Abstract
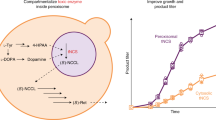
The protein complexes of the mitochondrial electron transport chain exist in isolation and in higher order assemblies termed supercomplexes (SCs) or respirasomes (SC I+III2+IV). The association of complexes I, III and IV into the respirasome is regulated by unknown mechanisms. Here, we designed a nanoluciferase complementation reporter for complex III and IV proximity to determine in vivo respirasome levels. In a chemical screen, we found that inhibitors of the de novo pyrimidine synthesis enzyme dihydroorotate dehydrogenase (DHODH) potently increased respirasome assembly and activity. By-passing DHODH inhibition via uridine supplementation decreases SC assembly by altering mitochondrial phospholipid composition, specifically elevated peroxisomal-derived ether phospholipids. Cell growth rates upon DHODH inhibition depend on ether lipid synthesis and SC assembly. These data reveal that nucleotide pools signal to peroxisomes to modulate synthesis and transport of ether phospholipids to mitochondria for SC assembly, which are necessary for optimal cell growth in conditions of nucleotide limitation.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Datasets generated during this current study are included in the published article as supplementary data or as publicly available datasets. Chemical screen data has been deposited in PubChem and proteomic data in PRIDE with accession no PXD023882. Source data are provided with this paper. Any additional data not included in this manuscript is available from the corresponding author upon request.
References
Mitchell, P. Coupling of phosphorylation to electron and hydrogen transfer by a chemi-osmotic type of mechanism. Nature 191, 144–148 (1961).
Schagger, H. & Pfeiffer, K. Supercomplexes in the respiratory chains of yeast and mammalian mitochondria. EMBO J. 19, 1777–1783 (2000).
Schagger, H. & Pfeiffer, K. The ratio of oxidative phosphorylation complexes I–V in bovine heart mitochondria and the composition of respiratory chain supercomplexes. J. Biol. Chem. 276, 37861–37867 (2001).
Letts, J. A., Fiedorczuk, K. & Sazanov, L. A. The architecture of respiratory supercomplexes. Nature 537, 644–648 (2016).
Gu, J. et al. The architecture of the mammalian respirasome. Nature 537, 639–643 (2016).
Sousa, J. S., Mills, D. J., Vonck, J. & Kuhlbrandt, W. Functional asymmetry and electron flow in the bovine respirasome. Elife 5, https://doi.org/10.7554/eLife.21290 (2016).
Wu, M., Gu, J., Guo, R., Huang, Y. & Yang, M. Structure of mammalian respiratory supercomplex I1III2IV1. Cell 167, 1598–1609.e10 (2016).
Guo, R., Zong, S., Wu, M., Gu, J. & Yang, M. Architecture of human mitochondrial respiratory megacomplex I2III2IV2. Cell 170, 1247–1257.e12 (2017).
Acin-Perez, R., Fernandez-Silva, P., Peleato, M. L., Perez-Martos, A. & Enriquez, J. A. Respiratory active mitochondrial supercomplexes. Mol. Cell 32, 529–539 (2008).
Shinzawa-Itoh, K. et al. Purification of active respiratory supercomplex from bovine heart mitochondria enables functional studies. J. Biol. Chem. 291, 4178–4184 (2016).
Calvo, E. et al. Functional role of respiratory supercomplexes in mice: SCAF1 relevance and segmentation of the Qpool. Sci. Adv. 6, eaba7509 (2020).
Garcia-Poyatos, C. et al. Scaf1 promotes respiratory supercomplexes and metabolic efficiency in zebrafish. EMBO Rep. 21, e50287 (2020).
Berndtsson, J. et al. Respiratory supercomplexes enhance electron transport by decreasing cytochrome c diffusion distance. EMBO Rep. 21, https://doi.org/10.15252/embr.202051015 (2020).
Chen, Y. C. et al. Identification of a protein mediating respiratory supercomplex stability. Cell Metab. 15, 348–360 (2012).
Hatle, K. M. et al. MCJ/DnaJC15, an endogenous mitochondrial repressor of the respiratory chain that controls metabolic alterations. Mol. Cell Biol. 33, 2302–2314 (2013).
Desmurs, M. et al. C11orf83, a mitochondrial cardiolipin-binding protein involved in bc1 complex assembly and supercomplex stabilization. Mol. Cell Biol. 35, 1139–1156 (2015).
Mitsopoulos, P. et al. Stomatin-like protein 2 is required for in vivo mitochondrial respiratory chain supercomplex formation and optimal cell function. Mol. Cell Biol. 35, 1838–1847 (2015).
Nagano, H. et al. p53-inducible DPYSL4 associates with mitochondrial supercomplexes and regulates energy metabolism in adipocytes and cancer cells. Proc. Natl Acad. Sci. USA 115, 8370–8375 (2018).
Ikeda, K., Shiba, S., Horie-Inoue, K., Shimokata, K. & Inoue, S. A stabilizing factor for mitochondrial respiratory supercomplex assembly regulates energy metabolism in muscle. Nat. Commun. 4, 2147 (2013).
Lapuente-Brun, E. et al. Supercomplex assembly determines electron flux in the mitochondrial electron transport chain. Science 340, 1567–1570 (2013).
Milenkovic, D., Blaza, J. N., Larsson, N. G. & Hirst, J. The enigma of the respiratory chain supercomplex. Cell Metab. 25, 765–776 (2017).
Cogliati, S. et al. Mechanism of super-assembly of respiratory complexes III and IV. Nature 539, 579–582 (2016).
Mourier, A., Matic, S., Ruzzenente, B., Larsson, N. G. & Milenkovic, D. The respiratory chain supercomplex organization is independent of COX7a2l isoforms. Cell Metab. 20, 1069–1075 (2014).
Perez-Perez, R. et al. COX7A2L is a mitochondrial complex III binding protein that stabilizes the III2+IV supercomplex without affecting respirasome formation. Cell Rep. 16, 2387–2398 (2016).
Pfeiffer, K. et al. Cardiolipin stabilizes respiratory chain supercomplexes. J. Biol. Chem. 278, 52873–52880 (2003).
Bottinger, L. et al. Phosphatidylethanolamine and cardiolipin differentially affect the stability of mitochondrial respiratory chain supercomplexes. J. Mol. Biol. 423, 677–686 (2012).
Das, S. et al. ATP citrate lyase improves mitochondrial function in skeletal muscle. Cell Metab. 21, 868–876 (2015).
Baker, C. D., Basu Ball, W., Pryce, E. N. & Gohil, V. M. Specific requirements of nonbilayer phospholipids in mitochondrial respiratory chain function and formation. Mol. Biol. Cell 27, 2161–2171 (2016).
Tasseva, G. et al. Phosphatidylethanolamine deficiency in mammalian mitochondria impairs oxidative phosphorylation and alters mitochondrial morphology. J. Biol. Chem. 288, 4158–4173 (2013).
Basu Ball, W., Neff, J. K. & Gohil, V. M. The role of nonbilayer phospholipids in mitochondrial structure and function. FEBS Lett. 592, 1273–1290 (2018).
Dixon, A. S. et al. NanoLuc complementation reporter optimized for accurate measurement of protein interactions in cells. ACS Chem. Biol. 11, 400–408 (2016).
Ohashi, K., Kiuchi, T., Shoji, K., Sampei, K. & Mizuno, K. Visualization of cofilin-actin and Ras-Raf interactions by bimolecular fluorescence complementation assays using a new pair of split Venus fragments. Biotechniques 52, 45–50 (2012).
Balsa, E. et al. ER and nutrient stress promote assembly of respiratory chain supercomplexes through the PERK-eIF2α axis. Mol. Cell 74, 877–890.e6 (2019).
Madak, J. T., Bankhead, A. 3rd, Cuthbertson, C. R., Showalter, H. D. & Neamati, N. Revisiting the role of dihydroorotate dehydrogenase as a therapeutic target for cancer. Pharmacol. Ther. 195, 111–131 (2019).
Schlame, M. & Ren, M. Barth syndrome, a human disorder of cardiolipin metabolism. FEBS Lett. 580, 5450–5455 (2006).
McKenzie, M., Lazarou, M., Thorburn, D. R. & Ryan, M. T. Mitochondrial respiratory chain supercomplexes are destabilized in Barth syndrome patients. J. Mol. Biol. 361, 462–469 (2006).
Dudek, J. et al. Cardiolipin deficiency affects respiratory chain function and organization in an induced pluripotent stem cell model of Barth syndrome. Stem Cell Res. 11, 806–819 (2013).
Breitkopf, S. B. et al. A relative quantitative positive/negative ion switching method for untargeted lipidomics via high resolution LC-MS/MS from any biological source. Metabolomics 13, https://doi.org/10.1007/s11306-016-1157-8 (2017).
Braverman, N. E. & Moser, A. B. Functions of plasmalogen lipids in health and disease. Biochim. Biophys. Acta 1822, 1442–1452 (2012).
Dean, J. M. & Lodhi, I. J. Structural and functional roles of ether lipids. Protein Cell 9, 196–206 (2018).
Honsho, M., Asaoku, S. & Fujiki, Y. Posttranslational regulation of fatty acyl-CoA reductase 1, Far1, controls ether glycerophospholipid synthesis. J. Biol. Chem. 285, 8537–8542 (2010).
Kimura, T. et al. Substantial decrease in plasmalogen in the heart associated with tafazzin deficiency. Biochemistry 57, 2162–2175 (2018).
Letts, J. A. & Sazanov, L. A. Clarifying the supercomplex: the higher-order organization of the mitochondrial electron transport chain. Nat. Struct. Mol. Biol. 24, 800–808 (2017).
Horibata, Y. et al. EPT1 (selenoprotein I) is critical for the neural development and maintenance of plasmalogen in humans. J. Lipid Res. 59, 1015–1026 (2018).
Kimura, T. et al. Plasmalogen loss caused by remodeling deficiency in mitochondria. Life Sci. Alliance 2, e201900348 (2019).
Greggio, C. et al. Enhanced respiratory chain supercomplex formation in response to exercise in human skeletal muscle. Cell Metab. 25, 301–311 (2017).
Hollinshead, K. E. R. et al. Respiratory supercomplexes promote mitochondrial efficiency and growth in severely hypoxic pancreatic cancer. Cell Rep. 33, 108231 (2020).
Ikeda, K. et al. Mitochondrial supercomplex assembly promotes breast and endometrial tumorigenesis by metabolic alterations and enhanced hypoxia tolerance. Nat. Commun. 10, 4108 (2019).
Jain, I. H. et al. Genetic screen for cell fitness in high or low oxygen highlights mitochondrial and lipid metabolism. Cell 181, 716–727.e11 (2020).
Benjamin, D. I. et al. Ether lipid generating enzyme AGPS alters the balance of structural and signaling lipids to fuel cancer pathogenicity. Proc. Natl Acad. Sci. USA 110, 14912–14917 (2013).
Zou, Y. et al. Plasticity of ether lipids promotes ferroptosis susceptibility and evasion. Nature 585, 603–608 (2020).
Rhee, H.-W. et al. Proteomic mapping of mitochondria in living cells via spatially restricted enzymatic tagging. Science 339, 1328–1331 (2013).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Shalem, O. et al. Genome-scale CRISPR–Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Spinazzi, M., Casarin, A., Pertegato, V., Salviati, L. & Angelini, C. Assessment of mitochondrial respiratory chain enzymatic activities on tissues and cultured cells. Nat. Protoc. 7, 1235–1246 (2012).
Yuan, M., Breitkopf, S. B., Yang, X. & Asara, J. M. A positive/negative ion-switching, targeted mass spectrometry-based metabolomics platform for bodily fluids, cells, and fresh and fixed tissue. Nat. Protoc. 7, 872–881 (2012).
Yuan, M. et al. Ex vivo and in vivo stable isotope labelling of central carbon metabolism and related pathways with analysis by LC–MS/MS. Nat. Protoc. 14, 313–330 (2019).
Honsho, M., Yagita, Y., Kinoshita, N. & Fujiki, Y. Isolation and characterization of mutant animal cell line defective in alkyl-dihydroxyacetonephosphate synthase: localization and transport of plasmalogens to post-Golgi compartments. Biochim. Biophys. Acta 1783, 1857–1865 (2008).
Meyers, R. M. et al. Computational correction of copy number effect improves specificity of CRISPR–Cas9 essentiality screens in cancer cells. Nat. Genet. 49, 1779–1784 (2017).
Dempster, J. M. et al. Extracting biological insights from the project achilles genome-scale CRISPR screens in cancer cell lines. Preprint at bioRxiv https://doi.org/10.1101/720243 (2019).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Acknowledgements
We thank members of the Puigserver laboratory for helpful discussions regarding this project. We would like to acknowledge the Nikon Imaging Center at Harvard Medical School for assistance with fluorescence microscopy, the ICCB-Longwood Screening Facility at Harvard Medical School for facilitating our screening efforts, J. M. Asara and M. Yuan at the Beth Israel Deaconess Medical Center Mass Spectrometry Core for providing metabolomics and nonpolar lipidomics profiling data and Promega for providing us with a nanoluciferase polyclonal antibody. This work was supported by NIH R01 DK089883-07 NIDDK, R01 DK081418-11 NIDDK and R01 GM121452 NIGMS to P.P., NIH F32 GM125243-01A1 NIGMS to C.F.B. and NIH F30 DE028206-01A1 NIDR to E.A.P., EMBO postdoctoral fellowship and MDA Development Grant to E.B., Human Frontier Science Program (LT-000033/2019-L) to P.L.M., and American Heart Associate postdoctoral fellowship to C.L.R.
Author information
Authors and Affiliations
Contributions
Conceptualization was performed by C.F.B., E.B. and P.P. Methodology development was carried out by C.F.B. Formal analysis was performed by C.F.B. Investigation was carried out by C.F.B., K.E.O., E.A.P., E.B., P.L.-M., C.L.R., C.L. and M.J. Resources were provided by M.J. and S.P.G. Writing of the original draft was completed by C.F.B. and P.P. Further editing was completed by C.F.B., P.P., K.E.O., E.A.P., E.B., P.L.-M., C.L.R., C.L. and M.J. Data presentation and visualization was done by C.F.B. Supervision of research and funding acquisition was carried out by C.F.B. and P.P.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Chemical Biology thanks Clay Semenkovich and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 A high-throughput chemical screen for respirasome assembly using an in vivo NanoBiT reporter.
a, BiFC SC reporter fluorescence co-localizes with MitoTracker Red. Scale bar, 25 µm (n = 1 experiment). b, NanoBiT SC reporter luminescence is proportional to cell number. Cells expressing UQCRB-LgBit and COX5B-SmBit were grown for 24 hours after seeding at different densities (n = 3 biologically independent samples). c, Luminescent measurements of NanoBiT reporter cells (n = 128 biologically independent samples, * left to right q < 0.0001, <0.0001) and d, In-gel Nanoluciferase activity of mitochondria fractions treated with tunicamycin or galactose media for 48 hours (n = 3 experiments). e, Mitochondrial respiration of cells expressing NanoBiT reporter (n = 5 biologically independent samples). f, Cell proliferation in glucose (n = 6 biologically independent samples over 2 independent experiments) or galactose media (n = 6 biologically independent samples over 3 independent experiments, * left to right q = 0.0214, 0.0292, 0.0278, 0.0013, 0.0214) of NanoBiT reporter cells. g, Overview of high-throughput chemical screen. NanoBiT SC reporter cells are plated in 384-well plates with compound libraries, grown for 48 hours, and measured for luminescence and cell number. Compounds that increased the NanoBiT/cell number ratio were scored positive and a subset were validated by BN-PAGE. h, Luminescent measurements of NanoBiT reporter cells treated with lowest dose effective compound concentration from primary screen, see Supplementary Table 2 (DMSO, n = 60; drug treatments, n = 8 biologically independent samples, * q < 0.0001). i, BN-PAGE and j, Quantification of respirasome abundance (MTCO1/ATP5A intensity) of mitochondrial fractions isolated from cells treated with Vidofludimus (10 µM), Genistein (10 µM), Entinostat (2 µM), or OTX15 (0.4 µM) for 48 hours (n = 4 independent experiments). Data are presented as mean values + /- s.e.m., Student’s t-test with a two-stage linear step-up procedure of Benjamini, Krieger and Yekutieli, with Q = 5%.
Extended Data Fig. 2 Inhibitors of nucleotide synthesis promote respirasome assembly.
BN-PAGE of mitochondrial fractions isolated from U2OS cells treated with MTX (0.1 µM) (n = 2 experiments). b, Luminescence of NanoBiT reporter cells treated with MTX (DMSO, n = 36; MTX, n = 12 biologically independent samples, 100 nM q < 0.0001, 50 nM q < 0.0001). c, BN-PAGE of mitochondrial fractions isolated from 293 T and A375P cells treated with Brequinar (0.5 µM). d-g, Volcano plots of metabolite changes in cells treated with vidofludimus, Brequinar, CB-839, or methotrexate. h-k, Proliferation of cells grown in the presence of indicated compound (n = 11 biologically independent samples). l, BN-PAGE of mitochondrial fractions isolated from cells treated with 6-MP (25 µM) (n = 3 experiments). m,n, Heatmaps of metabolite changes in cells treated with 6-MP (25 µM) (n = 4 biologically independent samples). Cells were treated with compound for 48 hours, heatmap row/column groupings are Pearson’s correlations, data are presented as mean values +/− s.e.m., Student’s t-test with a two-stage linear step-up procedure of Benjamini, Krieger and Yekutieli, with Q = 5. 6-MP (6-mercaptopurine), MTX (methotrexate).
Extended Data Fig. 3 DHODH controls mitochondrial respirasome assembly through pyrimidine nucleotides.
a, BN-PAGE of mitochondrial fractions from cells treated with Vidofludimus in the presence or absence of uridine (n = 2 experiments). b, Luminescent measurements of NanoBiT reporter cells treated with Vidofludimus in the presence or absence of uridine (DMSO, n = 32; Vido, n = 16 biologically independent samples, * q < 0.0001). c, In-gel CI and CIV activities of mitochondrial fractions isolated from cells treated with Vidofludimus (n = 2 experiments). d, Western blot of proteins isolated from sgDHODH cells (n = 1 experiment). e, BN-PAGE of mitochondrial fractions from sgDHODH cells in the presence or absence of uridine (n = 2 experiments). f, Protein quantification of respiratory chain subunits in cells treated with Brequinar (n = 3 biologically independent samples). g, Proliferation measurements of cells treated with Vidofludimus for 3 days, initial seed of 1×105 cells (n = 3 biologically independent samples, Vidofludimus q = <0.0001, Vidofludimus + uridine q = 0.0004). h, Proliferation measurements of sgDHODH cells in the absence of uridine for 3 days, initial seed of 1×105 cells (n = 3 biologically independent samples, sgDHODH q < 0.0001, sgDHODH + uridine q < 0.0001). i, Volcano plot of metabolite changes in cells treated with Brequinar, relates to Fig. 3g. j, Heatmaps of hexosamine and phospholipid metabolites from cells treated with Brequinar (n = 3 biologically independent samples). k, Residual plot of alterations in metabolite levels of Brequinar treated cells with uridine supplementation. Cells were treated with compound for 48 hours unless otherwise noted, data are presented as mean values +/- s.e.m., Student’s t-test with a two-stage linear step-up procedure of Benjamini, Krieger and Yekutieli, with Q = 5%.
Extended Data Fig. 4 Mitochondrial phospholipid composition is altered with DHODH inhibition.
a, Abundances of total PC, PE, PG, and PI in cells are differentially regulated with DHODH inhibition in a uridine-dependent manner (n = 3 biologically independent samples, * left to right Vido q = 0.0189, 0.0020, 0.0020, 0.0035, 0.0020, Vido + uri [compared to Vido] q = 0.0157, 0.0015, 0.0029, 0.0489, Breq q = 0.0258, 0.0003, 0.0359, 0.0247, 0.0247, 0.0258, Breq + uri [compared to Breq] q = 0.0017, 0.0096). b, Abundances of total phospholipid classes in mitochondrial fractions are not regulated with DHODH inhibition (n = 8 biologically independent samples over 2 independent experiments). c, Heatmaps of mitochondrial PG, PI, and PS species in cells treated with Brequinar. Data from experiments 1 (n = 3 biologically independent samples) and 2 (n = 5 biologically independent samples) are shown. d, Quantification of mitochondrial cardiolipin acyl chain composition (summation) in cells treated with Brequinar. Data from experiment 1 is shown due to better detection of cardiolipin species (n = 3 biologically independent samples, Breq sn1 q = 0.0195, Breq + uri [compared to Breq] sn1 q = 0.0073, 0.0132, 0.0073, 0.0203, 0.0062, 0.0150, 0.0160, sn3 q = 0.0078, sn4 q = 0.0077, 0.0077, 0.0166, 0.0003, 0.0077, 0.0128, 0.0236). e, Quantification of cardiolipin species with >3 double bonds (box and whisker plot, line at median, 5-95% C.I., n = 8 biologically independent samples over 2 experiments, * left to right q = 0.0025, 0.0025, 0.0210, 0.0117). f, Heatmap of Brequinar-dependent changes in CL (p < 0.15, two-sided Student’s t-test). Data from experiment 1 (n = 3 biologically independent samples). In this figure, cells were treated with drug for 48 hours, heatmap row/column groupings are Pearson’s correlations, data are presented as mean values + /- s.e.m., Student’s t-test with a two-stage linear step-up procedure of Benjamini, Krieger and Yekutieli, with Q = 5%.
Extended Data Fig. 5 Ether lipid synthesis is increased relative to diacyl-PLs upon CTP depletion.
a, Heatmap of mitochondrial PC (p < 0.10, two-sided Student’s t-test). Data from experiment 2 (n = 5 biologically independent samples). Ether lipids are bolded. b, Lipidomic analysis of PC and PE subclass abundances in cells treated with Vidofludimus or Brequinar (min-max box plot, line at mean, n = 3 biologically independent samples, * left to right Vido q = 0.0041, 0.0041, 0.0016, 0.0476, Vido + uri [compared to Vido] q = 0.0025, 0.0023, 0.0019, 0.0002, 0.0025, 0.0005, 0.0025, Breq q = 0.0005, 0.0005, 0.0043, 0.0040, Breq + uri [compared to Breq] q = 0.0011, 0.0011, 0.0436, 0.0277). c, Quantification of plasmenyl/diacyl PC or PE abundances in cells treated with Vidofludimus or Brequinar (n = 3 biologically independent samples, * left to right Vido q = 0.0002, 0.0306, Vido + uri [compared to Vido] q = 0.0005 0.0011, Breq q = 0.0018, 0.0286, Breq + uri [compared to Breq] q = 0.0053, 0.0199). d, 14C-ethanolamine incorporation into PE in cells treated with Brequinar (n = 2 experiments). Lipid fractions were treated with TCA to cleave the ether vinyl bond, resulting in lyso-PE. e, Western blot of CTPS1 from sgCTPS1 cells (n = 1 experiment). f, BN-PAGE of mitochondrial fractions from sgCTPS1 cells in the presence or absence of cytidine (50 µg/ml) (n = 3 experiments). g, BN-PAGE of mitochondrial fractions from Brequinar treated cells in the presence or absence of CTP (200 µM) (n = 1 experiment). In this figure, cells were treated with drug for 48 hours, heatmap row/column groupings are Pearson’s correlations, data are presented as mean values + /- s.e.m., Student’s t-test with a two-stage linear step-up procedure of Benjamini, Krieger and Yekutieli, with Q = 5%. Breq (Brequinar), uri (uridine), and cyt (cytidine).
Extended Data Fig. 6 Ether lipid synthesis is required for respirasome assembly from DHODH inhibition.
a, Western blot of FAR1 from cells treated with Brequinar in the presence or absence of uridine (n = 2 experiments). b, Western blot of FAR1 from sgFAR1 cells (n = 2 experiments). c, Lipidomic analysis of PC and PE subclass abundances in mitochondrial fractions isolated from sgFAR1 cells treated with Brequinar (min-max box plot, line at mean, n = 3 biologically independent samples, Student’s t-test with a two-stage linear step-up procedure of Benjamini, Krieger and Yekutieli, with Q = 5%, * left to right sgControl DMSO [compared to sgControl Breq] q = 0.0003, 0.0034, 0.0007, sgControl DMSO [compared to sgFAR1 DMSO] q = 0.0208, <0.0001, 0.0005, 0.0016, 0.0016, 0.0001, sgControl Breq [compared to sgFAR1 Breq] q = <0.0001, <0.0001, <0.0001, <0.0001, <0.0001). d, Western blot of FAR1 from sgFAR1 cells exogenously expressing CRISPR-resistant FAR1 (n = 2 experiments). e, BN-PAGE and f, quantification of respirasome abundance (MTCO1/ATP5A intensity) of mitochondria isolated from sgFAR1 + exogenous FAR1 cells treated with Brequinar (n = 3 independent experiments, two-sided paired Student’s t-test, * left to right p = 0.0460, 0.0142). g, Western blot of AGPS from sgAGPS cells (n = 2 experiment). h, BN-PAGE and i, quantification of respirasome abundance of mitochondria isolated from sgAGPS cells treated with Brequinar (n = 4 independent experiments, two-sided paired Student’s t-test, * left to right p = 0.0445, 0.0005). j-l, Heatmaps of plasmenyl PC and PE (that is plasmalogens) and cardiolipin species in sgFAR1 mitochondria (n = 4 biologically independent samples). In this figure, cells were treated with Brequinar for 48 hours, heatmap row/column groupings are Pearson’s correlations, data are presented as mean values +/- s.e.m.
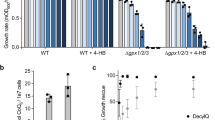
Extended Data Fig. 7 Proliferation under low nucleotide conditions requires increased mitochondrial respiratory capacity.
a, CI-dependent respiration of mitochondria isolated from sgFAR1 cells treated with Brequinar (n = 5 independent experiments, two-sided paired Student’s t-test, * left to right sgControl DMSO [compared to sgControl Breq] p = 0.0496, 0.0070, sgControl Breq [compared to sgFAR1 Breq] p = 0.0470, 0.0323, 0.0421, sgFAR1 DMSO [compared to sgFAR1 Breq] q = 0.0284). b, Proliferation of sgFAR1 cells treated with Brequinar (0.25 µM) (n = 12 biologically independent samples over 4 experiments, * left to right q = 0.0153, 0.0153). c, Western blot of HIGD2A from sgHIGD2A cells. d, Proliferation of sgHIGD2A cells treated with Brequinar (0.25 µM) (n = 12 biologically independent samples over 4 experiments, * left to right q = 0.0006, <0.0001, <0.0001). e, Pearson’s correlations of FAR1 CERES dependency score against other genes. Highlighted in green are ether lipid synthesis genes. f,g, Pearsons’s correlation of CERES dependency scores between DHODH and FAR1 or Ether lipid average (AGPS, FAR1, GNPAT, TMEM189). In this figure, cells were treated with Brequinar for 48 hours unless otherwise noted, data are presented as mean values +/− s.e.m., Student’s t-test with a two-stage linear step-up procedure of Benjamini, Krieger and Yekutieli, with Q = 5%.
Supplementary information
Supplementary Information
Supplementary Table 1. DNA sequences of expression constructs and CRISPR guides. Supplementary Table 2. Small-molecule screening data. Supplementary Fig. 1. Gating strategy for flow cytometry.
Supplementary Data 1
High-throughput small-molecule screen identifies compounds that increase NanoBiT SC reporter.
Supplementary Data 2
Metabolomic datasets for Vidofludimus, Brequinar, CB-839, methotrexate.
Supplementary Data 3
Proteomic dataset for Brequinar-treated cells supplemented with uridine.
Supplementary Data 4
Lipidomic datasets for Brequinar-treated cells supplemented with uridine.
Supplementary Data 5
Lipidomic dataset for sgFAR1 cells treated with Brequinar.
Supplementary Data 6
Lipidomic dataset for sgFAR1 mitochondria treated with Brequinar.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 1
Unprocessed western blots and/or gels.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 2
Unprocessed western blots and/or gels.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 3
Unprocessed western blots and/or gels.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 2
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 6
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 7
Unprocessed western blots and/or gels.
Rights and permissions
About this article
Cite this article
Bennett, C.F., O’Malley, K.E., Perry, E.A. et al. Peroxisomal-derived ether phospholipids link nucleotides to respirasome assembly. Nat Chem Biol 17, 703–710 (2021). https://doi.org/10.1038/s41589-021-00772-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-021-00772-z
This article is cited by
-
Tissue-specific roles of peroxisomes revealed by expression meta-analysis
Biology Direct (2024)
-
A FRET-based respirasome assembly screen identifies spleen tyrosine kinase as a target to improve muscle mitochondrial respiration and exercise performance in mice
Nature Communications (2023)
-
Multifaceted mitochondria: moving mitochondrial science beyond function and dysfunction
Nature Metabolism (2023)
-
TIM-4 orchestrates mitochondrial homeostasis to promote lung cancer progression via ANXA2/PI3K/AKT/OPA1 axis
Cell Death & Disease (2023)
-
Ether phospholipids are required for mitochondrial reactive oxygen species homeostasis
Nature Communications (2023)