A. B., Demo, G. & Korostelev, A. A. Cryo-EM of elongating ribosome with EF-Tu• GTP elucidates tRNA proofreading. Nature 584, 640–645 (2020).
Rozov, A., Westhof, E., Yusupov, M. & Yusupova, G. The ribosome prohibits the G*U wobble geometry at the first position of the codon–anticodon helix. Nucl. Acids Res. 44, 6434–6441 (2016).
Rozov, A., Demeshkina, N., Westhof, E., Yusupov, M. & Yusupova, G. Structural insights into the translational infidelity mechanism. Nat. Commun. 6, 7251 (2015).
Demeshkina, N., Jenner, L., Westhof, E., Yusupov, M. & Yusupova, G. A new understanding of the decoding principle on the ribosome. Nature 484, 256–259 (2012).
Nilsson, E. M. & Alexander, R. W. Bacterial wobble modifications of NNA-decoding tRNAs. IUBMB Life 71, 1158–1166 (2019).
Klassen, R., Bruch, A. & Schaffrath, R. Independent suppression of ribosomal +1 frameshifts by different tRNA anticodon loop modifications. RNA Biol. 14, 1252–1259 (2017).
Powell, C. A. et al. TRMT5 mutations cause a defect in post-transcriptional modification of mitochondrial tRNA associated with multiple respiratory-chain deficiencies. Am. J. Hum. Genet. 97, 319–328 (2015).
Motorin, Y. & Helm, M. tRNA stabilization by modified nucleotides. Biochemistry 49, 4934–4944 (2010).
Voigts-Hoffmann, F. et al. A methyl group controls conformational equilibrium in human mitochondrial tRNA(Lys). J. Am. Chem. Soc. 129, 13382–13383 (2007).
Alexandrov, A. et al. Rapid tRNA decay can result from lack of nonessential modifications. Mol. Cell 21, 87–96 (2006).
Kadaba, S. et al. Nuclear surveillance and degradation of hypomodified initiator tRNAMet in S. cerevisiae. Genes Dev. 18, 1227–1240 (2004).
Madore, E. et al. Effect of modified nucleotides on Escherichia coli tRNAGlu structure and on its aminoacylation by glutamyl-tRNA synthetase. Predominant and distinct roles of the mnm5 and s2 modifications of U34. Eur. J. Biochem. 266, 1128–1135 (1999).
Helm, M., Giege, R. & Florentz, C. A Watson–Crick base-pair-disrupting methyl group (m1A9) is sufficient for cloverleaf folding of human mitochondrial tRNALys. Biochemistry 38, 13338–13346 (1999).
Kruger, M. K. & Sorensen, M. A. Aminoacylation of hypomodified tRNAGlu in vivo. J. Mol. Biol. 284, 609–620 (1998).
Sylvers, L. A., Rogers, K. C., Shimizu, M., Ohtsuka, E. & Soll, D. A 2-thiouridine derivative in tRNAGlu is a positive determinant for aminoacylation by Escherichia coli glutamyl-tRNA synthetase. Biochemistry 32, 3836–3841 (1993).
Rogers, M. J. et al. Selectivity and specificity in the recognition of tRNA by E coli glutaminyl-tRNA synthetase. Biochimie 75, 1083–1090 (1993).
Bjork, G. R., Wikstrom, P. M. & Bystrom, A. S. Prevention of translational frameshifting by the modified nucleoside 1-methylguanosine. Science 244, 986–989 (1989).
Weixlbaumer, A. et al. Mechanism for expanding the decoding capacity of transfer RNAs by modification of uridines. Nat. Struct. Mol. Biol. 14, 498–502 (2007).
Nasvall, S. J., Chen, P. & Bjork, G. R. The wobble hypothesis revisited: uridine-5-oxyacetic acid is critical for reading of G-ending codons. RNA 13, 2151–2164 (2007).
Nasvall, S. J., Chen, P. & Bjork, G. R. The modified wobble nucleoside uridine-5-oxyacetic acid in tRNAPro(cmo5UGG) promotes reading of all four proline codons in vivo. RNA 10, 1662–1673 (2004).
Murphy, F. V. T. & Ramakrishnan, V. Structure of a purine-purine wobble base pair in the decoding center of the ribosome. Nat. Struct. Mol. Biol. 11, 1251–1252 (2004).
Crick, F. H. Codon–anticodon pairing: the wobble hypothesis. J. Mol. Biol. 19, 548–555 (1966).
Parker, J. Errors and alternatives in reading the universal genetic code. Microbiol. Rev. 53, 273–298 (1989).
Soma, A. et al. An RNA-modifying enzyme that governs both the codon and amino acid specificities of isoleucine tRNA. Mol. Cell 12, 689–698 (2003).
Nakanishi, K. et al. Structural basis for translational fidelity ensured by transfer RNA lysidine synthetase. Nature 461, 1144–1148 (2009).
Muramatsu, T. et al. A novel lysine-substituted nucleoside in the first position of the anticodon of minor isoleucine tRNA from Escherichia coli. J. Biol. Chem. 263, 9261–9267 (1988).
Köhrer, C. et al. Identification and characterization of a tRNA decoding the rare AUA codon in Haloarcula marismortui. RNA 14, 117–126 (2008).
Ikeuchi, Y. et al. Agmatine-conjugated cytidine in a tRNA anticodon is essential for AUA decoding in archaea. Nat. Chem. Biol. 6, 277–282 (2010).
Mandal, D. et al. Agmatidine, a modified cytidine in the anticodon of archaeal tRNAIle, base pairs with adenosine but not with guanosine. Proc. Natl Acad. Sci. USA 107, 2872–2877 (2010).
Voorhees, R. M. et al. The structural basis for specific decoding of AUA by isoleucine tRNA on the ribosome. Nat. Struct. Mol. Biol. 20, 641–643 (2013).
Punjani, A. & Fleet, D. J. 3D variability analysis: resolving continuous flexibility and discrete heterogeneity from single particle cryo-EM. J. Struct. Biol. 213, 107702 (2021).
von Loeffelholz, O. et al. Focused classification and refinement in high-resolution cryo-EM structural analysis of ribosome complexes. Curr. Opin. Struct. Biol. 46, 140–148 (2017).
Serna, M. Hands on methods for high resolution cryo-electron microscopy structures of heterogeneous macromolecular complexes. Front. Mol. Biosci. 6, 33 (2019).
Rozov, A. et al. Importance of potassium ions for ribosome structure and function revealed by long-wavelength X-ray diffraction. Nat. Commun. 10, 2519 (2019).
Schmeing, T. M., Voorhees, R. M., Kelley, A. C. & Ramakrishnan, V. How mutations in tRNA distant from the anticodon affect the fidelity of decoding. Nat. Struct. Mol. Biol. 18, 432–436 (2011).
Kajander, T. et al. Buried charged surface in proteins. Structure 8, 1203–1214 (2000).
Dulic, M., Cvetesic, N., Perona, J. J. & Gruic-Sovulj, I. Partitioning of tRNA-dependent editing between pre- and post-transfer pathways in class I aminoacyl-tRNA synthetases. J. Biol. Chem. 285, 23799–23809 (2010).
Zhang, C., Yashiro, Y., Sakaguchi, Y., Suzuki, T. & Tomita, K. Substrate specificities of Escherichia coli ItaT that acetylates aminoacyl-tRNAs. Nucl. Acids Res. 48, 7532–7544 (2020).
Jünemann, R. et al. In vivo deuteration of transfer RNAs: overexpression and large-scale purification of deuterated specific tRNAs. Nucl. Acids Res. 24, 907–913 (1996).
Basu, R. S., Sherman, M. B. & Gagnon, M. G. Compact IF2 allows initiator tRNA accommodation into the P site and gates the ribosome to elongation. Nat. Commun. 13, 3388 (2022).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Watson, Z. L. et al. Structure of the bacterial ribosome at 2 Å resolution. eLife 9, e60482 (2020).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. 66, 486–501 (2010).
Leroy, E. C., Perry, T. N., Renault, T. T. & Innis, C. A. Tetracenomycin X sequesters peptidyl-tRNA during translation of QK motifs. Nat. Chem. Biol. 19, 1091–1096 (2023).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D. 75, 861–877 (2019).
Afonine, P. V. et al. New tools for the analysis and validation of cryo-EM maps and atomic models. Acta Crystallogr. D. 74, 814–840 (2018).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Iudin, A. et al. EMPIAR: the electron microscopy public image archive. Nucleic Acids Res. 51, D1503–D1511 (2023).
Cardone, G., Heymann, J. B. & Steven, A. C. One number does not fit all: mapping local variations in resolution in cryo-EM reconstructions. J. Struct. Biol. 184, 226–236 (2013).
Acknowledgements
We thank R. Basu for critical reading of the paper and useful suggestions. We thank the staff of the Sealy Center for Structural Biology and Molecular Biophysics cryo-electron microscopy facility at the University of Texas Medical Branch for advice and support. Special thanks to K.-Y. (Clem) Wong and J. Perkyns (University of Texas Medical Branch) for computational support and to the Sealy and Smith Foundation for supporting the Sealy Center for Structural Biology at the University of Texas Medical Branch. We are grateful to T. Suzuki and N. Akiyama for kindly sharing their unpublished cryo-EM data of the ribosome complexed with the lysidine-modified tRNAIleLAU. This work was supported by NIH grant R01GM136936 (to M.G.G.), the Welch Foundation grant H-2032-20230405 (to M.G.G.), startup funds from the University of Texas Medical Branch (to M.G.G.), Rising Science and Technology Acquisition and Retention Program award from the University of Texas system (to M.G.G.), NIH grant P41GM103311 (to the Resource for Biocomputing, Visualization, and Informatics at the University of California, San Francisco) for developing UCSF Chimera, and NIH grant R01GM129325 (to the Resource for Biocomputing, Visualization, and Informatics at the University of California, San Francisco) and the Office of Cyber Infrastructure and Computational Biology, National Institute of Allergy and Infectious Diseases for developing UCSF ChimeraX.
Author information
Authors and Affiliations
Contributions
M.Y.R. and M.G.G. designed the project. M.Y.R. purified ribosomes, tRNAIle, TilS, IleRS and EF-Tu, prepared the samples for structure determination, and collected the cryo-EM data. M.Y.R. and M.G.G. processed the cryo-EM data and built the molecular models. M.Y.R. made the figures. M.Y.R. and M.G.G. wrote the paper. Both authors reviewed, edited and approved the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks the anonymous reviewer(s) for their contribution to the peer review of this work. Peer review reports are available. Primary Handling Editor: Sara Osman, in collaboration with the Nature Structural & Molecular Biology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Cryo-EM data processing and particle classification workflow for structure I.
In this 70S ribosome complex, EF-Tu•GDPCP•Ile-tRNAIleLAU is bound to the cognate AUA codon in the A site. See Methods for details.
Extended Data Fig. 2 Cryo-EM data processing and particle classification workflow for structure II.
In this 70S ribosome complex, EF-Tu•GDPCP•Ile-tRNAIleLAU is bound to the near-cognate AUG codon in the A site in the presence of paromomycin. See Methods for details.
Extended Data Fig. 3 Local resolution estimation and Fourier Shell Correlation (FSC) validation.
Local resolution heat maps on slices of density from structures I (a, EF-Tu•GDPCP•Ile-tRNAIleLAU with A-site AUA), II (b, EF-Tu•GDPCP•Ile-tRNAIleLAU with A-site AUG), III (c, tRNAIleLAU with A-site AUA), and IV (d, tRNAIleLAU with P-site AUA) shown in the range of 2–6 Å resolution, calculated with cryoSPARC 3.3.2 implementation of BlocRes56. The gold-standard FSC curves of each half-map (red), using a ‘soft mask’ excluding solvent and model-map (green), are plotted across resolution. Map and model validation were performed in PHENIX 1.19.2 (ref. 53).
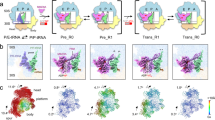
Extended Data Fig. 4 Cryo-EM density of the EF-Tu ternary complex in structures I (A-site AUA) and II (A-site AUG).
a, Well-defined density of EF-Tu, Ile-tRNAIleLAU and GDPCP in structure I upon decoding of the cognate AUA codon in the A site. b, In structure II with the near-cognate AUG codon, the presence of paromomycin stabilizes the anticodon loop in the decoding center, while the density for the other parts of Ile-tRNAIleLAU and EF-Tu is poorly resolved, suggesting unstable binding to the ribosome. The Coulomb potential density (shown as mesh and contoured at 4.0σ) of EF-Tu·GDPCP is magenta, that of the A/T-Ile-tRNAIleLAU is green, and that of the AUA- or AUG-mRNA is blue. The isoleucine attached to residue A76 of tRNAIleLAU and the lysidine 34 modification in the anticodon loop are indicated.
Extended Data Fig. 5 Base pair geometries in the codon-anticodon helix.
First (a), second (b), or third (c) position of the codon upon decoding of the cognate AUA (left, structure I) or the near-cognate AUG (right, structure II) codon. The gap that forms at the second and third positions in the complex with the AUG codon is indicated with a gray arrow. The hydrogen bonds between nucleotides are shown in Fig. 4. d, Potential non-favorable interactions between G6 in the usual syn-conformation and lysidine 34 during decoding of the near-cognate AUG codon. The curved dashed lines indicate repulsive forces between two donors of hydrogen bond.
Extended Data Fig. 6 Comparison of the Ile-tRNAIleLAU in the A/T state and bound to EF-Tu in structures I (cognate A-site AUA) and II (near-cognate A-site AUG), and tRNAIleLAU in the A/A state in structure III (cognate A-site AUA).
a, Structure alignment based on the 23S rRNA shows that the Ile-tRNAIleLAU has the exact same conformation in structures I and II. The conformation of the mRNA bases in the A site diverges between structures I and II at the second and third position of the codon (inset). b, Close-up view of the second and third positions of the A-site codon. Relative to A6 in the AUA codon (structure I), syn-G6 inclines by ~15° to avoid a collision between the exocyclic amine and the phosphate oxygen of the mRNA backbone in the AUG codon (structure II). Through π-π stacking interactions, the effects of the inclined conformation of syn-G6 propagate to the neighboring bases, weakening the codon-anticodon base pairs with Ile-tRNAIleLAU. c, Structure alignment based on 23S rRNA showing that AUA-mRNA has the same conformation in structures I and III, regardless of the presence (structure III) or the absence (structure I) of paromomycin in the decoding center, confirming that paromomycin has no obvious effect on the conformation of the decoding center.
Extended Data Fig. 7 Cryo-EM density of the codon-anticodon region.
a, Structure I with the cognate AUA codon in the A site. Note that nucleotide A6 at the third position of the A-site codon is in the usual anti-conformation. b, Structure II with the near-cognate AUG codon in the A site. In this structure, nucleotide G6 at the wobble position of the A-site codon adopts the unusual syn-conformation. c, Nucleotide A7 immediately downstream of the A-site codon also adopts the syn-conformation in structure II, which maximizes stacking with syn-G6. d, EM map of the decoding center in structure III with accommodated A-site tRNAIleLAU bound to the cognate AUA codon. The Coulomb potential density is contoured at 2.9σ. e-f, The amine of the lysidine side chain in structures I (with the A/T-Ile-tRNAIleLAU) and III (with the A/A-tRNAIleLAU) may interact with the 2’OH group of A7 immediately downstream of the A-site codon. The putative water 'W' is shown as a yellow sphere and the gray dashed lines depict hydrogen bonds.
Extended Data Fig. 8 Comparison of the EF-Tu•GDPCP•Ile-tRNAIleLAU ternary complex in structure I with that from a previous study (5UYM1).
The conformation of EF-Tu and tRNA bound in the A/T state is the same as in the 70S ribosome•EF-Tu•GDPCP•Phe-tRNAPhe complex upon decoding of the cognate phenylalanine codon (PDB 5UYM)1. The structures are aligned based on the sarcin-ricin loop (SRL) region in the 50S subunit. a, The conformation of the EF-Tu•GDPCP•Ile-tRNAIleLAU ternary complex in structure I is in the GTP-activated state as reported previously1. b, The catalytic His84 in the G-domain of EF-Tu is within interaction distance from A2662 in the SRL. c, The quality of the EM map for EF-Tu in structure I allows to unambiguously model amino acid side chains. The density of the P-loop region is shown with magenta mesh, and that of the ordered switch I region with orange mesh. d, The EM density (gray mesh) of the isoleucine residue attached to nucleotide A76 of Ile-tRNAIleLAU bound to EF-Tu in structure I. e, Nucleotide-binding pocket in the G-domain of EF-Tu. The EM density of the GDPCP nucleotide is shown with gray mesh. In panels c-e, the Coulomb potential density is contoured at 2.8σ.
Extended Data Fig. 9 Cryo-EM data processing and particle classification workflow for structure III.
In this 70S ribosome complex, tRNAIleLAU is bound to the cognate AUA codon in the A site in the presence of paromomycin. See Methods for details.
Extended Data Fig. 10 Cryo-EM data processing and particle classification workflow for structure IV.
In this 70S ribosome complex, tRNAIleLAU is bound to the cognate AUA codon in the P site. See Methods for details.
Supplementary information
Supplementary Information
Supplementary Figs. 1–4 and references.
Supplementary Data 1
Micrograph_Structure_I.
Supplementary Data 2
Micrograph_Structure_II.
Supplementary Data 3
Micrograph_Structure_III.
Supplementary Data 4
Micrograph_Structure_IV.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Rybak, M.Y., Gagnon, M.G. Structures of the ribosome bound to EF-Tu–isoleucine tRNA elucidate the mechanism of AUG avoidance. Nat Struct Mol Biol (2024). https://doi.org/10.1038/s41594-024-01236-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41594-024-01236-3