Abstract
RNA-based gene therapy requires therapeutic RNA to function inside target cells without eliciting unwanted immune responses. RNA can be ferried into cells using non-viral drug delivery systems, which circumvent the limitations of viral delivery vectors. Here, we review the growing number of RNA therapeutic classes, their molecular mechanisms of action, and the design considerations for their respective delivery platforms. We describe polymer-based, lipid-based, and conjugate-based drug delivery systems, differentiating between those that passively and those that actively target specific cell types. Finally, we describe the path from preclinical drug delivery research to clinical approval, highlighting opportunities to improve the efficiency with which new drug delivery systems are discovered.
Similar content being viewed by others
Introduction
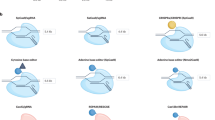
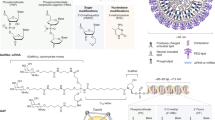
RNA therapies can manipulate gene expression or produce therapeutic proteins, making these drugs suitable for pathologies with established genetic targets, including infectious diseases, cancers, immune diseases and Mendelian disorders (including neurological disorders). Moreover, the ability to sequence hundreds of thousands of genomes, analyse gene expression at the single-cell level, and manipulate genes with programmable nucleases is driving the discovery of new targets for gene therapies. Yet the ability to manipulate these targets, especially non-coding DNA and the 85% of the genome that might be undruggable using small molecules1, is lessened without the capacity to deliver therapeutic RNA to diseased cells. In this Review, therapeutic RNA refers to antisense oligonucleotides (ASOs), such as gapmers, which contain DNA nucleotides flanked by RNA2, small interfering RNAs (siRNAs), or large RNAs, such as messenger RNA (mRNA) (Fig. 1). These RNA therapies act by targeting RNA or proteins, by encoding missing or defective proteins, or by mediating DNA or RNA editing. Irrespective of their therapeutic mechanism of action, the large size of some therapeutic RNAs, such as mRNAs, their anionic charge, and their susceptibility to RNases present in both the bloodstream and tissues make it difficult for therapeutic RNA to enter cells efficiently and function on its own.
a | One class of RNA therapeutics requires delivery of small RNA molecules. Small interfering RNAs (siRNAs) can reduce gene expression via RNA-induced silencing complex (RISC)-mediated mRNA degradation, antisense oligonucleotides (ASOs) can alter isoforms by binding to splice sites, and adenosine deaminase acting on RNA ASOs (ADAR-oligonucleotides) can edit RNA. In all three cases, these small RNAs can be designed with site-specific chemical modifications using solid-phase synthesis and can be delivered using nanoparticles or conjugate delivery systems. In this figure, the blue molecule represents the small therapeutic RNA being ferried into the cell. b | A second class of RNA therapeutics requires delivery of large RNA molecules. In vitro transcribed mRNA consists of a 5′ cap, 5′ and 3′ untranslated regions (UTRs), an open reading frame encoding antigen(s), and a 3′ poly(A) tail. c | mRNA payloads can encode nucleases that mediate DNA or RNA editing. mRNA can also be used to replace dysfunctional protein or encode antigens that confer longer-term immunity to a pathogen, such as SARS-CoV-2. mRNAs are transcribed in vitro and thus cannot currently be made with site-specific chemical modifications. In this figure, the blue molecule represents the protein encoded by the mRNA. Cas, CRISPR-associated protein; crRNA, CRISPR RNA; dCas9, dead Cas9; GFP, green fluorescent protein; pegRNA, prime editing guide RNA; sgRNA; single-guide RNA.
To overcome the barriers to safe and effective RNA delivery, scientists have developed both viral-vector-based and non-viral delivery systems that protect the RNA from degradation, maximize delivery to on-target cells and minimize exposure to off-target cells. Viral gene therapies3 have generated successful clinical readouts4,5,6,7,8,9, but the effectiveness of these approaches can be limited by pre-existing immunity10, viral-induced immunogenicity11, unwanted genomic integration12, payload size constraints13, the inability to re-dose, complications involved in upscaling14, and expensive vector production. Although scientists are overcoming some of these limitations15, they have fuelled the search for alternative drug delivery vehicles. Concurrent advances in the development of synthetic materials that encapsulate RNA, such as polymers, lipids and lipid nanoparticles (LNPs), have invigorated research into non-viral-based delivery systems, leading to US Food and Drug Administration (FDA) approval of subcutaneously administered N-acetylgalactosamine (GalNAc)–siRNA conjugates that target hepatocytes16,17,18, intravenously administered LNP-based siRNA drugs that target hepatocytes19, and emergency use authorization (EUA) and FDA approval20 for intramuscularly administered LNP-based mRNA COVID vaccines21,22. These approvals suggest that improved delivery to non-liver tissues (also known as extrahepatic tissues) as well as local delivery to the central nervous system, eye and ear could result in new drugs. Nanoparticle-based drug delivery systems may also be useful for non-viral DNA delivery, which has been reviewed elsewhere23.
Here, we detail the expanding number of therapeutic RNA payloads and highlight how the downstream biochemical mechanism of action influences delivery. After reviewing the chemistries commonly used in drug delivery systems as well as the approaches to targeting specific cells, we describe the series of experiments used to characterize nanoparticles in preclinical studies (hereafter referred to as the nanoparticle discovery pipeline) and identify opportunities to improve the efficiency of this pipeline. Finally, we describe the advantages, disadvantages and hallmarks of existing FDA-approved and EUA-approved RNA delivery systems.
Therapeutic RNA payloads
RNA drugs are often classified by the biochemical mechanism of action used to manipulate genes or gene expression (Fig. 1). When designing drug delivery vehicles, it is important to consider how these mechanisms of action influence the requirements for clinically relevant drug delivery. Oligonucleotide drugs, such as ASOs and siRNAs, that utilize enzymes endogenous to eukaryotic cells, such as RNAse H1 or the RNA-induced silencing complex (RISC), respectively, facilitate delivery by not requiring the delivery of large enzymes. Important improvements have been made in recent years in terms of the delivery of small molecules and macromolecules24, but most therapeutic oligonucleotides must still be maintained at high concentrations over time in order to manipulate the target gene25. For example, givosiran16 is administered subcutaneously at 2.5 mg/kg monthly, and lumasiran18 is administered subcutaneously at 3.0 mg/kg monthly for 3 months, then 3 mg/kg once every 3 months. More promising, however, is inclisiran26, which is administered as a 284 mg dose given subcutaneously as a single injection on day 1, day 90 and every 6 months thereafter. With additional improvements to delivery or siRNA design, it may become feasible to inject patients annually, which could coincide with a yearly checkup. By contrast, DNA nucleases, including many clustered regularly interspaced short palindromic repeat (CRISPR)-based systems, can cause long-term effects in cells even if the construct is only expressed transiently27. MicroRNAs (miRNAs) recruit RISC to complementary mRNA sequences, thereby facilitating targeted RNA interference. As a result, miRNA mimics, which are designed to increase native miRNA activity, and anti-miRNAs or antago-miRNAs, which inhibit miRNA activity, have been studied in animal models and used in clinical trials28. One phase I clinical trial investigated the use of MRX34, which uses liposomes to deliver a double-stranded miRNA-34a mimic, for the treatment of advanced solid tumours29. In a phase II clinical trial, the anti-miRNA-122 miravirsen, which binds miRNA-122 and leads to its subsequent inactivity, was tested for the treatment of hepatitis C30. Using a similar approach, RG-101, which antagonizes miRNA-122, was reported to reduce viral load by several logs in patients with chronic hepatitis C in a phase Ib clinical trial31. However, RG-101 was discontinued after the drug was found to cause hyperbilirubinaemia32. Finally, mRNA drugs promise a simple and flexible way to treat a litany of diseases; these therapeutics encompass antigen production, including the COVID-19 vaccine, protein replacement therapies and genome engineering.
siRNA
One biochemical mechanism of action that has been safely used in humans is siRNA-mediated gene silencing. These double-stranded RNAs with a molecular weight of approximately 13 kDa suppress protein translation by recruiting RISC to mRNA via Watson–Crick base pairing (Fig. 1a). Through the action of the catalytic RISC protein Ago2, a member of the Argonaute family, the target mRNA is cleaved. Alternatively, other Ago proteins (Ago1, Ago3 and Ago4) catalyse endonuclease-mediated nonspecific mRNA degradation by localizing the bound mRNA in processing (P)-bodies33,34. siRNA can reduce the expression of any protein-coding gene and has been approved by the FDA or the European Medicines Agency (EMA) in the form of the following drugs: patisiran, which is used to treat hereditary transthyretin-mediated amyloidosis (hATTR)19; givosiran, which is used to treat acute hepatic porphyria16; lumasiran, which is used to treat primary hyperoxaluria type 118; and inclisiran, which is used to treat hypercholesterolaemia17. In addition, the FDA has accepted a new drug application for vutrisiran35, an investigational RNA interference (RNAi) therapeutic for the treatment of hATTR amyloidosis with polyneuropathy in adults, following a successful phase III clinical trial36. The FDA approved the first siRNA drug 20 years19 after the first report of RNAi in eukaryotic cells37. This speedy clinical implementation of siRNA has been made easier by three traits. First, its small number of nucleotides enables scientists to use solid-phase synthesis to manufacture siRNA with site-specific chemical modifications, usually in the phosphate backbone and sugar rings. A whole suite of chemistries has been developed for a variety of ribose modifications, such as 2′-O-methyl (2′-OMe), 2′-methoxyethyl (2′-MOE), 2′-fluor (2′-F), locked nucleic acid oligonucleotides, constrained ethyl oligonucleotides (cEt), and tricyclo-DNA oligonucleotides (tc-DNA)38. Complementing these modifications are phosphate backbone modifications including phosphorothioates, phosphorodiamidate morpholino oligonucleotides (PMO), peptide nucleic acid oligonucleotides (PNA), and nucleobase modifications, such as 5-methylcytosine (m5C). Changing the combination of different chemical modifications of the siRNA has enabled scientists to improve the pharmacokinetic properties, innate immune response and stability39. Second, siRNA utilizes RISC, which is endogenous to eukaryotic cells and thus does not require the delivery of large enzymes with nuclease domains. Finally, given that siRNA interferes with mature mRNA, it requires only cytoplasmic delivery, which is easier to achieve than nuclear delivery.
Antisense oligonucleotides
ASOs are a second class of RNA therapeutics, and are oligonucleotides with a molecular weight of 6–9 kDa (ref.40). ASOs have the same manufacturing advantages as siRNA and have been approved by the FDA to treat familial hypercholesterolaemia41, hATTR amyloidosis with polyneuropathy42, specific subtypes of Duchenne muscular dystrophy43,44, and infantile-onset spinal muscular atrophy45. ASOs can act through three mechanisms of action (Fig. 1a). First, similar to siRNAs, ASOs bind mRNA via Watson–Crick base pairing, but unlike siRNAs, the ASO DNA–RNA heteroduplex recruits RNase H1 rather than RISC. RNase H1-dependent ASOs are also known as gapmers and lead to cleavage of the target RNA46. Second, ASOs can also interfere with splicing machinery by interacting with pre-mRNA, thereby promoting alternative splicing46 and increasing target protein expression47. Thus, unlike siRNA, which silences target genes, ASOs can be used to increase protein activity in diseases including Duchenne muscular dystrophy and spinal muscular atrophy. However, these two mechanisms require nuclear delivery so that the ASO–RNase H1 complex can interact with pre-mRNA. The third mode, which causes downregulation of mRNA expression, is the translational arrest of the targeted protein through binding with the translation initiation codon of the target mRNA48.
ASO chemical modification can influence the mechanism of action, as well as the target sequence binding affinity. Gapmers have a general chimeric structure of regions of five nucleotides of RNA-like residues flanking a central 10-nucleotide DNA region49. The gapmer can improve hybridization and nuclease resistance, while still retaining RNase H1 activation. As described above, other modifications, such as locked nucleic acid oligonucleotides, can be used instead of the 2′-modified designs, but although these modifications can improve potency, they can also increase toxicity50,51. The ASOs that involve steric blocking are RNAse H1-independent and are entirely made of RNA bases, and not DNA. Notably, ASOs often have a full phosphorothioate backbone, which can facilitate their transport into the nucleus52,53.
Chemically modified oligonucleotides with an antisense target binding motif have been designed with an engineered hairpin domain that recruits the endogenous RNA-editing enzyme adenosine deaminase acting on RNA (ADAR)54,55 (Fig. 1a). These ADAR-oligonucleotides, which have a molecular weight of approximately 10–35 kDa and can be manufactured with site-specific chemical modifications, use their single-stranded RNA domain to bind a target mRNA via Watson–Crick base pairing. The other domain recruits ADAR to the RNA, where ADAR converts adenosines to inosines; the inosines are subsequently read as guanosine by the endogenous translational machinery, leading to A to I to G RNA editing. A to I to G editing can occur on mature mRNA, suggesting that cytoplasmic delivery of ADAR-oligonucleotides can be sufficient to achieve an effect. Thus, ADAR-oligonucleotides represent an emerging class of oligonucleotides for treating genetic disease56.
mRNA
Another type of RNA therapeutic is mRNA, which can encode proteins that have therapeutic activity. Because of their size, mRNAs are in vitro transcribed and cannot currently be made with site-specific chemical modifications using solid-state synthesis (Fig. 1b). mRNA can be used to replace protein, using replacement therapies57; to reduce protein levels, using Cas9 cutting approaches58; or to repair protein mutations at the DNA level, using base editing59,60. In 2021, researchers reported the successful use of LNPs encapsulating Streptococcus pyogenes Cas9 (CRISPR-associated endonuclease Cas9) mRNA and a CRISPR guide RNA in six patients with hATTR amyloidosis with polyneuropathy; a single 0.3 mg/kg dose resulted in a mean reduction from baseline of blood transthyretin (TTR) levels of 87% at 28 days post-dose58. The gene product TTR is responsible for transporting vitamin A and thyroxine throughout the body, and mutations in this gene cause hATTR58. This milestone for a new therapeutic modality comes alongside the remarkable success of the now FDA-approved20, mRNA-based vaccine against SARS-CoV2 (Box 1)61,62. Other clinical examples of mRNA-mediated protein replacement include efforts to treat cystic fibrosis as well as ornithine transcarbamylase deficiency. Translate Bio is continuing its trial of mRNA-mediated protein replacement for cystic fibrosis, despite not showing much improved lung function in patients with cystic fibrosis63, and discontinuing its trial for ornithine transcarbamylase deficiency, owing to an undesirable pharmacokinetic and safety profile64. Similarly, Arcturus Therapeutics have received regulatory approval to initiate a phase II clinical trial of an mRNA therapeutic to treat ornithine transcarbamylase deficiency65. In addition to gene replacement, transient expression of myelin oligodendrocyte glycoprotein has led to immunological tolerance and subsequent treatment of experimental autoimmune encephalomyelitis in mice66. The antithesis of this tolerance mechanism is mRNA-mediated expression of an antigen to generate long-lasting immunity against the antigen, the so-called mRNA vaccines. Fundamental mRNA vaccine research67 focused on viruses such as Zika68, HIV69 and influenza70, or diseases such as melanoma71.
Intramuscular injections of human carcinoembryonic antigen mRNA have been used to induce antigen-specific immune responses in mice as far back as 1995 (ref.72). Since then, strides have been made and multiple therapeutic cancer vaccines are currently undergoing clinical trials. BNT111 from BioNTech’s FixVac platform targets a fixed combination of four mRNA-encoded melanoma-associated antigens (NY-ESO-1, MAGEA3, tyrosinase and TPTE). Phase I trials of BNT111 in patients with advanced melanoma showed partial responses and shrinkage of metastases after eight vaccinations given via intravenous injections73. Aside from vaccination, mRNA can be used to deliver co-stimulatory immune checkpoint molecules, such as OX40L for the treatment of solid tumours74. mRNA-2416 from Moderna was injected intratumorally every 2 weeks for up to 12 doses and was shown to be tolerable with increased OX40L protein expression and elevated pro-inflammatory activity and PD-L1 levels in patients with locally advanced, recurrent or metastatic solid malignancy or lymphoma74.
mRNA can also transiently express zinc finger nucleases, transcription activator-like nucleases, or nucleases derived from CRISPR–Cas systems75. Nucleases that are designed to manipulate DNA are particularly well suited for mRNA-based therapeutics, which produce protein for hours to days76 instead of weeks, as would be the case with DNA-based therapeutics; a short-lived DNA nuclease can create a long-lived gene editing event77. A clinical trial of an adeno-associated viral vector-based SaCas9 has been initiated78, but two arguments suggest that mRNA-based or protein-based transient Cas expression might be preferable in cases when the enzyme is an active DNA nuclease. First, long-term expression of an active DNA nuclease might lead to increased editing events in off-target loci27. Second, adeno-associated viral vector-based Cas9 delivery has been reported to lead to vector integration following DNA double-stranded breaks79.
Cas enzymes are also amenable to three types of biochemistry-driven improvements that expand their possible therapeutic potential80. In the first type of improvement, Cas enzymes can be rationally designed81 or evolved82 to target DNA next to different protospacer-adjacent motif sites. A second type of improvement is that Cas enzymes evolved to make double-stranded cuts in target DNA can be mutated in one or two inactive nuclease domains, leading to ‘nickases’ or ‘dead’ Cas (dCas9) enzymes, respectively. A third type of improvement is that Cas enzymes can be appended with domains that lead to transcriptional activation83, epigenome editing84,85, base editing86, changes to mitochondrial DNA87, reverse transcriptase-mediated gene insertion88, and transposition89. Notably, these changes can also be made to Cas12a enzymes90, which can require a shorter guide RNA and lead to staggered double-stranded cuts.
Complementing these DNA nucleases are RNA nucleases that bind and cleave RNA91,92 or, alternatively, can be engineered with an ADAR domain to enable RNA base editing93,94 (Fig. 1c). Compared with DNA-editing nucleases, non-viral-mediated delivery of an RNA nuclease facilitates transient, shorter-lived changes in gene expression93. Thus, whereas DNA nucleases may be well suited for chronic diseases, RNA nucleases may be more useful for short-term diseases, such as transient inflammation, or diseases driven by RNA pathogens such as coronaviruses95,96. Delivering CRISPR therapeutics is challenging because both the Cas protein and the single-guide RNA (sgRNA) must be present at sufficient concentrations to form intracellular ribonucleoproteins. In preclinical studies, this two-payload problem has been solved by concurrently delivering mRNA and sgRNA in nanoparticles77,96,97,98,99,100,101,102,103,104, by constitutively expressing sgRNA via adeno-associated virus (AAV) before injecting LNPs loaded with Cas9 mRNA105, or by injecting pre-complexed RNPs106,107,108. An alternative solution is to reduce the size of the Cas enzyme, which might make it easier to package alongside the sgRNA in the same delivery system. Several compact nucleases have been reported, including a Cas12j (termed Casϕ)109, Cas12f110,111, Cas13bt112 and Cas13ct112.
Synthetic vehicles for RNA delivery
Although different RNA payloads can have different biochemical mechanisms of action, all of them must avoid clearance by off-target organs, must access the correct tissue, must interact with the desired cell type in a complex tissue microenvironment, must be taken up by endocytosis, and must exit the endosome, without eliciting a deleterious immune response113. Although small oligonucleotide RNA therapeutics, including ASOs, siRNAs and ADAR-oligonucleotides, can be modified using stable chemistries and delivered using conjugates, mRNA-based and DNA-based therapeutics require a vehicle for entry into a cell. To facilitate this process, scientists have developed several RNA delivery systems using a range of materials, including polymers and LNPs.
Lipids and lipid-based nanoparticles
LNPs are a key class of drug delivery system that includes nanoparticles approved by the FDA for liver siRNA delivery19 and for mRNA vaccine delivery61,62. On the basis of the size of the hydrophilic head group relative to the size of the hydrophobic tail or tails114, lipids form distinct structures including micelles, liposomes and LNPs (Fig. 2a). FDA-approved LNPs contain variations of four basic components: a cationic or ionizable lipid, cholesterol, a helper lipid, and a poly(ethylene glycol) (PEG)-lipid (Fig. 2b,c). Scientists have investigated the structure of lipid-based delivery systems complexed with nucleic acid115,116 and demonstrated that lipid structure alters how LNPs interact with cells117. Given that lipid structure influences delivery and that lipids can be easily synthesized using chemistries including Michael addition-based, epoxide-based and alcohol-based reactions, scientists have created libraries of dozens to thousands of chemically distinct lipid delivery systems118,119. Many of these efforts focused on improving siRNA delivery to hepatocytes in mice120. This work, along with a more rational approach to lipid design117, reduced the dose required for robust in vivo hepatocyte gene silencing in mice from approximately 1.0 mg/kg (ref.121) to 0.002 mg/kg (ref.122). Key lipids that delivered siRNA in non-human primates included C12-200, which was synthesized using epoxide–amine chemistry120; cKK-E12, a peptide-like lipid compound122; DLin-KC2-DMA, an ionizable lipid identified using rational design117; and DLin-MC3-DMA123, which was used in patisiran to treat hATTR19 (Fig. 2c,d).
a | Lipid-based structures can include micelles, which consist of a lipid monolayer, or liposomes, which consist of a bilayer. Lipid nanoparticles are composed of multiple lipid layers as well as microdomains of lipid and nucleic acid. b,c | In addition to the RNA payload, LNPs often consist of cholesterol, a helper lipid, a PEG-lipid (all shown in part b), and a cationic or ionizable lipid (part c). Despite the variety of chemical structures and lipid lengths, all the lipids in part c contain amine groups, which become positively charged at lower pH; this charge binds the anionic backbone of the RNA, providing a driving force for the formation of a stable LNP. d | The molar ratios of the four components making up the FDA-approved Acuitas/BioNTech/Pfizer COVID vaccine and patisiran, which delivers siRNA to the liver.
LNPs have also delivered mRNA to the liver in mice, in non-human primates and in humans. In some cases, LNPs utilized lipids previously developed for siRNA delivery. For example, in mice, LNPs formulated with cKK-E12124,125, C12-200126, and DLin-MC3-DMA127 delivered mRNA to the liver. Newer lipids reported, such as LP0177(Intellia Therapeutics), Lipid H128(Moderna), and FTT5103(Ohio State and Beam Therapeutics), have also delivered mRNA to the mouse liver. Recently, two LNPs formulated with an unreported cationic or ionizable lipid, PEG-lipid, cholesterol and 1,2-distearoyl-sn-glycero-3-phosphocholine (DSPC) delivered mRNA encoding a base-editing Cas9 and sgRNA targeting PCSK9 to the liver in non-human primates59,60. A single LNP administration led to months of sustained PCSK9 silencing. Moreover, long-term PCSK9 silencing mediated by antibodies129 or siRNA17 has shown beneficial effects in cardiovascular disease in humans. Separately, Beam Therapeutics reported that LNP-mediated delivery of base editors led to sustained effects in the liver of non-human primates130. In addition to these preclinical studies, Intellia has released data in patients dosed with its NTLA-2001, used to inactivate the TTR gene. Inactivation of TTR has also previously been validated in humans; siRNA or ASOs targeting TTR slowed the progression of hATTR amyloidosis with polyneuropathy36,42.
In addition to the RNA payload, the Alnylam, Moderna and Pfizer/BioNTech/Acuitas LNPs comprise four components: the cationic or ionizable lipids DLin-MC3-DMA (Alnylam), SM-102 (Moderna), or ALC-0315 (Pfizer/BioNTech/Acuitas), cholesterol, the PEG-lipids PEG-2000-C-DMG (Alnylam), PEG-2000-DMG (Moderna), or ALC-0159 (Pfizer/BioNTech/Acuitas), and DSPC (Fig. 2b–d). Although most preclinical studies have evaluated how the structure of the cationic or ionizable lipid influences delivery, the other three components can also affect delivery131,132. For example, by changing the cholesterol, PEG-lipid or ‘helper’ lipid, an LNP that delivered siRNA to pulmonary and cardiovascular endothelial cells in mice133 and non-human primates134 was retargeted to deliver siRNA135, sgRNA136 or mRNA99 to bone marrow, hepatic and splenic endothelial cells after intravenous administration as well as lung epithelial cells after nebulization137. In additional examples, changing the PEG-lipid structure or its molar percentage altered LNP pharmacokinetics and liver siRNA delivery138 in mice and affected delivery within the eye139. Both the PEG and lipid components of the PEG-lipid affect how it interacts with the LNP and cells: the lipid ‘anchors’ the PEG-lipid into the LNP, while the hydrophilic PEG interacts with water in the blood, thereby creating an aqueous barrier similar to other PEGylated nanomedicines140. Likewise, although most LNPs have been formulated with unmodified cholesterols, LNP delivery in cell cultures and in mice has been shown to be improved by use of oxidized cholesterols124, esterified cholesterols136, or cholesterol analogues such as phytosterols116. Although the mechanism behind cholesterol-mediated improvements in delivery remains unknown, incorporating modified cholesterol into LNPs can change their structure141. Researchers have demonstrated that replacing DSPC with other lipids can promote LNP delivery to the spleen or lungs125,142. Similarly, by adding another lipid to the LNP, thereby changing the LNP from a four-component system to a five-component system, LNPs were targeted to the lung and spleen in a process termed selective organ targeting101. Finally, both Intellia143 and Beam Therapeutics130 have shared information that LNPs can be made to deliver mRNA to haematopoietic stem and progenitor cells in mice, with the hope of developing in vivo haematopoietic stem cell-targeting therapies.
Polymers and polymer-based nanoparticles
Many non-viral RNA delivery systems also utilize polymers and polymeric nanoparticles144 (Fig. 3a). Chemists can vary polymer traits including charge, degradability and molecular weight, all of which influence how polymers deliver RNA into cells145,146. One frequently used polymer is poly(lactic-co-glycolic acid) (PLGA). PLGA drug delivery systems have been approved by the FDA for the delivery of small-molecule drugs but not for the delivery of nucleic acids147. At neutral pH, PLGA does not have the positive charge required to complex the anionic RNA phosphodiester backbone. Thus, to utilize PLGA as an RNA delivery system, scientists have added cationic chemical groups such as chitosan to deliver siRNA in mice148.
a | Polymeric nanoparticles and polymers based on poly(ethylenimine) (PEI), poly(l-lysine) (PLL), and poly(beta-amino-ester) (PBAE) use cationic amine groups to complex the anionic phosphodiester backbone of RNA. Polymers based on poly(lactic-co-glycolic acid) (PLGA) are typically engineered to contain separate cationic groups. b | Dendrimers are polymeric structures with a defined number of molecules emanating from a core. PAMAM, poly(amidoamine).
Polymers that contain amine groups that can become cationic, such as polyethylenimine (PEI) and poly(l-lysine) (PLL), can complex with RNA via electrostatic interactions and deliver it into cells149,150. However, unmodified PEI and PLL are not always well tolerated151, and PEI transfection capability and toxicity increase with molecular weight152. Thus, PEI and PLL have been chemically modified to improve in vivo efficacy and tolerability. For example, nanoparticles made with PEG-grafted PEI have been used to deliver mRNA to immune cells in the lungs153, and cyclodextrin-PEI conjugates have been used to deliver an mRNA vaccine in vivo154. Similarly, iron oxide nanoparticles were surface-modified with PLL to deliver genes to the central nervous system in mice155.
Another cationic polymer class is the poly(beta-amino ester)s (PBAEs), which are synthesized by conjugating amine monomers to diacrylates. These polymers, which were designed to have improved biodegradation and cytotoxicity relative to PEI and PLL156, contain cationic amines as well as biodegradable ester bonds157. Early research used Michael addition chemistry to synthesize hundreds of chemically distinct PBAEs and then evaluated how these nanoparticles delivered DNA158,159 and RNA in cell culture160. These PBAE ‘libraries’ enabled researchers to study how PBAE chemical structure influences drug delivery, thereby generating design rules for subsequent PBAEs161,162. Initial design rules suggested that effective polymers were almost always hydrophobic, had monoalcohol or dialcohol side groups, and had linear bis (secondary) amines161. A follow-up study using these design criteria showed that top polymers were all formed from amino alcohols and were similar chemically, only differing by one carbon162. More generally, these studies established the feasibility of high-throughput chemical synthesis followed by high-throughput drug delivery studies, which has been applied to other nanoparticle chemical classes, including LNPs118. PBAEs have been used for the delivery of DNA vectors to pulmonary cells after nebulization163, the delivery of mRNA intranasally164, and the delivery of siRNA to a human orthotopic glioblastoma tumour model in mice165. More recently, researchers have also used PBAE-based polymers to deliver Cas13a mRNA and to guide RNA to the respiratory tract of mice and hamsters via nebulization for the treatment of SARS-Cov-2 (ref.96).
Researchers have also synthesized lipid–polymer hybrids and found that adding lipids to PBAEs improved serum stability and delivery166,167. An additional polymer class used for RNA delivery is dendrimers, which consist of a defined number of branched monomers emanating from a central core molecule (Fig. 3b). Dendrimers synthesized with cationic groups, such as poly(amidoamine) (PAMAM) or PLL, can form complexes and deliver RNA into cells. Dendrimers have delivered RNA to the central nervous system168, acted as intramuscular vaccines against the Ebola and H1N1 viruses169, and delivered siRNA to hepatic endothelial cells170. Dendrimer structure has also been modified to protect nucleic acids from enzymatic degradation171 and to enhance endosomal escape172.
Active versus passive tissue targeting
Passive tissue targeting
As outlined in the section ‘Lipids and lipid-based nanoparticles’, research has demonstrated that LNPs originally developed for liver siRNA or mRNA delivery can be redirected to other organs without the need for antibody fragments, peptides, aptamers or other active targeting ligands that bind specific receptors on the surface of target cells173. For the purposes of this Review, we define such retargeting, probably driven by interactions between the nanoparticle and serum proteins in endogenous trafficking pathways, as a process termed ‘passive targeting’ or ‘endogenous targeting’ (Fig. 4a). As a sphere becomes smaller, its surface area to volume ratio increases; as a result, nanoparticles have large surface areas174 and when a nanoparticle comes into contact with a biological milieu, many biomolecules can coat its surface175. By covering the nanoparticle surface, these coronas (proteins that bind to the nanoparticle surface) change the chemical and biological molecules at the surface of the nanoparticle, thereby altering how the nanoparticle interacts with immune cells176 and on-target tissues177. In one clinical example178, apolipoprotein E (ApoE) adsorption was required for ionizable, but not cationic, lipids to deliver siRNA to hepatocytes179. Researchers have found LNPs that can be trafficked via an albumin-dependent mechanism180 as well as LNPs with variable dependence on low-density lipoprotein (LDL), very low-density lipoprotein (VLDL) and caveolin-1 receptors136,181. In addition to the effect that the protein corona can have on LNP tropism, LNP size can also contribute to passive tissue targeting. In one example, nanoparticle size influenced the potency of siRNA-based LNPs in vivo182. In another example, nanoparticle size governed the frequency with which an LNP was cleared by immune cells in the lymph node183. Similarly, several groups have demonstrated that LNP charge can affect delivery. In one example, scientists developed a lipoplex that delivered mRNA to the spleen by altering its charge142. Similar systems have delivered therapeutic mRNA in different animal models66,184. Likewise, by adding a cationic lipid to the LNP, scientists retargeted an LNP with liver tropism to the lung125. These splenic and pulmonary delivery datasets were subsequently observed using LNPs that facilitated mRNA delivery and Cas9-mediated gene editing101.
a–d | Delivery vehicles can reach desired cells using passive or endogenous targeting (part a), which leads to adsorption of serum biomolecules onto the outside of the lipid nanoparticle (LNP) in the bloodstream. For example, the serum lipoprotein ApoE binds to LNPs, leading to delivery via low-density lipoprotein receptor (LDLR) expressed on hepatocytes. Active targeting employs a ligand directly conjugated to a nucleic acid (part b), an antibody directly conjugated to a nucleic acid (part c) or a ligand or antibody conjugated to a nanoparticle to target a receptor expressed on the cell (part d). ASGPR, asialoglycoprotein receptor; GalNAc, N-acetylgalactosamine. ApoE, apolipoprotein E.
Active targeting
RNA can also be ferried into on-target cells using ‘active targeting’. In this strategy, a ligand that binds a specific biomolecule is added to the delivery system (Fig. 4b–d). The most clinically validated examples are GalNAc–siRNA and GalNAc–ASO conjugates, which have led to the FDA-approved drugs givosiran16 and lumasiran18, as well as the EMA-approved drug inclisiran17. GalNAc is a carbohydrate-derived trivalent ligand that binds the asialoglycoprotein receptor (ASGPR) (Fig. 4b). ASGPR is an ideal receptor for active targeting: it is highly expressed on target cells (in this case hepatocytes), not expressed on other cell types, leads to rapid endocytosis upon GalNAc binding, and is rapidly recycled to the cell surface following endocytosis. GalNAc also exhibits several traits that make it an ideal targeting ligand. First, it has a molecular weight of <2 kDa, which is many times less than the molecular weight of the ASO or siRNA it delivers185. This low molecular weight ensures that most of the drug being administered is siRNA and not GalNAc. Second, improvements can be made to its chemical structure to enhance silencing in vivo185,186, and its interactions with serum proteins are well understood187; in addition, site-specific modifications made to the ribose and phosphate moieties have reduced nuclease-mediated siRNA degradation as well as off-target mRNA binding187,188,189. The combination of this ideal receptor and optimized ligand has resulted in subcutaneous delivery of siRNA and ASO at doses far below those that cause toxicity in large-animal models190,191.
Small ligands have also been developed for extrahepatic delivery. Specifically, by measuring the biodistribution and delivery mediated by a library of different siRNA–lipid conjugates, researchers found that hydrophobic conjugates accumulated in the liver, whereas less hydrophobic conjugates accumulated in the kidneys192. In the same study, the authors found that when compared with cholesterol (a well studied hepatic conjugate), both dichloroacetic acid alone and dichloroacetic acid with a phosphocholine polar head group improved siRNA delivery to extrahepatic tissues such as the lung and heart and, to a lesser degree, improved siRNA delivery to skeletal muscle and fat. In a follow-up study, researchers identified two lipid transport pathways that could be guiding delivery: hydrophobic siRNA conjugates tended to spontaneously associate with LDL, an association known to improve trafficking to the liver, whereas less-hydrophobic conjugates bound to high-density lipoprotein (HDL), part of the reverse cholesterol transport pathway193. A 2021 study reported that cholesterol-functionalized DNA–RNA heteroduplexes were capable of crossing the blood–brain barrier in mice and rats after systemic administration. Once again, the chemical structure of the lipid and conjugate was critical for cell targeting, gene silencing and pharmacokinetics194. RNA aptamers, which are RNAs that fold into defined three-dimensional structures195, have also emerged as ligands for specific cell-targeted delivery of RNAi-based therapeutics. In one example, researchers conjugated an siRNA-targeting STAT3, a key regulator of glioblastoma, to the receptor tyrosine kinase anti-PDGFRα RNA aptamer. This delivery system inhibited the expression of target genes and showed strong reduction of cell viability in vitro196.
A distinct targeting mechanism requires the use of antibody fragments or antibodies to target cell types. For example, siRNA and ASO have also been delivered to extrahepatic tissues using antibody–siRNA conjugates (Fig. 4c). In one example, an anti-CD71 antibody fragment delivered siRNA to the heart and skeletal muscle, leading to extended on-target gene silencing197. Data using either anti-CD71 fragments197 or monoclonal antibodies198 have demonstrated long-term muscle silencing in preclinical models, including 12 weeks of silencing in non-human primates after a single administration198. As a result, Avidity Biosciences initiated a phase I/II clinical trial delivering siRNA against myotonic dystrophy protein kinase to treat myotonic dystrophy type I in 2021.
Although ASOs and siRNAs are small enough to be delivered using a small conjugate or antibody, mRNAs are too large. For this reason, mRNA has been formulated inside nanoparticles that are decorated with antibodies (Fig. 4d). Researchers have developed a cell-targeting platform known as Anchored Secondary scFv Enabling Targeting (ASSET) that used monoclonal antibody-coated LNPs to deliver siRNA or mRNA. This system relies on a membrane-anchored lipoprotein within the LNP that binds to an antibody’s fragment (Fc) domain; switching out the variable region enables specific targeting to different cell subsets199. ASSET was used to facilitate LNP binding to extrahepatic cell types for the purpose of treating inflammatory bowel disease200. These approaches can also be targeted to receptors in specific conformations, as demonstrated by recent data using ASSET to deliver LNPs to a specific, high-affinity conformation of α4β7 (ref.201). Finally, LNPs covalently conjugated to an antibody that binds to plasmalemma vesicle-associated protein have been used to target lung cells by facilitating specific caveolin-mediated endocytosis202, and LNPs conjugated to mannose have been used to immunize mice against H1N1203.
The pathway to clinical RNA delivery
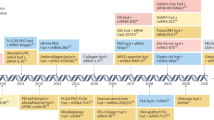
Independently of their chemical structure or targeting mechanism, nanoparticle-based RNA delivery systems are selected for clinical trials using a series of preclinical experiments here termed ‘the nanoparticle discovery pipeline’. This pipeline often starts with initial high-throughput studies in cell culture. Based on these in vitro delivery data, which are used to optimize nanoparticle traits, a small number of nanoparticles is selected for mouse studies before a smaller number is tested in rats and, finally, an even smaller number in non-human primates (Fig. 5a). If the nanoparticle has a sufficiently safe toxicity profile in non-human primates, the delivery system can then be considered for subsequent clinical trials.
a | Thousands of nanoparticles are often synthesized and tested in vitro before a few nanoparticles are evaluated in mice. However, in vitro delivery can be a poor predictor of in vivo delivery. A smaller number of nanoparticles are then often tested in rats or non-human primates (NHPs). Given that NHPs are the best preclinical models for human delivery, one key opportunity to improve the efficiency of this pipeline is to understand which small-animal model is most predictive of NHP efficacy and tolerability; this species-to-species drug delivery relationship is understudied. b | Tissue weight as a percentage of total animal body weight (bw) in female animals. Relative organ size varies across species, which may alter nanoparticle on-target and off-target delivery.
High-throughput nanoparticle discovery pipeline
Although the nanoparticle discovery pipeline has identified nanoparticles that deliver RNA to immune cells after intramuscular administration21,22 and hepatocytes after systemic administration19, opportunities do exist to improve the efficiency of the pipeline. For example, chemists can synthesize thousands of nanoparticles, but running a multi-thousand-mouse experiment is infeasible. As a result, nanoparticles are first evaluated in cell culture, even though delivery in cell culture can poorly predict delivery in vivo204. To address this problem, DNA barcoding assays to test thousands of nanoparticles in vivo have been developed205,206,207,208. In these assays, LNP-1 is formulated to carry DNA barcode 1 and a nucleic acid (such as siRNA or mRNA), whereas LNP-N is formulated to carry DNA barcode N and the same nucleic acid. After pooling the LNPs together and administering them to mice, cells in which functional RNA delivery occurs — such as mRNA-mediated protein production99 or siRNA-mediated gene silencing135 — are isolated and all N barcodes are quantified. This approach has been used to deliver RNA to non-hepatocytes without targeting ligands99,124,136,209,210,211. A second way to improve the nanoparticle discovery pipeline is to understand which small animals best predict delivery in a non-human primate. Species variability can derive from physiological differences in basal metabolic rates212, which can affect nanoparticle delivery213,214,215, or differences in serum lipids216, which can affect the colloidal stability, biodistribution, cellular interactions, toxicity and clearance of the resulting nanoparticle coronas217. Additional species variability probably derives from the mass of a given tissue, relative to the total animal weight, which is not conserved (Fig. 5b). For example, mouse218,219,220,221,222 and rat223 livers are larger, normalized to overall body mass, than those of non-human primates224 and humans225,226,227,228. These differences may increase the effective off-target liver delivery in mice and rats, relative to non-human primates and humans. To address some of these concerns, researchers have developed species-agnostic nanoparticle delivery screening (SANDS)229. Using SANDS, the authors tested how dozens of LNPs delivered mRNA to normal mice, mice with primatized livers, and mice with humanized livers. After quantifying the correlation between delivery in murine, non-human primate and human cells in vivo, and observing species-dependent delivery, the authors used RNA sequencing to identify genes that might differentiate delivery across species. This approach might enable scientists to compare how many chemically distinct nanoparticles deliver drugs across different species.
Hallmarks of clinically relevant delivery systems
As the nanoparticle pipeline continues to improve, next-generation delivery vehicles might benefit from recapitulating traits shared among current clinical RNA delivery vehicles. For example, as part of its clinical trials, detailed pharmacokinetic and clearance studies of patisiran in humans have been published230,231. In addition to describing sustained gene silencing over several months, these studies demonstrated that patisiran had consistent mean serum concentration and dynamics over 18 months with repeated administration. Similarly, the FDA Center for Drug Evaluation and Research report on patisiran232 describes the accepted animal models, toxicology readouts and time points used in the approval process. These data provide a roadmap for the clinical characterization of nanoparticle RNA drugs. More generally, the three RNA drug delivery systems approved by the FDA19,20 or granted an EUA21 to date tend to share six characteristics that taken together could constitute hallmarks of approved delivery vehicles (Fig. 6).
First, clinical delivery systems tend to be synthesized using scalable chemistry that is often biodegradable. For example, adding ester bonds, which can degrade in water, to ionizable lipids improved LNP safety233; the Moderna, Acuitas and Alnylam lipids used in humans contain esters (Fig. 2c). Second, the drug delivery system should be chemically simple enough to be manufactured at human scale; for example, in a hypothetical clinical trial injecting 100 kg patients with 6 mg/kg lipid and 0.3 mg/kg RNA (that is, a lipid:RNA mass ratio of 20:1), assuming lipid loss during the synthesis and formulation process, nearly 1 g of lipid would need to be synthesized per injection. GalNAc conjugates can be synthesized and, separately, conjugated to siRNAs or ASOs at human scale (that is, with large-batch manufacturing capability that is compliant with Current Good Manufacturing Practice). We note that clinically approved LNPs have so far included four lipid components and have not included targeting ligands. Determining the advantages and disadvantages of adding targeting ligands in clinical nanoparticles will be important234. Third, the drug delivery system must have an acceptable ratio of on-target to off-target delivery. On-target and off-target delivery should be measured both as biodistribution (that is, where does the delivery system go?) and as function (that is, where does the payload affect cell function?). As over 95% of RNA can be retained in endosomes235, biodistribution is necessary, but not sufficient, for functional cytoplasmic RNA delivery. Research has also shown that small copy numbers of siRNAs are sufficient for gene knockdown in vitro236, implying that siRNA acts in a catalytic manner. Fourth, the dose required for RNA efficacy must be substantially lower than the dose at which toxicity occurs. Ideally, this finding is observed in non-human primates, as mice have historically not been rigorous models of RNA toxicity, for yet-to-be-determined reasons. Fifth, the activity of the drug should be consistent across many batches, even after shipping. To this end, scientists are developing methods of stabilizing mRNA drugs; a CureVac mRNA platform was active after lyophilization and storage at 40 °C for up to 6 months and room temperature for 3 years237. However, for LNP-encapsulated mRNA, lyophilization might decrease stability by promoting lipid crystallization, unless cryoprotectants such as sucrose are used238. For example, by adding 10% sucrose, lipid nanoparticles complexed with RNA can be lyophilized and stored at room temperature for ≥8 months239. Sixth, in most clinical settings, re-dosing the RNA drug without losing efficacy or safety will be necessary to maintain the biological effect or ‘dose to effect’. siRNA drugs have been safely re-dosed in patients when doses have been given 3 weeks apart19. mRNA vaccines have been safely dosed twice, with doses either 3 weeks or 4 weeks apart21,22. Re-dosing mRNA weekly has been reported to reduce its efficacy in mice240, with this effect being driven by a subset of B lymphocytes.
FDA-approved and EMA-approved RNA therapeutics
Although the criteria constituting hallmarks of approved delivery vehicles make clinically relevant delivery challenging, the number of successful RNA drugs is increasing. A leading example is the GalNAc–siRNA conjugates, which have demonstrated efficacy and safety in patients. Like the mRNA-LNP vaccines, GalNAc–siRNA conjugates can easily be redesigned to treat different diseases, since only the siRNA sequence needs to change. As a result, the GalNAc–siRNA conjugates givosiran, which treats acute intermittent hepatic porphyria, and lumasiran, which treats primary hyperoxaluria type 1, have been approved by the FDA. Inclisiran, a subcutaneous therapeutic used to treat hypercholesterolaemia or mixed dyslipidaemia by inhibiting hepatic synthesis of proprotein convertase subtilisin–kexin type 9 (PCSK9)17, has been approved by the EMA, but not by the FDA (which delayed approval owing to issues at a manufacturing plant). The GalNAc–siRNA conjugates fitusiran and vutrisiran have also been used to generate promising clinical data. Fitusiran is used to treat haemophilia A or B241 by targeting antithrombin mRNA in the liver241. Vutrisiran (see above) is used to treat hATTR amyloidosis with polyneuropathy in adults, and generated positive top-line results in a phase III study242. Arrowhead, Silence Therapeutics and Dicerna are also utilizing GalNAc–siRNA conjugates in ongoing phase I/II trials for the treatment of hepatitis B, hereditary haemochromatosis, and primary hyperoxaluria, among other diseases62, and Ionis Pharmaceuticals is using GalNAc to deliver ASOs243. For all of these drugs, long-term gene silencing mediated by GalNAc conjugates is important for their eventual clinical use and to improve patient compliance. For example, inclisiran reduces PCSK9 levels for up to 6 months after administration in humans26.
Conclusions
In the past decade, preclinical and clinical data have hinted at the potential for RNA therapies to treat disease. However, to fully reach their potential, several advances are needed. One advance will be to understand how RNA payloads interact with delivery vehicles and how these interactions affect targeting and tolerability. For example, emerging evidence suggests that nanoparticle tropism can change with RNA payload99. This effect may be caused by changes in the nanoparticle and therefore the biomolecules with which it interacts in the body. Alternatively, payload-dependent tropism may be affected by cell state244, since a cell that is optimized for the production of exogenous mRNA might not be optimized for siRNA-based mRNA silencing. Similarly, although LNP interactions with ApoE are known to drive liver delivery178, a need remains to identify trafficking mechanisms that promote delivery to extrahepatic tissues; it is feasible that disease-specific trafficking could be exploited to enhance extrahepatic delivery. One potential approach is to use naturally occurring systems such as PEG10, which package mRNA in human cells245 or extracellular vesicles, which are outside the scope of this manuscript and have been reviewed elsewhere246. Another related advance is the need to understand how chemical modifications made to the RNA payload influence RNA stability, avoidance of intracellular off-target effects, such as binding based on partial complementarity247, or activation of the innate immune system248. For example, CureVac recently reported that its mRNA vaccine, which utilizes unmodified mRNA, did not provide robust protection against COVID-19249. Unmodified mRNA programmes led by Translate Bio have also been ineffective when nebulized to treat cystic fibrosis63 or systemically administered to treat ornithine transcarbamylase deficiency64. These findings, alongside evidence that chemical modification patterns boost siRNA and ASO efficacy250 and that the untranslated regions of mRNA can be engineered for improved251 or cell-type-specific activity252, suggest that mRNA payloads can be improved in future therapies. Finally, we need an improved understanding of how efficacy and tolerability in smaller animal models (such as mice and rats) predict efficacy and tolerability in non-human primates and humans. Given the major ethical issues with testing many potential drug delivery systems in non-human primates, one key advance would be the identification of smaller animal models that are maximally predictive of delivery in non-human primates253. In cancer biology, researchers identified genetically engineered mice that more accurately recreated clinical outcomes from human trials carried out in parallel254. If we are thereby able to expand our understanding of how the RNA drug, drug delivery system and body all interact with one another, patients will benefit from effective next-generation gene therapies.
References
Hopkins, A. L. & Groom, C. R. The druggable genome. Nat. Rev. Drug. Discov. 1, 727–730 (2002).
Roberts, T. C., Langer, R. & Wood, M. J. A. Advances in oligonucleotide drug delivery. Nat. Rev. Drug Discov. 19, 673–694 (2020).
High, K. A. & Roncarolo, M. G. Gene therapy. N. Engl. J. Med. 381, 455–464 (2019).
Pasi, K. J. et al. Multiyear follow-up of AAV5-hFVIII-SQ gene therapy for hemophilia A. N. Engl. J. Med. 382, 29–40 (2020).
Mendell, J. R. et al. Single-dose gene-replacement therapy for spinal muscular atrophy. N. Engl. J. Med. 377, 1713–1722 (2017).
Frangoul, H. et al. CRISPR-Cas9 gene editing for sickle cell disease and β-thalassemia. N. Engl. J. Med. 384, 252–260 (2021).
Esrick, E. B. et al. Post-transcriptional genetic silencing of BCL11A to treat sickle cell disease. N. Engl. J. Med. 384, 205–215 (2021).
Kohn, D. B. et al. Autologous ex vivo lentiviral gene therapy for adenosine deaminase deficiency. N. Engl. J. Med. 384, 2002–2013 (2021).
Russell, S. et al. Efficacy and safety of voretigene neparvovec (AAV2-hRPE65v2) in patients with RPE65-mediated inherited retinal dystrophy: a randomised, controlled, open-label, phase 3 trial. Lancet 390, 849–860 (2017).
Aronson, S. J. et al. Prevalence and relevance of pre-existing anti-adeno-associated virus immunity in the context of gene therapy for Crigler–Najjar syndrome. Hum. Gene Ther. 30, 1297–1305 (2019).
Bryson, T. E., Anglin, C. M., Bridges, P. H. & Cottle, R. N. Nuclease-mediated gene therapies for inherited metabolic diseases of the liver. Yale J. Biol. Med. 90, 553–566 (2017).
Nguyen, G. N. et al. A long-term study of AAV gene therapy in dogs with hemophilia A identifies clonal expansions of transduced liver cells. Nat. Biotechnol. 39, 47–55 (2021).
Wu, Z., Yang, H. & Colosi, P. Effect of genome size on AAV vector packaging. Mol. Ther. 18, 80–86 (2010).
Chandler, M., Panigaj, M., Rolband, L. A. & Afonin, K. A. Challenges to optimizing RNA nanostructures for large scale production and controlled therapeutic properties. Nanomedicine 15, 1331–1340 (2020).
Leborgne, C. et al. IgG-cleaving endopeptidase enables in vivo gene therapy in the presence of anti-AAV neutralizing antibodies. Nat. Med. 26, 1096–1101 (2020).
Balwani, M. et al. Phase 3 trial of RNAi therapeutic givosiran for acute intermittent porphyria. N. Engl. J. Med. 382, 2289–2301 (2020).
Ray, K. K. et al. Two phase 3 trials of inclisiran in patients with elevated LDL cholesterol. N. Engl. J. Med. 382, 1507–1519 (2020).
Garrelfs, S. F. et al. Lumasiran, an RNAi therapeutic for primary hyperoxaluria type 1. N. Engl. J. Med. 384, 1216–1226 (2021).
Adams, D. et al. Patisiran, an RNAi therapeutic, for hereditary transthyretin amyloidosis. N. Engl. J. Med. 379, 11–21 (2018).
Parums, D. V. Editorial: first full regulatory approval of a COVID-19 vaccine, the BNT162b2 Pfizer-BioNTech vaccine, and the real-world implications for Public Health Policy. Med. Sci. Monit. 27, e934625 (2021).
Baden, L. R. et al. Efficacy and safety of the mRNA-1273 SARS-CoV-2 vaccine. N. Engl. J. Med. 384, 403–416 (2021).
Polack, F. P. et al. Safety and efficacy of the BNT162b2 mRNA Covid-19 vaccine. N. Engl. J. Med. 383, 2603–2615 (2020).
Buck, J., Grossen, P., Cullis, P. R., Huwyler, J. & Witzigmann, D. Lipid-based DNA therapeutics: hallmarks of non-viral gene delivery. ACS Nano 13, 3754–3782 (2019).
Vargason, A. M., Anselmo, A. C. & Mitragotri, S. The evolution of commercial drug delivery technologies. Nat. Biomed. Eng. 5, 951–967 (2021).
Watts, J. K. & Corey, D. R. Silencing disease genes in the laboratory and the clinic. J. Pathol. 226, 365–379 (2012).
Kosmas, C. E. et al. Inclisiran for the treatment of cardiovascular disease: a short review on the emerging data and therapeutic potential. Ther. Clin. Risk Manag. 16, 1031–1037 (2020).
Chen, F., Alphonse, M. & Liu, Q. Strategies for nonviral nanoparticle-based delivery of CRISPR/Cas9 therapeutics. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 12, e1609 (2020).
Hanna, J., Hossain, G. S. & Kocerha, J. The potential for microRNA therapeutics and clinical research. Front. Genet. 10, 478 (2019).
Hong, D. S. et al. Phase 1 study of MRX34, a liposomal miR-34a mimic, in patients with advanced solid tumours. Br. J. Cancer 122, 1630–1637 (2020).
van der Ree, M. H. et al. Miravirsen dosing in chronic hepatitis C patients results in decreased microRNA-122 levels without affecting other microRNAs in plasma. Aliment. Pharmacol. Ther. 43, 102–113 (2016).
van der Ree, M. H. et al. Safety, tolerability, and antiviral effect of RG-101 in patients with chronic hepatitis C: a phase 1B, double-blind, randomised controlled trial. Lancet 389, 709–717 (2017).
Regulus announces pipeline updates and advancements. Regulus http://ir.regulusrx.com/news-releases/news-release-details/regulus-announces-pipeline-updates-and-advancements (2017).
Wilson, R. C. & Doudna, J. A. Molecular mechanisms of RNA interference. Annu. Rev. Biophys. 42, 217–239 (2013).
Liu, J., Valencia-Sanchez, M. A., Hannon, G. J. & Parker, R. MicroRNA-dependent localization of targeted mRNAs to mammalian P-bodies. Nat. Cell Biol. 7, 719–723 (2005).
Alnylam announces U.S. Food and Drug Administration acceptance of new drug application for investigational vutrisiran for the treatment of the polyneuropathy of hereditary ATTR amyloidosis. Alnylam https://investors.alnylam.com/press-release?id=25811 (2021).
HELIOS-A: 9-month results from the phase 3 study of vutrisiran in patients with hereditary transthyretin-mediated amyloidosis with polyneuropathy. Alnylam https://www.alnylam.com/wp-content/uploads/2021/04/Adams_HELIOS-A-9-Month-Results.pdf (2021).
Fire, A. et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806–811 (1998).
Adachi, H., Hengesbach, M., Yu, Y. T. & Morais, P. From antisense RNA to RNA modification: therapeutic potential of RNA-based technologies. Biomedicines 9, 550 (2021).
Humphreys, S. C. et al. Emerging siRNA design principles and consequences for biotransformation and disposition in drug development. J. Med. Chem. 63, 6407–6422 (2020).
Evers, M. M., Toonen, L. J. & van Roon-Mom, W. M. Antisense oligonucleotides in therapy for neurodegenerative disorders. Adv. Drug Deliv. Rev. 87, 90–103 (2015).
Santos, R. D. et al. Mipomersen, an antisense oligonucleotide to apolipoprotein B-100, reduces lipoprotein(a) in various populations with hypercholesterolemia: results of 4 phase III trials. Arterioscler. Thromb. Vasc. Biol. 35, 689–699 (2015).
Benson, M. D. et al. Inotersen treatment for patients with hereditary transthyretin amyloidosis. N. Engl. J. Med. 379, 22–31 (2018).
Lim, K. R., Maruyama, R. & Yokota, T. Eteplirsen in the treatment of Duchenne muscular dystrophy. Drug Des. Devel Ther. 11, 533–545 (2017).
Frank, D. E. et al. Increased dystrophin production with golodirsen in patients with Duchenne muscular dystrophy. Neurology 94, e2270–e2282 (2020).
Finkel, R. S. et al. Nusinersen versus sham control in infantile-onset spinal muscular atrophy. N. Engl. J. Med. 377, 1723–1732 (2017).
Crooke, S. T. Molecular mechanisms of antisense oligonucleotides. Nucleic Acid Ther. 27, 70–77 (2017).
Lim, K. H. et al. Antisense oligonucleotide modulation of non-productive alternative splicing upregulates gene expression. Nat. Commun. 11, 3501 (2020).
Kilanowska, A. & Studzińska, S. In vivo and in vitro studies of antisense oligonucleotides — a review. RSC Adv. 10, 34501–34516 (2020).
Bennett, C. F., Baker, B. F., Pham, N., Swayze, E. & Geary, R. S. Pharmacology of antisense drugs. Annu. Rev. Pharmacol. Toxicol. 57, 81–105 (2017).
Burdick, A. D. et al. Sequence motifs associated with hepatotoxicity of locked nucleic acid — modified antisense oligonucleotides. Nucleic Acids Res. 42, 4882–4891 (2014).
Yamamoto, T. et al. Highly potent GalNAc-conjugated tiny LNA anti-miRNA-122 antisense oligonucleotides. Pharmaceutics 13, 817 (2021).
Shen, W. et al. Chemical modification of PS-ASO therapeutics reduces cellular protein-binding and improves the therapeutic index. Nat. Biotechnol. 37, 640–650 (2019).
Miller, C. M. et al. Stabilin-1 and stabilin-2 are specific receptors for the cellular internalization of phosphorothioate-modified antisense oligonucleotides (ASOs) in the liver. Nucleic Acids Res. 44, 2782–2794 (2016).
Merkle, T. et al. Precise RNA editing by recruiting endogenous ADARs with antisense oligonucleotides. Nat. Biotechnol. 37, 133–138 (2019).
Qu, L. et al. Programmable RNA editing by recruiting endogenous ADAR using engineered RNAs. Nat. Biotechnol. 37, 1059–1069 (2019).
Aquino-Jarquin, G. Novel engineered programmable systems for ADAR-mediated RNA editing. Mol. Ther. Nucleic Acids 19, 1065–1072 (2020).
Da Silva Sanchez, A., Paunovska, K., Cristian, A. & Dahlman, J. E. Treating cystic fibrosis with mRNA and CRISPR. Hum. Gene Ther. 31, 940–955 (2020).
Gillmore, J. D. et al. CRISPR-Cas9 in vivo gene editing for transthyretin amyloidosis. N. Engl. J. Med. 385, 493–502 (2021).
Musunuru, K. et al. In vivo CRISPR base editing of PCSK9 durably lowers cholesterol in primates. Nature 593, 429–434 (2021).
Rothgangl, T. et al. In vivo adenine base editing of PCSK9 in macaques reduces LDL cholesterol levels. Nat. Biotechnol. 39, 949–957 (2021).
Thompson, M. G. et al. Interim estimates of vaccine effectiveness of BNT162b2 and mRNA-1273 COVID-19 vaccines in preventing SARS-CoV-2 infection among health care personnel, first responders, and other essential and frontline workers — eight U.S. locations, December 2020-March 2021. MMWR 70, 495–500 (2021).
Dobrowolski, C., Paunovska, K., Hatit, M. Z. C., Lokugamage, M. P. & Dahlman, J. E. Therapeutic RNA delivery for COVID and other diseases. Adv. Health. Mater. 10, e2002022 (2021).
Translate Bio announces results from second interim data analysis from ongoing phase 1/2 clinical trial of MRT5005 in patients with cystic fibrosis (CF). Translate Bio https://investors.translate.bio/news-releases/news-release-details/translate-bio-announces-results-second-interim-data-analysis (2021).
Translate Bio announces pipeline program update. Translate Bio https://investors.translate.bio/news-releases/news-release-details/translate-bio-announces-pipeline-program-update (2021).
Arcturus Therapeutics announces first quarter 2021 company overview and financial results and provides new clinical data. Arcturus Therapeutics https://ir.arcturusrx.com/news-releases/news-release-details/arcturus-therapeutics-announces-first-quarter-2021-company (2021).
Krienke, C. et al. A noninflammatory mRNA vaccine for treatment of experimental autoimmune encephalomyelitis. Science 371, 145–153 (2021).
Pardi, N., Hogan, M. J., Porter, F. W. & Weissman, D. mRNA vaccines — a new era in vaccinology. Nat. Rev. Drug Discov. 17, 261–279 (2018).
Luisi, K. et al. Development of a potent Zika virus vaccine using self-amplifying messenger RNA. Sci. Adv. 6, eaba5068 (2020).
Leal, L. et al. Phase I clinical trial of an intranodally administered mRNA-based therapeutic vaccine against HIV-1 infection. AIDS 32, 2533–2545 (2018).
Feldman, R. A. et al. mRNA vaccines against H10N8 and H7N9 influenza viruses of pandemic potential are immunogenic and well tolerated in healthy adults in phase 1 randomized clinical trials. Vaccine 37, 3326–3334 (2019).
Sahin, U. et al. Personalized RNA mutanome vaccines mobilize poly-specific therapeutic immunity against cancer. Nature 547, 222–226 (2017).
Conry, R. M. et al. Characterization of a messenger RNA polynucleotide vaccine vector. Cancer Res. 55, 1397–1400 (1995).
Sahin, U. et al. An RNA vaccine drives immunity in checkpoint-inhibitor-treated melanoma. Nature 585, 107–112 (2020).
Jimeno, A. et al. Abstract CT032: A phase 1/2, open-label, multicenter, dose escalation and efficacy study of mRNA-2416, a lipid nanoparticle encapsulated mRNA encoding human OX40L, for intratumoral injection alone or in combination with durvalumab for patients with advanced malignancies. Cancer Res. 80, CT032 (2020).
Zhang, H. X., Zhang, Y. & Yin, H. Genome editing with mRNA encoding ZFN, TALEN, and Cas9. Mol. Ther. 27, 735–746 (2019).
Pardi, N. et al. Expression kinetics of nucleoside-modified mRNA delivered in lipid nanoparticles to mice by various routes. J. Controlled Rel. 217, 345–351 (2015).
Finn, J. D. et al. A single administration of CRISPR/Cas9 lipid nanoparticles achieves robust and persistent in vivo genome editing. Cell Rep. 22, 2227–2235 (2018).
Allergan and Editas Medicine announce dosing of first patient in landmark phase 1/2 clinical trial of CRISPR medicine AGN-151587 (EDIT-101) for the treatment of LCA10. Editas Medicine https://ir.editasmedicine.com/news-releases/news-release-details/allergan-and-editas-medicine-announce-dosing-first-patient (2020).
Hanlon, K. S. et al. High levels of AAV vector integration into CRISPR-induced DNA breaks. Nat. Commun. 10, 4439 (2019).
Jiang, F. & Doudna, J. A. CRISPR-Cas9 structures and mechanisms. Annu. Rev. Biophys. 46, 505–529 (2017).
Slaymaker, I. M. et al. Rationally engineered Cas9 nucleases with improved specificity. Science 351, 84–88 (2016).
Kleinstiver, B. P. et al. Engineered CRISPR-Cas9 nucleases with altered PAM specificities. Nature 523, 481–485 (2015).
Gilbert, L. A. et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 154, 442–451 (2013).
Thakore, P. I., Black, J. B., Hilton, I. B. & Gersbach, C. A. Editing the epigenome: technologies for programmable transcription and epigenetic modulation. Nat. Methods 13, 127–137 (2016).
Nuñez, J. K. et al. Genome-wide programmable transcriptional memory by CRISPR-based epigenome editing. Cell 184, 2503–2519.e2517 (2021).
Porto, E. M., Komor, A. C., Slaymaker, I. M. & Yeo, G. W. Base editing: advances and therapeutic opportunities. Nat. Rev. Drug Discov. 19, 839–859 (2020).
Mok, B. Y. et al. A bacterial cytidine deaminase toxin enables CRISPR-free mitochondrial base editing. Nature 583, 631–637 (2020).
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019).
Saito, M. et al. Dual modes of CRISPR-associated transposon homing. Cell 9, 2441–2453.e18 (2021).
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 163, 759–771 (2015).
Abudayyeh, O. O. et al. C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 353, aaf5573 (2016).
Özcan, A. et al. Programmable RNA targeting with the single-protein CRISPR effector Cas7-11. Nature 597, 720–725 (2021).
Cox, D. B. T. et al. RNA editing with CRISPR-Cas13. Science 358, 1019–1027 (2017).
Abudayyeh, O. O. et al. A cytosine deaminase for programmable single-base RNA editing. Science 365, 382–386 (2019).
Abbott, T. R. et al. Development of CRISPR as an antiviral strategy to combat SARS-CoV-2 and influenza. Cell 181, 865–876.e812 (2020).
Blanchard, E. L. et al. Treatment of influenza and SARS-CoV-2 infections via mRNA-encoded Cas13a in rodents. Nat. Biotechnol. 39, 717–726 (2021).
Miller, J. B. et al. Non-viral CRISPR/Cas gene editing in vitro and in vivo enabled by synthetic nanoparticle co-delivery of Cas9 mRNA and sgRNA. Angew. Chem. Int. Edn Engl. 56, 1059–1063 (2017).
Jiang, C. et al. A non-viral CRISPR/Cas9 delivery system for therapeutically targeting HBV DNA and pcsk9 in vivo. Cell Res. 27, 440–443 (2017).
Sago, C. D. et al. High-throughput in vivo screen of functional mRNA delivery identifies nanoparticles for endothelial cell gene editing. Proc. Natl Acad. Sci. USA 115, E9944–E9952 (2018).
Yin, H. et al. Structure-guided chemical modification of guide RNA enables potent non-viral in vivo genome editing. Nat. Biotechnol. 35, 1179–1187 (2017).
Cheng, Q. et al. Selective organ targeting (SORT) nanoparticles for tissue-specific mRNA delivery and CRISPR-Cas gene editing. Nat. Nanotechnol. 15, 313–320 (2020).
Rosenblum, D. et al. CRISPR-Cas9 genome editing using targeted lipid nanoparticles for cancer therapy. Sci. Adv. 6, eabc9450 (2020).
Zhang, X. et al. Functionalized lipid-like nanoparticles for in vivo mRNA delivery and base editing. Sci. Adv. 6, eabc2315 (2020).
Qiu, M. et al. Lipid nanoparticle-mediated codelivery of Cas9 mRNA and single-guide RNA achieves liver-specific in vivo genome editing of Angptl3. Proc. Natl Acad. Sci. USA 118, e2020401118 (2021).
Yin, H. et al. Therapeutic genome editing by combined viral and non-viral delivery of CRISPR system components in vivo. Nat. Biotechnol. 34, 328–333 (2016).
Lee, B. et al. Nanoparticle delivery of CRISPR into the brain rescues a mouse model of fragile X syndrome from exaggerated repetitive behaviours. Nat. Biomed. Eng. 2, 497–507 (2018).
Lee, K. et al. Nanoparticle delivery of Cas9 ribonucleoprotein and donor DNA in vivo induces homology-directed DNA repair. Nat. Biomed. Eng. 1, 889–901 (2017).
Wei, T., Cheng, Q., Min, Y.-L., Olson, E. N. & Siegwart, D. J. Systemic nanoparticle delivery of CRISPR-Cas9 ribonucleoproteins for effective tissue specific genome editing. Nat. Commun. 11, 3232 (2020).
Pausch, P. et al. CRISPR-CasΦ from huge phages is a hypercompact genome editor. Science 369, 333–337 (2020).
Kim, D. Y. et al. Efficient CRISPR editing with a hypercompact Cas12f1 and engineered guide RNAs delivered by adeno-associated virus. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-01009-z (2021).
Xu, X. et al. Engineered miniature CRISPR-Cas system for mammalian genome regulation and editing. Mol. Cell 81, 4333–4345.e4 (2021).
Kannan, S. et al. Compact RNA editors with small Cas13 proteins. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-01030-2 (2021).
Cheng, C. J., Tietjen, G. T., Saucier-Sawyer, J. K. & Saltzman, W. M. A holistic approach to targeting disease with polymeric nanoparticles. Nat. Rev. Drug Discov. 14, 239–247 (2015).
Israelachvili, J. N., Mitchell, D. J. & Ninham, B. W. Theory of self-assembly of lipid bilayers and vesicles. Biochim. Biophys. Acta 470, 185–201 (1977).
Kulkarni, J. A. et al. On the formation and morphology of lipid nanoparticles containing ionizable cationic lipids and siRNA. ACS Nano 12, 4787–4795 (2018).
Herrera, M., Kim, J., Eygeris, Y., Jozic, A. & Sahay, G. Illuminating endosomal escape of polymorphic lipid nanoparticles that boost mRNA delivery. Biomater. Sci. 9, 4289–4300 (2021).
Semple, S. C. et al. Rational design of cationic lipids for siRNA delivery. Nat. Biotechnol. 28, 172–176 (2010).
Altınoglu, S., Wang, M. & Xu, Q. Combinatorial library strategies for synthesis of cationic lipid-like nanoparticles and their potential medical applications. Nanomedicine 10, 643–657 (2015).
Zhang, Y., Sun, C., Wang, C., Jankovic, K. E. & Dong, Y. Lipids and lipid derivatives for RNA delivery. Chem. Rev. 121, 12181–12277 (2021).
Love, K. T. et al. Lipid-like materials for low-dose, in vivo gene silencing. Proc. Natl Acad. Sci. USA 107, 1864–1869 (2010).
Zimmermann, T. S. et al. RNAi-mediated gene silencing in non-human primates. Nature 441, 111–114 (2006).
Dong, Y. et al. Lipopeptide nanoparticles for potent and selective siRNA delivery in rodents and nonhuman primates. Proc. Natl Acad. Sci. USA 111, 3955–3960 (2014).
Jayaraman, M. et al. Maximizing the potency of siRNA lipid nanoparticles for hepatic gene silencing in vivo. Angew. Chem. Int. Edn Engl. 51, 8529–8533 (2012).
Paunovska, K. et al. Nanoparticles containing oxidized cholesterol deliver mRNA to the liver microenvironment at clinically relevant doses. Adv. Mater. 31, 1807748 (2019).
Kauffman, K. J. et al. Rapid, single-cell analysis and discovery of vectored mRNA transfection in vivo with a loxP-flanked tdtomato reporter mouse. molecular therapy. Nucleic Acids 10, 55–63 (2018).
Kauffman, K. J. et al. Optimization of lipid nanoparticle formulations for mRNA delivery in vivo with fractional factorial and definitive screening designs. Nano Lett. 15, 7300–7306 (2015).
Sedic, M. et al. Safety evaluation of lipid nanoparticle-formulated modified mRNA in the Sprague–Dawley rat and cynomolgus monkey. Vet. Pathol. 55, 341–354 (2018).
ModernaTx. Compounds and compositions for intracellular delivery of therapeutic agents. US patent US20170210697A1 (2021).
Sabatine, M. S. et al. Evolocumab and clinical outcomes in patients with cardiovascular disease. N. Engl. J. Med. 376, 1713–1722 (2017).
Beam Therapeutics announces updated preclinical data highlighting optimized LNP delivery approaches for in vivo base editing to the liver and other tissues. Beam Therapeutics https://investors.beamtx.com/news-releases/news-release-details/beam-therapeutics-announces-updated-preclinical-data (2021).
Kulkarni, J. A., Cullis, P. R. & van der Meel, R. Lipid nanoparticles enabling gene therapies: from concepts to clinical utility. Nucleic Acid. Ther. 28, 146–157 (2018).
Cheng, X. & Lee, R. J. The role of helper lipids in lipid nanoparticles (LNPs) designed for oligonucleotide delivery. Adv. Drug Deliv. Rev. 99, 129–137 (2016).
Dahlman, J. E. et al. In vivo endothelial siRNA delivery using polymeric nanoparticles with low molecular weight. Nat. Nano 9, 648–655 (2014).
Khan, O. F. et al. Endothelial siRNA delivery in nonhuman primates using ionizable low-molecular weight polymeric nanoparticles. Sci. Adv. 4, eaar8409 (2018).
Sago, C. D. et al. Nanoparticles that deliver RNA to bone marrow identified by in vivo directed evolution. J. Am. Chem. Soc. 140, 17095–17105 (2018).
Paunovska, K. et al. Analyzing 2000 in vivo drug delivery data points reveals cholesterol structure impacts nanoparticle delivery. ACS Nano 12, 8341–8349 (2018).
Lokugamage, M. P. et al. Optimization of lipid nanoparticles for the delivery of nebulized therapeutic mRNA to the lungs. Nat. Biomed. Eng. 5, 1059–1068 (2021).
Mui, B. L. et al. Influence of polyethylene glycol lipid desorption rates on pharmacokinetics and pharmacodynamics of siRNA lipid nanoparticles. Mol. Ther. Nucleic acids 2, e139 (2013).
Ryals, R. C. et al. The effects of PEGylation on LNP based mRNA delivery to the eye. PLoS ONE 15, e0241006 (2020).
Suk, J. S., Xu, Q., Kim, N., Hanes, J. & Ensign, L. M. PEGylation as a strategy for improving nanoparticle-based drug and gene delivery. Adv. Drug Deliv. Rev. 99, 28–51 (2016).
Eygeris, Y., Patel, S., Jozic, A. & Sahay, G. Deconvoluting lipid nanoparticle structure for messenger RNA delivery. Nano Lett. 20, 4543–4549 (2020).
Kranz, L. M. et al. Systemic RNA delivery to dendritic cells exploits antiviral defence for cancer immunotherapy. Nature 534, 396–401 (2016).
Intellia Therapeutics presents preclinical proof of concept for CRISPR-based in vivo editing of bone marrow at Keystone eSymposium. Intellia Therapeutics https://ir.intelliatx.com/news-releases/news-release-details/intellia-therapeutics-presents-preclinical-proof-concept-crispr (2021).
Rai, R., Alwani, S. & Badea, I. Polymeric nanoparticles in gene therapy: new avenues of design and optimization for delivery applications. Polymers 11, 745 (2019).
Kamaly, N., Yameen, B., Wu, J. & Farokhzad, O. C. Degradable controlled-release polymers and polymeric nanoparticles: mechanisms of controlling drug release. Chem. Rev. 116, 2602–2663 (2016).
Crucho, C. I. C. & Barros, M. T. Polymeric nanoparticles: A study on the preparation variables and characterization methods. Mater. Sci. Eng. C 80, 771–784 (2017).
Zhong, H., Chan, G., Hu, Y., Hu, H. & Ouyang, D. A comprehensive map of FDA-approved pharmaceutical products. Pharmaceutics 10, 263 (2018).
Xiao, B. et al. Combination therapy for ulcerative colitis: orally targeted nanoparticles prevent mucosal damage and relieve inflammation. Theranostics 6, 2250–2266 (2016).
Harada-Shiba, M. et al. Polyion complex micelles as vectors in gene therapy — pharmacokinetics and in vivo gene transfer. Gene Ther. 9, 407–414 (2002).
Ewe, A. et al. Optimized polyethylenimine (PEI)-based nanoparticles for siRNA delivery, analyzed in vitro and in an ex vivo tumor tissue slice culture model. Drug Deliv. Transl. Res. 7, 206–216 (2017).
Gao, X. et al. The association of autophagy with polyethylenimine-induced cytotoxicity in nephritic and hepatic cell lines. Biomaterials 32, 8613–8625 (2011).
Breunig, M., Lungwitz, U., Liebl, R. & Goepferich, A. Breaking up the correlation between efficacy and toxicity for nonviral gene delivery. Proc. Natl Acad. Sci. USA 104, 14454–14459 (2007).
Ke, X. et al. Surface-functionalized PEGylated nanoparticles deliver messenger RNA to pulmonary immune cells. ACS Appl. Mater. Interf. 12, 35835–35844 (2020).
Tan, L. et al. Optimization of an mRNA vaccine assisted with cyclodextrin–polyethyleneimine conjugates. Drug. Deliv. Transl. Res. 10, 678–689 (2020).
Xiang, J. J. et al. IONP-PLL: a novel non-viral vector for efficient gene delivery. J. Gene Med. 5, 803–817 (2003).
Yin, H. et al. Non-viral vectors for gene-based therapy. Nat. Rev. Genet. 15, 541–555 (2014).
Choi, J. et al. Nonviral polymeric nanoparticles for gene therapy in pediatric CNS malignancies. Nanomedicine 23, 102115 (2020).
Akinc, A., Lynn, D. M., Anderson, D. G. & Langer, R. Parallel synthesis and biophysical characterization of a degradable polymer library for gene delivery. J. Am. Chem. Soc. 125, 5316–5323 (2003).
Green, J. J., Langer, R. & Anderson, D. G. A combinatorial polymer library approach yields insight into nonviral gene delivery. Acc. Chem. Res. 41, 749–759 (2008).
Vandenbroucke, R. E. et al. Prolonged gene silencing in hepatoma cells and primary hepatocytes after small interfering RNA delivery with biodegradable poly(beta-amino esters). J. Gene Med. 10, 783–794 (2008).
Anderson, D. G., Lynn, D. M. & Langer, R. Semi-automated synthesis and screening of a large library of degradable cationic polymers for gene delivery. Angew. Chem. Int. Edn Engl. 42, 3153–3158 (2003).
Anderson, D. G., Akinc, A., Hossain, N. & Langer, R. Structure/property studies of polymeric gene delivery using a library of poly(beta-amino esters). Mol. Ther. 11, 426–434 (2005).
Mastorakos, P. et al. Highly compacted biodegradable DNA nanoparticles capable of overcoming the mucus barrier for inhaled lung gene therapy. Proc. Natl Acad. Sci. USA 112, 8720–8725 (2015).
Su, X., Fricke, J., Kavanagh, D. G. & Irvine, D. J. In vitro and in vivo mRNA delivery using lipid-enveloped pH-responsive polymer nanoparticles. Mol. Pharm. 8, 774–787 (2011).
Kozielski, K. L. et al. Cancer-selective nanoparticles for combinatorial siRNA delivery to primary human GBM in vitro and in vivo. Biomaterials 209, 79–87 (2019).
Eltoukhy, A. A., Chen, D., Alabi, C. A., Langer, R. & Anderson, D. G. Degradable terpolymers with alkyl side chains demonstrate enhanced gene delivery potency and nanoparticle stability. Adv. Mater. 25, 1487–1493 (2013).
Kaczmarek, J. C. et al. Polymer–lipid nanoparticles for systemic delivery of mRNA to the lungs. Angew. Chem. Int. Edn Engl. 55, 13808–13812 (2016).
Xu, L., Zhang, H. & Wu, Y. Dendrimer advances for the central nervous system delivery of therapeutics. ACS Chem. Neurosci. 5, 2–13 (2014).
Chahal, J. S. et al. Dendrimer-RNA nanoparticles generate protective immunity against lethal Ebola, H1N1 influenza, and Toxoplasma gondii challenges with a single dose. Proc. Natl Acad. Sci. USA 113, E4133–E4142 (2016).
Khan, O. F. et al. Ionizable amphiphilic dendrimer-based nanomaterials with alkyl-chain-substituted amines for tunable siRNA delivery to the liver endothelium in vivo. Angew. Chem. Int. Edn Engl. 53, 14397–14401 (2014).
Bielinska, A. U., Kukowska-Latallo, J. F. & Baker, J. R. Jr The interaction of plasmid DNA with polyamidoamine dendrimers: mechanism of complex formation and analysis of alterations induced in nuclease sensitivity and transcriptional activity of the complexed DNA. Biochim. Biophys. Acta 1353, 180–190 (1997).
Sonawane, N. D., Szoka, F. C. Jr & Verkman, A. S. Chloride accumulation and swelling in endosomes enhances DNA transfer by polyamine-DNA polyplexes. J. Biol. Chem. 278, 44826–44831 (2003).
Yoo, J., Park, C., Yi, G., Lee, D. & Koo, H. Active targeting strategies using biological ligands for nanoparticle drug delivery systems. Cancers 11, 640 (2019).
Nel, A. E. et al. Understanding biophysicochemical interactions at the nano-bio interface. Nat. Mater. 8, 543–557 (2009).
Dawson, K. A. & Yan, Y. Current understanding of biological identity at the nanoscale and future prospects. Nat. Nanotechnol. 16, 229–242 (2021).
Schöttler, S. et al. Protein adsorption is required for stealth effect of poly(ethylene glycol)- and poly(phosphoester)-coated nanocarriers. Nat. Nanotechnol. 11, 372–377 (2016).
Salvati, A. et al. Transferrin-functionalized nanoparticles lose their targeting capabilities when a biomolecule corona adsorbs on the surface. Nat. Nanotechnol. 8, 137–143 (2013).
Akinc, A. et al. The Onpattro story and the clinical translation of nanomedicines containing nucleic acid-based drugs. Nat. Nanotechnol. 14, 1084–1087 (2019).
Akinc, A. et al. Targeted delivery of RNAi therapeutics with endogenous and exogenous ligand-based mechanisms. Mol. Ther. 18, 1357–1364 (2010).
Miao, L. et al. Synergistic lipid compositions for albumin receptor mediated delivery of mRNA to the liver. Nat. Commun. 11, 2424 (2020).
Sago, C. D. et al. Modifying a commonly expressed endocytic receptor retargets nanoparticles in vivo. Nano Lett. 18, 7590–7600 (2018).
Chen, S. et al. Influence of particle size on the in vivo potency of lipid nanoparticle formulations of siRNA. J. Control. Rel. 235, 236–244 (2016).
Nakamura, T. et al. The effect of size and charge of lipid nanoparticles prepared by microfluidic mixing on their lymph node transitivity and distribution. Mol. Pharm. 17, 944–953 (2020).
Reinhard, K. et al. An RNA vaccine drives expansion and efficacy of claudin-CAR-T cells against solid tumors. Science 367, 446–453 (2020).
Nair, J. K. et al. Multivalent N-acetylgalactosamine-conjugated siRNA localizes in hepatocytes and elicits robust RNAi-mediated gene silencing. J. Am. Chem. Soc. 136, 16958–16961 (2014).
Prakash, T. P. et al. Targeted delivery of antisense oligonucleotides to hepatocytes using triantennary N-acetyl galactosamine improves potency 10-fold in mice. Nucleic Acids Res. 42, 8796–8807 (2014).
Agarwal, S. et al. Impact of serum proteins on the uptake and RNA interference activity of N-acetylgalactosamine-conjugated small interfering RNAs. Nucleic Acid Ther. 31, 309–315 (2021).
Foster, D. J. et al. Advanced siRNA designs further improve in vivo performance of GalNAc-siRNA conjugates. Mol. Ther. 26, 708–717 (2018).
Nair, J. K. et al. Impact of enhanced metabolic stability on pharmacokinetics and pharmacodynamics of GalNAc-siRNA conjugates. Nucleic Acids Res. 45, 10969–10977 (2017).
Zanardi, T. A. et al. Safety, pharmacokinetic, and pharmacodynamic evaluation of a 2′-(2-methoxyethyl)-d-ribose antisense oligonucleotide-triantenarry N-acetyl-galactosamine conjugate that targets the human transmembrane protease serine 6. J. Pharmacol. Exp. Ther. 377, 51–63 (2021).
Janas, M. M. et al. The nonclinical safety profile of GalNAc-conjugated RNAi therapeutics in subacute studies. Toxicol. Pathol. 46, 735–745 (2018).
Biscans, A. et al. Diverse lipid conjugates for functional extra-hepatic siRNA delivery in vivo. Nucleic Acids Res. 47, 1082–1096 (2019).
Osborn, M. F. et al. Hydrophobicity drives the systemic distribution of lipid-conjugated siRNAs via lipid transport pathways. Nucleic Acids Res. 47, 1070–1081 (2019).
Nagata, T. et al. Cholesterol-functionalized DNA/RNA heteroduplexes cross the blood–brain barrier and knock down genes in the rodent CNS. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-00972-x (2021).
Zhou, J. & Rossi, J. Aptamers as targeted therapeutics: current potential and challenges. Nat. Rev. Drug Discov. 16, 181–202 (2017).
Yoon, S., Wu, X., Armstrong, B., Habib, N. & Rossi, J. J. An RNA aptamer targeting the receptor tyrosine kinase PDGFRα induces anti-tumor effects through STAT3 and p53 in glioblastoma. Mol. Ther. Nucleic Acids 14, 131–141 (2019).
Sugo, T. et al. Development of antibody-siRNA conjugate targeted to cardiac and skeletal muscles. J. Controlled Rel. 237, 1–13 (2016).
Avidity corporate presentation. Avidity Biosciences https://aviditybiosciences.investorroom.com/events-and-presentations (2021).
Kedmi, R. et al. A modular platform for targeted RNAi therapeutics. Nat. Nanotechnol. 13, 214–219 (2018).
Veiga, N. et al. Cell specific delivery of modified mRNA expressing therapeutic proteins to leukocytes. Nat. Commun. 9, 4493 (2018).
Dammes, N. et al. Conformation-sensitive targeting of lipid nanoparticles for RNA therapeutics. Nat. Nanotechnol. https://doi.org/10.1038/s41565-021-00928-x (2021).
Li, Q. et al. Engineering caveolae-targeted lipid nanoparticles to deliver mRNA to the lungs. ACS Chem. Biol. 15, 830–836 (2020).
Zhuang, X. et al. mRNA vaccines encoding the HA protein of influenza A H1N1 virus delivered by cationic lipid nanoparticles induce protective immune responses in mice. Vaccines 8, 123 (2020).
Paunovska, K. et al. A direct comparison of in vitro and in vivo nucleic acid delivery mediated by hundreds of nanoparticles reveals a weak correlation. Nano Lett. 18, 2148–2157 (2018).
Paunovska, K., Loughrey, D., Sago, C. D., Langer, R. & Dahlman, J. E. Using large datasets to understand nanotechnology. Adv. Mater. 31, e1902798 (2019).
Lokugamage, M. P., Sago, C. D. & Dahlman, J. E. Testing thousands of nanoparticles in vivo using DNA barcodes. Curr. Opin. Biomed. Eng. 7, 1–8 (2018).
Yaari, Z. et al. Theranostic barcoded nanoparticles for personalized cancer medicine. Nat. Commun. 7, 13325 (2016).
Dahlman, J. E. et al. Barcoded nanoparticles for high throughput in vivo discovery of targeted therapeutics. Proc. Natl Acad. Sci. USA 114, 2060–2065 (2017).
Lokugamage, M. P., Sago, C. D., Gan, Z., Krupczak, B. R. & Dahlman, J. E. Constrained nanoparticles deliver siRNA and sgRNA to T cells in vivo without targeting ligands. Adv. Mater. 31, e1902251 (2019).
Lokugamage, M. P. et al. Mild innate immune activation overrides efficient nanoparticle-mediated RNA delivery. Adv. Mater. 32, 1904905 (2019).
Riley, R. S. et al. Ionizable lipid nanoparticles for in utero mRNA delivery. Sci. Adv. 7, eaba1028 (2021).
Havel, P. J., Kievit, P., Comuzzie, A. G. & Bremer, A. A. Use and importance of nonhuman primates in metabolic disease research: current state of the field. ILAR J. 58, 251–268 (2017).
Paunovska, K. et al. Increased PIP3 activity blocks nanoparticle mRNA delivery. Sci. Adv. 6, eaba5672 (2020).
Li, R. et al. Therapeutically reprogrammed nutrient signalling enhances nanoparticulate albumin bound drug uptake and efficacy in KRAS-mutant cancer. Nat. Nanotechnol. 16, 830–839 (2021).
Patel, S. et al. Boosting intracellular delivery of lipid nanoparticle-encapsulated mRNA. Nano Lett. 17, 5711–5718 (2017).
Yin, W. et al. Plasma lipid profiling across species for the identification of optimal animal models of human dyslipidemia. J. Lipid Res. 53, 51–65 (2012).
Rampado, R., Crotti, S., Caliceti, P., Pucciarelli, S. & Agostini, M. Recent advances in understanding the protein corona of nanoparticles and in the formulation of “stealthy” nanomaterials. Front. Bioeng. Biotechnol. 8, 166 (2020).
Delprato, A. et al. Systems genetic analysis of hippocampal neuroanatomy and spatial learning in mice. Genes Brain Behav. 14, 591–606 (2015).
Harrill, A. H. et al. A mouse diversity panel approach reveals the potential for clinical kidney injury due to DB289 not predicted by classical rodent models. Toxicol. Sci. 130, 416–426 (2012).
Church, R. J. et al. A systems biology approach utilizing a mouse diversity panel identifies genetic differences influencing isoniazid-induced microvesicular steatosis. Toxicol. Sci. 140, 481–492 (2014).
Leist, S. R. et al. Influenza H3N2 infection of the collaborative cross founder strains reveals highly divergent host responses and identifies a unique phenotype in CAST/EiJ mice. BMC Genomics 17, 143 (2016).
Jaxpheno2 project protocol: morphometric (organ weight) survey of 11 strains of mice (2006). Mouse Phenome Database at the Jackson Laboratory https://phenome.jax.org/projects/Jaxpheno2/protocol?method=organ+weights (2006).
Sugimoto, K. et al. Background data on organ weights and histopathological lesions in Cej:CD(SD)IGS rats for 4-, 13- and 26-weeks repeated-dose toxicity studies. Biological reference data on CD(SD)IGS rats. In IGS Databook 2000 79–87 (Charles River Laboratory, 2000).
Durbin, P. W., Jeung, N., Williams, M. H., Kullgren, B. & Parrott, M. W. Weights of bones and tissues at maturity and growth of the skeleton of rhesus (Macaca mullata and cynomolgus (Macaca fascicularis) monkeys. escholarship https://escholarship.org/content/qt6kw7682s/qt6kw7682s.pdf (1996).
Molina, D. K. & DiMaio, V. J. Normal organ weights in men. Part II — the brain, lungs, liver, spleen, and kidneys. Am. J. Forensic Med. Pathol. 33, 368–372 (2012).
Molina, D. K. & DiMaio, V. J. Normal organ weights in women. Part II — the brain, lungs, liver, spleen, and kidneys. Am. J. Forensic Med. Pathol. 36, 182–187 (2015).
Molina, D. K. & DiMaio, V. J. Normal organ weights in women. Part I — the heart. Am. J. Forensic Med. Pathol. 36, 176–181 (2015).
Molina, D. K. & DiMaio, V. J. Normal organ weights in men. Part I — the heart. Am. J. Forensic Med. Pathol. 33, 362–367 (2012).
Hatit, M. Z. C. et al. Species-dependent in vivo mRNA delivery and cellular responses to nanoparticles. Nat. Nanotechnol. https://doi.org/10.1038/s41565-021-01030-y (2021).
Zhang, X., Goel, V. & Robbie, G. J. Pharmacokinetics of patisiran, the first approved RNA interference therapy in patients with hereditary transthyretin-mediated amyloidosis. J. Clin. Pharmacol. 60, 573–585 (2019).
Zhang, X. et al. Patisiran pharmacokinetics, pharmacodynamics, and exposure-response analyses in the phase 3 APOLLO trial in patients with hereditary transthyretin-mediated (hATTR) amyloidosis. J. Clin. Pharmacol. 60, 37–49 (2020).
Center for Drug Evaluation and Research application number: 210922Orig1s000. FDA https://www.accessdata.fda.gov/drugsatfda_docs/nda/2018/210922Orig1s000MultiR.pdf (2018).
Maier, M. A. et al. Biodegradable lipids enabling rapidly eliminated lipid nanoparticles for systemic delivery of RNAi therapeutics. Mol. Ther. 21, 1570–1578 (2013).
Cheng, Z., Al Zaki, A., Hui, J. Z., Muzykantov, V. R. & Tsourkas, A. Multifunctional nanoparticles: cost versus benefit of adding targeting and imaging capabilities. Science 338, 903–910 (2012).
Gilleron, J. et al. Image-based analysis of lipid nanoparticle-mediated siRNA delivery, intracellular trafficking and endosomal escape. Nat. Biotechnol. 31, 638–646 (2013).
Wittrup, A. et al. Visualizing lipid-formulated siRNA release from endosomes and target gene knockdown. Nat. Biotechnol. 33, 870–876 (2015).
Alberer, M. et al. Safety and immunogenicity of a mRNA rabies vaccine in healthy adults: an open-label, non-randomised, prospective, first-in-human phase 1 clinical trial. Lancet 390, 1511–1520 (2017).
Zhao, P. et al. Long-term storage of lipid-like nanoparticles for mRNA delivery. Bioact. Mater. 5, 358–363 (2020).
Gerhardt, A. et al. A thermostable, flexible RNA vaccine delivery platform for pandemic response. Preprint at bioRxiv https://doi.org/10.1101/2021.02.01.429283 (2021).
Besin, G. et al. Accelerated blood clearance of lipid nanoparticles entails a biphasic humoral response of B-1 followed by B-2 lymphocytes to distinct antigenic moieties. Immunohorizons 3, 282–293 (2019).
Machin, N. & Ragni, M. V. An investigational RNAi therapeutic targeting antithrombin for the treatment of hemophilia A and B. J. Blood Med. 9, 135–140 (2018).
Habtemariam, B. A. et al. Single-dose pharmacokinetics and pharmacodynamics of transthyretin targeting N-acetylgalactosamine–small interfering ribonucleic acid conjugate, vutrisiran, in healthy subjects. Clin. Pharmacol. Ther. 109, 372–382 (2021).
Wang, Y., Yu, R. Z., Henry, S. & Geary, R. S. Pharmacokinetics and clinical pharmacology considerations of GalNAc(3)-conjugated antisense oligonucleotides. Expert Opin. Drug Metab. Toxicol. 15, 475–485 (2019).
Ferguson, C. M., Echeverria, D., Hassler, M., Ly, S. & Khvorova, A. Cell type impacts accessibility of mRNA to silencing by RNA interference. Mol. Ther. Nucleic Acids 21, 384–393 (2020).
Segel, M. et al. Mammalian retrovirus-like protein PEG10 packages its own mRNA and can be pseudotyped for mRNA delivery. Science 373, 882 (2021).
Herrmann, I. K., Wood, M. J. A. & Fuhrmann, G. Extracellular vesicles as a next-generation drug delivery platform. Nat. Nanotechnol. 16, 748–759 (2021).
Dhuri, K. et al. Antisense oligonucleotides: an emerging area in drug discovery and development. J. Clin. Med. 9, 2004 (2020)
Hung, G. et al. Characterization of target mRNA reduction through in situ RNA hybridization in multiple organ systems following systemic antisense treatment in animals. Nucleic Acid. Ther. 23, 369–378 (2013).
CureVac provides update on phase 2b/3 trial of first-generation COVID-19 vaccine candidate, CVnCoV. CureVac https://www.curevac.com/en/2021/06/16/curevac-provides-update-on-phase-2b-3-trial-of-first-generation-covid-19-vaccine-candidate-cvncov/ (2021).
McKenzie, L. K., El-Khoury, R., Thorpe, J. D., Damha, M. J. & Hollenstein, M. Recent progress in non-native nucleic acid modifications. Chem. Soc. Rev. 50, 5126–5164 (2021).
Orlandini von Niessen, A. G. et al. Improving mRNA-based therapeutic gene delivery by expression-augmenting 3′ UTRs identified by cellular library screening. Mol. Ther. 27, 824–836 (2019).
Jain, R. et al. MicroRNAs enable mRNA therapeutics to selectively program cancer cells to self-destruct. Nucleic Acid Ther. 28, 285–296 (2018).
Hrkach, J. et al. Preclinical development and clinical translation of a PSMA-targeted docetaxel nanoparticle with a differentiated pharmacological profile. Sci. Transl. Med. 4, 128ra139 (2012).
Chen, Z. et al. A murine lung cancer co-clinical trial identifies genetic modifiers of therapeutic response. Nature 483, 613–617 (2012).
Kojima, N., Turner, I. & Klausner, J. D. The Covid-19 vaccine-development multiverse. N. Engl. J. Med. 384, 681–682 (2021).
Levine-Tiefenbrun, M. et al. Initial report of decreased SARS-CoV-2 viral load after inoculation with the BNT162b2 vaccine. Nat. Med. 27, 790–792 (2021).
Dagan, N. et al. BNT162b2 mRNA Covid-19 vaccine in a nationwide mass vaccination setting. N. Engl. J. Med. 384, 1412–1423 (2021).
Lindsay, K. E. et al. Visualization of early events in mRNA vaccine delivery in non-human primates via PET-CT and near-infrared imaging. Nat. Biomed. Eng. 3, 371–380 (2019).
Pardi, N. et al. Nucleoside-modified mRNA vaccines induce potent T follicular helper and germinal center B cell responses. J. Exp. Med. 215, 1571–1588 (2018).
Laczkó, D. et al. A single immunization with nucleoside-modified mRNA vaccines elicits strong cellular and humoral immune responses against SARS-CoV-2 in mice. Immunity 53, 724–732.e727 (2020).
Wu, K. et al. Serum neutralizing activity elicited by mRNA-1273 vaccine. N. Engl. J. Med. https://doi.org/10.1056/NEJMc2102179 (2021).
Callaway, E. & Ledford, H. How to redesign COVID vaccines so they protect against variants. Nature 590, 15–16 (2021).
Hacisuleyman, E. et al. Vaccine breakthrough infections with SARS-CoV-2 variants. N. Engl. J. Med. https://doi.org/10.1056/NEJMoa2105000 (2021).
Han, B. A., Kramer, A. M. & Drake, J. M. Global patterns of zoonotic disease in mammals. Trends Parasitol. 32, 565–577 (2016).
Acknowledgements
The authors thank K. Tiegren at the Georgia Institute of Technology for copyediting the manuscript.
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding author
Ethics declarations
Competing interests
J.E.D. is a consultant for GV. The other authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Genetics thanks Yizhou Dong and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Paunovska, K., Loughrey, D. & Dahlman, J.E. Drug delivery systems for RNA therapeutics. Nat Rev Genet 23, 265–280 (2022). https://doi.org/10.1038/s41576-021-00439-4
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41576-021-00439-4
This article is cited by
-
tRNA-derived small RNAs in human cancers: roles, mechanisms, and clinical application
Molecular Cancer (2024)
-
Recent progress of non-linear topological structure polymers: synthesis, and gene delivery
Journal of Nanobiotechnology (2024)
-
Breaking the mold with RNA—a “RNAissance” of life science
npj Genomic Medicine (2024)
-
Light-field flow cytometry for high-resolution, volumetric and multiparametric 3D single-cell analysis
Nature Communications (2024)
-
Genetics of human brain development
Nature Reviews Genetics (2024)