Abstract
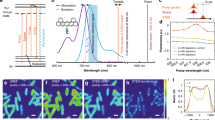
Bioimaging harnessing optical contrasts and chemical specificity is of vital importance in probing complex biology. Vibrational spectroscopy based on mid-infrared excitation can reveal rich chemical information about molecular distributions. However, its full potential for bioimaging is hindered by the achievable sensitivity. Here we report bond-selective fluorescence-detected infrared-excited (BonFIRE) spectro-microscopy. BonFIRE employs two-photon excitation in the mid- and near-infrared to upconvert vibrational excitations to electronic states for fluorescence detection, thus encoding vibrational information into fluorescence. The system utilizes tunable narrowband picosecond pulses to ensure high sensitivity, biocompatibility and robustness for bond-selective biological interrogations over a wide spectrum of reporter molecules. We demonstrate BonFIRE spectral imaging in both fingerprint and cell-silent spectroscopic windows with single-molecule sensitivity for common fluorescent dyes. We then demonstrate BonFIRE imaging on various intracellular targets in fixed and live cells, neurons and tissues, with promise for further vibrational multiplexing. For dynamic bioanalysis in living systems, we implement a high-frequency modulation scheme and demonstrate time-lapse BonFIRE microscopy of live HeLa cells. We expect BonFIRE to expand the bioimaging toolbox by providing a new level of bond-specific vibrational information and facilitate functional imaging and sensing for biological investigations.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The minimum dataset necessary to reproduce the results is available from the corresponding author.
Code availability
The codes for experimental data acquisition, physical model simulation and Franck–Condon factor calculation are available from the corresponding author.
References
Dean, K. M. & Palmer, A. E. Advances in fluorescence labeling strategies for dynamic cellular imaging. Nat. Chem. Biol. 10, 512–523 (2014).
Möckl, L. & Moerner, W. E. Super-resolution microscopy with single molecules in biology and beyond—essentials, current trends and future challenges. J. Am. Chem. Soc. 142, 17828–17844 (2020).
Wu, J., Ji, N. & Tsia, K. K. Speed scaling in multiphoton fluorescence microscopy. Nat. Photonics 15, 800–812 (2021).
Cheng, J.-X. & Xie, X. S. Vibrational spectroscopic imaging of living systems: an emerging platform for biology and medicine. Science 350, aaa8870 (2015).
Wei, L. et al. Super-multiplex vibrational imaging. Nature 544, 465–470 (2017).
Bhargava, R. Infrared spectroscopic imaging: the next generation. Appl. Spectrosc. 66, 1091–1120 (2012).
Ma, J., Pazos, I. M., Zhang, W., Culik, R. M. & Gai, F. Site-specific infrared probes of proteins. Annu. Rev. Phys. Chem. 66, 357–377 (2015).
Ostrander, J. S., Serrano, A. L., Ghosh, A. & Zanni, M. T. Spatially resolved two-dimensional infrared spectroscopy via wide-field microscopy. ACS Photonics 3, 1315–1323 (2016).
Shi, L. et al. Mid-infrared metabolic imaging with vibrational probes. Nat. Methods 17, 844–851 (2020).
Dazzi, A. & Prater, C. B. AFM-IR: technology and applications in nanoscale infrared spectroscopy and chemical imaging. Chem. Rev. 117, 5146–5173 (2017).
Pleitez, M. A. et al. Label-free metabolic imaging by mid-infrared optoacoustic microscopy in living cells. Nat. Biotechnol. 38, 293–296 (2020).
Bai, Y., Yin, J. & Cheng, J.-X. Bond-selective imaging by optically sensing the mid-infrared photothermal effect. Sci. Adv. 7, eabg1559 (2021).
Zhang, D. et al. Depth-resolved mid-infrared photothermal imaging of living cells and organisms with submicrometer spatial resolution. Sci. Adv. 2, e1600521 (2016).
Li, Z., Aleshire, K., Kuno, M. & Hartland, G. V. Super-resolution far-field infrared imaging by photothermal heterodyne imaging. J. Phys. Chem. B 121, 8838–8846 (2017).
Lim, J. M. et al. Cytoplasmic protein imaging with mid-infrared photothermal microscopy: cellular dynamics of live neurons and oligodendrocytes. J. Phys. Chem. Lett. 10, 2857–2861 (2019).
Schnell, M. et al. All-digital histopathology by infrared-optical hybrid microscopy. Proc. Natl Acad. Sci. USA 117, 3388–3396 (2020).
Shi, J. et al. High-resolution, high-contrast mid-infrared imaging of fresh biological samples with ultraviolet-localized photoacoustic microscopy. Nat. Photonics 13, 609–615 (2019).
Ruggeri, F. S., Mannini, B., Schmid, R., Vendruscolo, M. & Knowles, T. P. J. Single molecule secondary structure determination of proteins through infrared absorption nanospectroscopy. Nat. Commun. 11, 2945 (2020).
Laubereau, A., Seilmeier, A. & Kaiser, W. A new technique to measure ultrashort vibrational relaxation times in liquid systems. Chem. Phys. Lett. 36, 232–237 (1975).
Kryukov, P. G., Letokhov, V. S., Matveets, Y. A., Nikogosyan, D. N. & Sharkov, A. V. Selective two-stage excitation of an electronic state of organic molecules in aqueous solution by picosecond light pulse. Sov. J. Quantum Electron. 8, 1405–1407 (1978).
Sakai, M., Kawashima, Y., Takeda, A., Ohmori, T. & Fujii, M. Far-field infrared super-resolution microscopy using picosecond time-resolved transient fluorescence detected IR spectroscopy. Chem. Phys. Lett. 439, 171–176 (2007).
Whaley-Mayda, L., Guha, A., Penwell, S. B. & Tokmakoff, A. Fluorescence-encoded infrared vibrational spectroscopy with single-molecule sensitivity. J. Am. Chem. Soc. 143, 3060–3064 (2021).
Whaley-Mayda, L., Guha, A. & Tokmakoff, A. Resonance conditions, detection quality and single-molecule sensitivity in fluorescence-encoded infrared vibrational spectroscopy. J. Chem. Phys. 156, 174202 (2022).
Hopt, A. & Neher, E. Highly nonlinear photodamage in two-photon fluorescence microscopy. Biophys. J. 80, 2029–2036 (2001).
Talone, B. et al. Phototoxicity induced in living HeLa cells by focused femtosecond laser pulses: a data-driven approach. Biomed. Opt. Express 12, 7886–7905 (2021).
Xiong, H. et al. Stimulated Raman excited fluorescence spectroscopy and imaging. Nat. Photonics 13, 412–417 (2019).
Xiong, H., Qian, N., Miao, Y., Zhao, Z. & Min, W. Stimulated Raman excited fluorescence spectroscopy of visible dyes. J. Phys. Chem. Lett. 10, 3563–3570 (2019).
Kurochkin, D. V., Naraharisetty, S. R. G. & Rubtsov, I. V. A relaxation-assisted 2D IR spectroscopy method. Proc. Natl Acad. Sci. USA 104, 14209–14214 (2007).
Zhang, Y. et al. Fluorescence-detected mid-infrared photothermal microscopy. J. Am. Chem. Soc. 143, 11490–11499 (2021).
Li, M. et al. Fluorescence-detected mid-infrared photothermal microscopy. J. Am. Chem. Soc. 143, 10809–10815 (2021).
Watanabe, H., Hayazawa, N., Inouye, Y. & Kawata, S. DFT vibrational calculations of rhodamine 6G adsorbed on silver: analysis of tip-enhanced Raman spectroscopy. J. Phys. Chem. B 109, 5012–5020 (2005).
Hübner, H. J., Wörner, M., Kaiser, W. & Seilmeier, A. Subpicosecond vibrational relaxation of skeletal modes in polyatomic molecules. Chem. Phys. Lett. 182, 315–320 (1991).
Du, J., Wang, H. & Wei, L. Bringing vibrational imaging to chemical biology with molecular probes. ACS Chem. Biol. 17, 1621–1637 (2022).
Levinson, N. M. & Boxer, S. G. A conserved water-mediated hydrogen bond network defines bosutinib’s kinase selectivity. Nat. Chem. Biol. 10, 127–132 (2014).
Suydam, I. T., Snow, C. D., Pande, V. S. & Boxer, S. G. Electric fields at the active site of an enzyme: direct comparison of experiment with theory. Science 313, 200–204 (2006).
Seilmeier, A., Kaiser, W. & Laubereau, A. Vibrational combination states of polyatomic molecules investigated by ultrashort two-pulse spectroscopy. Opt. Commun. 26, 441–445 (1978).
Maier, J. P., Seilmeier, A. & Kaiser, W. Population lifetime of CH-stretching modes in medium-size molecules. Chem. Phys. Lett. 70, 591–596 (1980).
Clark, J. L., Miller, P. F. & Rumbles, G. Red edge photophysics of ethanolic rhodamine 101 and the observation of laser cooling in the condensed phase. J. Phys. Chem. A 102, 4428–4437 (1998).
Egner, A. et al. Fluorescence nanoscopy in whole cells by asynchronous localization of photoswitching emitters. Biophys. J. 93, 3285–3290 (2007).
Dempsey, G. T., Vaughan, J. C., Chen, K. H., Bates, M. & Zhuang, X. Evaluation of fluorophores for optimal performance in localization-based super-resolution imaging. Nat. Methods 8, 1027–1036 (2011).
Gaudioso, J. & Ho, W. Single-molecule vibrations, conformational changes and electronic conductivity of five-membered heterocycles. J. Am. Chem. Soc. 123, 10095–10098 (2001).
Acknowledgements
We thank Caltech Beckman Institute Laser Resource Center (BILRC) and the Office of Laboratory Animal Resources (OLAR) for research resources. We thank M. Okumura and S. Cushing for kindly sharing research resources. We thank X. Tao and T. Begusic for calculating the Frank–Condon factors and A. Colazo and P. Kocheril for helpful discussions. This work is supported by an NIH Director’s New Innovator Award (DP2 GM140919-01 for L.W.) and an Alfred P. Sloan Research Fellowship (L.W.). L.W. is a Heritage Principal Investigator supported by the Heritage Medical Research Institute at Caltech.
Author information
Authors and Affiliations
Contributions
H.W., D.L. and L.W. conceived and designed the research. H.W. and D.L. built the BonFIRE set-up and performed solution measurements. H.W. wrote the LabVIEW scripts and conducted single-molecule measurements. D.L. conducted biological sample preparation and bioimaging experiments. H.W., D.L. and Y.C. collected and analysed the data. X.B. and K.M. helped provide cell, neuronal and brain tissue samples. J.D. synthesized BF dyes. L.W. supervised the experiments. The manuscript was written by H.W., D.L. and L.W., with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors have filed a provisional patent application (63/462,131) based on this work.
Peer review
Peer review information
Nature Photonics thanks Marcus Cicerone, Delong Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Power outputs of the Idler and the DFG IR lasers.
The power is measured at the laser outputs using a thermopile power meter (919P-003-10, Newport). All wavenumbers between 800 and 4800 cm−1 (2.1–12 μm) are covered. The pulse width is 2 ps with a bandwidth of 8–10 cm−1 according to the manufacturer (APE Angewandte Physik & Elektronik GmbH, Berlin, Germany).
Extended Data Fig. 2 Reduced photothermal background in acetonitrile-d3.
The fluorescence vs. IR-probe delay (tD) of ATTO680 in DMSO (blue) and acetonitrile-d3 (orange) obtained using 1598 cm−1 IR frequency and 765-nm probe wavelength. While the signal-to-background (S/B) ratio is about 8% in DMSO, it increases to 56% in acetonitrile-d3.
Extended Data Fig. 3 Correlation between BonFIRE and FTIR spectra.
a, FTIR (grey) and BonFIRE (red) spectra obtained for dyes in the fingerprint region. The dye and the experimental conditions (either in 10 mM DMSO solutions or in KBr pellets) for obtaining FTIR spectrum are indicated in each spectrum. The BonFIRE spectra (red curves) and single-point BonFIRE measurements (red circles, BonFIRE vs. IR-probe delay obtained for the single wavenumber) are scaled and overplotted on top of the FTIR reference for better comparison. b, FTIR and BonFIRE spectra of four dyes in the cell-silent region. c, FTIR spectra of other dyes expected to be measured in BonFIRE (for grey dots in Fig. 1b). The FTIR spectra in (a) and (c) measured in DMSO are often complicated by the IR absorption of DMSO and water (DMSO is hygroscopic), whose spectra are plotted in (d) for reference. The spectra of ATTO680 and Rh800 are not shown here as they are shown in Fig. 2f and Extended Data Fig. 9.
Extended Data Fig. 4 Characterization of cell-silent BonFIRE.
a, BonFIRE signal dependence on the probe wavelength (orange). The BonFIRE excitation profile (blue, by horizontally shifting the raw data by adding the IR frequency of 2224 cm−1) is overplotted with the absorption spectrum of BF1 (purple). Data are presented as peak values +/− s.d. from the background (n = 14). b,c, BonFIRE signal dependence on the IR power (b) and the probe power (c) on sample, measured from 10 μM BF1 in PBS. The probe power dependence shows good linearity (R2 = 1.00) at low power levels but quickly saturates. Data are presented as peak values +/− s.d. from the background (n = 17).
Extended Data Fig. 5 Spatial resolution of BonFIRE microscopy.
a, BonFIRE images of 100-nm fluorescent beads (Invitrogen™ FluoSpheres™ carboxylate-modified microspheres, 0.1 μm, 715/755 nm, Fisher Scientific) embedded in PVA film on a CaF2 substrate. The image was obtained with 1592 cm−1 IR (~24 mW on sample) and 815 nm probe (~180 μW on sample). Scale bar, 1 μm. b, A sectional profile obtained from XY plane from a location indicated by the white dashed line in (a), a lateral resolution (FWHM) of 0.6 μm is obtained from Gaussian fitting. The Z-resolution of 1.8 μm is determined by fitting the sectional profile from the ZY plane (shown as inset). a.u., arbitrary unit.
Extended Data Fig. 6 Observation of multiple single-molecule isotopologues within the same diffraction-limited spot.
a, Colour-coded fluorescence imaging for a mixture of single-molecule BF1 isotopologues as shown in Fig. 4i. Red: BF1-13C ≡ 14N (2170 cm−1); green: BF1-12C ≡ 15N (2195 cm−1); and blue: BF1-12C ≡ 14N (2220 cm−1). b, In-situ BonFIRE spectra obtained from three spots indicated by the corresponding numbered arrowheads in (a). Gaussian fittings (and a linear background) are shown as solid curves. Spectra of spots 1 (orange), 2 (grey), and 3 (blue) contain all three vibrational peaks. Slight variations in peak positions are possibly due to local interactions41. c.p.ms, counts per millisecond. These results demonstrate BonFIRE can resolve multiple vibrational colours within the same diffraction-limited spot at the single-molecule level.
Extended Data Fig. 7 Evidence for robust single-molecule sensitivity of BonFIRE.
a, Raw data of single-molecule BonFIRE imaging of ATTO680 C = C in Fig. 4a, confirming that BonFIRE data from subtraction are not from single-molecule bleaching or blinking. The S/Bs reach >1 because of the much-reduced photothermal background of the dilute single-molecule sample and the IR-transparent PVA matrix. c.p.ms, counts per millisecond. b, Raw data of single-molecule BonFIRE imaging of 12C ≡ 15N labelled BF1, the same sample is used in Fig. 4e–g. Repeatable BonFIRE contrast and no photobleaching are confirmed by the continuous scan. c, Signal vs. IR-probe delay (tD) of a small region containing multiple single BF1 molecules. The S/Bs reach ~3. The signal on/off as the function of tD is another evidence of the BonFIRE’s single-molecule sensitivity. Scale bars, 1 µm.
Extended Data Fig. 8 BonFIRE concentration curves with PMT AC detection.
a, The concentration curve of C = C in ATTO680 in DMSO. The lowest concentration measured is 1 nM. The probe wavelength is 765 nm, and the IR frequency is 1592 cm−1. Data are presented as peak values +/− SD from the background (n = 9). a.u., arbitrary unit. b, Concentration curve of C ≡ N in Rh800 in DMSO. The lowest concentration measured is 8 nM. The probe wavelength is 830 nm, and the IR frequency is 2228 cm−1. Both minimum concentrations are close to or below the calculated single-molecule threshold of 5 nM. Data are presented as peak values +/− s.d. from the background (n = 22).
Extended Data Fig. 9 Broad spectral coverage of BonFIRE microscopy.
a, BonFIRE images of Rh800-stained HeLa cells targeting six different vibrational modes. Vibrations of phenolic C-O (1100 cm−1), aromatic C-N (1300 cm−1), conjugated C = C (1504 cm−1 and 1598 cm−1), conjugated C ≡ N (2228 cm−1), and C-H stretch (2860 cm−1) can all serve for BonFIRE imaging. a.u., arbitrary unit. Scale bars, 10 µm. b, BonFIRE spectrum (red) and FTIR spectrum (grey) of Rh800. BonFIRE spectrum is normalized to the most intense peak at 1598 cm−1. The C ≡ N peak around 2228 cm−1 and a weak broad peak of the C-H region (2700 cm−1–3200 cm−1) are enlarged in insets for better comparison.
Supplementary information
Supplementary Information
Supplementary Figs. 1–15 and Tables 1–5.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, H., Lee, D., Cao, Y. et al. Bond-selective fluorescence imaging with single-molecule sensitivity. Nat. Photon. 17, 846–855 (2023). https://doi.org/10.1038/s41566-023-01243-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41566-023-01243-8