Abstract
Apicomplexa are unicellular eukaryotes and obligate intracellular parasites, including Plasmodium (the causative agent of malaria) and Toxoplasma (one of the most widespread zoonotic pathogens). Rhoptries, one of their specialized secretory organelles, undergo regulated exocytosis during invasion1. Rhoptry proteins are injected directly into the host cell to support invasion and subversion of host immune function2. The mechanism by which they are discharged is unclear and appears distinct from those in bacteria, yeast, animals and plants. Here, we show that rhoptry secretion in Apicomplexa shares structural and genetic elements with the exocytic machinery of ciliates, their free-living relatives. Rhoptry exocytosis depends on intramembranous particles in the shape of a rosette embedded into the plasma membrane of the parasite apex. Formation of this rosette requires multiple non-discharge (Nd) proteins conserved and restricted to Ciliata, Dinoflagellata and Apicomplexa that together constitute the superphylum Alveolata. We identified Nd6 at the site of exocytosis in association with an apical vesicle. Sandwiched between the rosette and the tip of the rhoptry, this vesicle appears as a central element of the rhoptry secretion machine. Our results describe a conserved secretion system that was adapted to provide defence for free-living unicellular eukaryotes and host cell injection in intracellular parasites.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The mass spectrometry proteomics data generated during this study are available from the ProteomeXchange Consortium (http://proteomecentral.proteomexchange.org) via the PRIDE partner repository62 with the dataset identifiers PXD022713 (Nd9 dataset) and PXD022725 (NdP1 dataset). All data generated or analysed during this study are included within the published article and its Supplementary Information files. Source data are provided with this paper.
References
Dubremetz, J. F. Rhoptries are major players in Toxoplasma gondii invasion and host cell interaction. Cell Microbiol. 9, 841–848 (2007).
Boothroyd, J. C. & Dubremetz, J. F. Kiss and spit: the dual roles of Toxoplasma rhoptries. Nat. Rev. Microbiol. 6, 79–88 (2008).
Carruthers, V. B. & Sibley, L. D. Sequential protein secretion from three distinct organelles of Toxoplasma gondii accompanies invasion of human fibroblasts. Eur. J. Cell Biol. 73, 114–123 (1997).
Frenal, K., Dubremetz, J. F., Lebrun, M. & Soldati-Favre, D. Gliding motility powers invasion and egress in Apicomplexa. Nat. Rev. Microbiol. 15, 645–660 (2017).
Besteiro, S., Dubremetz, J. F. & Lebrun, M. The moving junction of apicomplexan parasites: a key structure for invasion. Cell Microbiol. 13, 797–805 (2011).
Ito, D., Schureck, M. A. & Desai, S. A. An essential dual-function complex mediates erythrocyte invasion and channel-mediated nutrient uptake in malaria parasites. eLife 6, e23485 (2017).
Counihan, N. A. et al. Plasmodium falciparum parasites deploy RhopH2 into the host erythrocyte to obtain nutrients, grow and replicate. eLife 6, e23217 (2017).
Nguitragool, W. et al. Malaria parasite clag3 genes determine channel-mediated nutrient uptake by infected red blood cells. Cell 145, 665–677 (2011).
Hakimi, M.-A., Olias, P. & Sibley, L. D. Toxoplasma effectors targeting host signaling and transcription. Clin. Microbiol. Rev. 30, 615–645 (2017).
Singh, S., Alam, M. M., Pal-Bhowmick, I., Brzostowski, J. A. & Chitnis, C. E. Distinct external signals trigger sequential release of apical organelles during erythrocyte invasion by malaria parasites. PLoS Pathog. 6, e1000746 (2010).
Kessler, H. et al. Microneme protein 8—a new essential invasion factor in Toxoplasma gondii. J. Cell Sci. 121, 947–956 (2008).
Nichols, B. A., Chiappino, M. L. & O’Connor, G. R. Secretion from the rhoptries of Toxoplasma gondii during host-cell invasion. J. Ultrastruct. Res. 83, 85–98 (1983).
Coleman, B. I. et al. A member of the ferlin calcium sensor family is essential for Toxoplasma gondii rhoptry secretion. mBio 9, e01510-18 (2018).
Suarez, C. et al. A lipid-binding protein mediates rhoptry discharge and invasion in Plasmodium falciparum and Toxoplasma gondii parasites. Nat. Commun. 10, 4041 (2019).
Tseng, T. T., Tyler, B. M. & Setubal, J. C. Protein secretion systems in bacterial–host associations, and their description in the Gene Ontology. BMC Microbiol. 9, S2 (2009).
Gubbels, M.-J. & Duraisingh, M. T. Evolution of apicomplexan secretory organelles. Int. J. Parasitol. 42, 1071–1081 (2012).
Plattner, H. Trichocysts—Paramecium’s projectile-like secretory organelles: reappraisal of their biogenesis, composition, intracellular transport, and possible functions. J. Eukaryot. Microbiol. 64, 106–133 (2017).
Plattner, H., Miller, F. & Bachmann, L. Membrane specializations in the form of regular membrane-to-membrane attachment sites in Paramecium. A correlated freeze-etching and ultrathin-sectioning analysis. J. Cell Sci. 13, 687–719 (1973).
Knoll, G., Braun, C. & Plattner, H. Quenched flow analysis of exocytosis in Paramecium cells: time course, changes in membrane structure, and calcium requirements revealed after rapid mixing and rapid freezing of intact cells. J. Cell Biol. 113, 1295–1304 (1991).
Dubremetz, J. F. & Torpier, G. Freeze fracture study of the pellicle of an Eimerian sporozoite (Protozoa, Coccidia). J. Ultrastruct. Res. 62, 94–109 (1978).
Porchet, E. & Torpier, G. [Freeze fracture study of Toxoplasma and Sarcocystis infective stages (author’s transl)]. Z. Parasitenkd. 54, 101–124 (1977).
Porchet-Hennere, E. & Nicolas, G. Are rhoptries of Coccidia really extrusomes? J. Ultrastruct. Res. 84, 194–203 (1983).
Dubremetz, J. F. & Entzeroth, R. in Advances in Cell and Molecular Biology of Membranes, Vol. 2A (Membrane Traffic in Protozoa) (ed. Tartakoff, A. M. & Plattner, H.) 83–98 (Elsevier Science, 1993).
Lefort-Tran, M., Aufderheide, K., Pouphile, M., Rossignol, M. & Beisson, J. Control of exocytotic processes: cytological and physiological studies of trichocyst mutants in Paramecium tetraurelia. J. Cell Biol. 88, 301–311 (1981).
Beisson, J., Lefort-Tran, M., Pouphile, M., Rossignol, M. & Satir, B. Genetic analysis of membrane differentiation in Paramecium. Freeze-fracture study of the trichocyst cycle in wild-type and mutant strains. J. Cell Biol. 69, 126–143 (1976).
Gogendeau, D., Keller, A. M., Yanagi, A., Cohen, J. & Koll, F. Nd6p, a novel protein with RCC1-like domains involved in exocytosis in Paramecium tetraurelia. Eukaryot. Cell 4, 2129–2139 (2005).
Froissard, M., Keller, A. M. & Cohen, J. ND9P, a novel protein with Armadillo-like repeats involved in exocytosis: physiological studies using allelic mutants in Paramecium. Genetics 157, 611–620 (2001).
Skouri, F. & Cohen, J. Genetic approach to regulated exocytosis using functional complementation in Paramecium: identification of the ND7 gene required for membrane fusion. Mol. Biol. Cell 8, 1063–1071 (1997).
Froissard, M., Keller, A. M., Dedieu, J. C. & Cohen, J. Novel secretory vesicle proteins essential for membrane fusion display extracellular-matrix domains. Traffic 5, 493–502 (2004).
Paredes-Santos, T. C., de Souza, W. & Attias, M. Dynamics and 3D organization of secretory organelles of Toxoplasma gondii. J. Struct. Biol. 177, 420–430 (2012).
Sidik, S. M. et al. A genome-wide CRISPR screen in Toxoplasma identifies essential apicomplexan genes. Cell 166, 1423–1435 (2016).
Long, S. et al. Calmodulin-like proteins localized to the conoid regulate motility and cell invasion by Toxoplasma gondii. PLoS Pathog. 13, e1006379 (2017).
Meissner, M., Schluter, D. & Soldati, D. Role of Toxoplasma gondii myosin A in powering parasite gliding and host cell invasion. Science 298, 837–840 (2002).
Sheiner, L. et al. A systematic screen to discover and analyze apicoplast proteins identifies a conserved and essential protein import factor. PLoS Pathog. 7, e1002392 (2011).
Lamarque, M. H. et al. Plasticity and redundancy among AMA–RON pairs ensure host cell entry of Toxoplasma parasites. Nat. Commun. 5, 4098 (2014).
Koshy, A. A. et al. Toxoplasma secreting Cre recombinase for analysis of host–parasite interactions. Nat. Methods 7, 307–309 (2010).
Collins, C. R. et al. Robust inducible Cre recombinase activity in the human malaria parasite Plasmodium falciparum enables efficient gene deletion within a single asexual erythrocytic growth cycle. Mol. Microbiol. 88, 687–701 (2013).
Zhang, M. et al. Uncovering the essential genes of the human malaria parasite Plasmodium falciparum by saturation mutagenesis. Science 360, eaap7847 (2018).
Briguglio, J. S., Kumar, S. & Turkewitz, A. P. Lysosomal sorting receptors are essential for secretory granule biogenesis in Tetrahymena. J. Cell Biol. 203, 537–550 (2013).
Mueller, C. et al. The Toxoplasma protein ARO mediates the apical positioning of rhoptry organelles, a prerequisite for host cell invasion. Cell Host Microbe 13, 289–301 (2013).
Beisson, J., Cohen, J., Lefort-Tran, M., Pouphile, M. & Rossignol, M. Control of membrane fusion in exocytosis. Physiological studies on a Paramecium mutant blocked in the final step of the trichocyst extrusion process. J. Cell Biol. 85, 213–227 (1980).
Bonnemain, H., Gulik-Krzywicki, T., Grandchamp, C. & Cohen, J. Interactions between genes involved in exocytotic membrane fusion in Paramecium. Genetics 130, 461–470 (1992).
Varghese, T. Fine structure of the endogenous stages of Eimeria labbeana. I. The first generation merozoites. J. Protozool. 22, 66–71 (1975).
Sheffield, H. G. Electron microscope study of the proliferative form of Besnoitia jellisoni. J. Parasitol. 52, 583–594 (1966).
Perkins, F. O. Zoospores of the oyster pathogen, Dermocystidium marinum. I. Fine structure of the conoid and other sporozoan-like organelles. J. Parasitol. 62, 959–974 (1976).
El Hajj, H. et al. Molecular signals in the trafficking of Toxoplasma gondii protein MIC3 to the micronemes. Eukaryot. Cell 7, 1019–1028 (2008).
Collins, C. R., Withers-Martinez, C., Hackett, F. & Blackman, M. J. An inhibitory antibody blocks interactions between components of the malarial invasion machinery. PLoS Pathog. 5, e1000273 (2009).
Burghaus, P. A. & Holder, A. A. Expression of the 19-kilodalton carboxy-terminal fragment of the Plasmodium falciparum merozoite surface protein-1 in Escherichia coli as a correctly folded protein. Mol. Biochem. Parasitol. 64, 165–169 (1994).
Douki, J. B. et al. Adhesion of normal and Plasmodium falciparum ring-infected erythrocytes to endothelial cells and the placenta involves the rhoptry-derived ring surface protein-2. Blood 101, 5025–5032 (2003).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Cerede, O. et al. Synergistic role of micronemal proteins in Toxoplasma gondii virulence. J. Exp. Med. 201, 453–463 (2005).
Branton, D. et al. Freeze-etching nomenclature. Science 190, 54–56 (1975).
Knuepfer, E., Napiorkowska, M., van Ooij, C. & Holder, A. A. Generating conditional gene knockouts in Plasmodium—a toolkit to produce stable DiCre recombinase-expressing parasite lines using CRISPR/Cas9. Sci. Rep. 7, 3881 (2017).
Iancu, C. V. et al. Electron cryotomography sample preparation using the Vitrobot. Nat. Protoc. 1, 2813–2819 (2006).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Lebrun, M. et al. The rhoptry neck protein RON4 relocalizes at the moving junction during Toxoplasma gondii invasion. Cell Microbiol. 7, 1823–1833 (2005).
Skorupa, A. et al. Angiogenin induces modifications in the astrocyte secretome: relevance to amyotrophic lateral sclerosis. J. Proteom. 91, 274–285 (2013).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Perez-Riverol, Y. et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res. 47, D442–D450 (2019).
Wilson, D., Madera, M., Vogel, C., Chothia, C. & Gough, J. The SUPERFAMILY database in 2007: families and functions. Nucleic Acids Res. 35, D308–D313 (2007).
Hadjebi, O., Casas-Terradellas, E., Garcia-Gonzalo, F. R. & Rosa, J. L. The RCC1 superfamily: from genes, to function, to disease. Biochim. Biophys. Acta 1783, 1467–1479 (2008).
Acknowledgements
We thank S. Lourido for the pU6-Universal plasmid; A. Scherf for the anti-PfRAP2 antibodies; D. Soldati-Favre for providing the anti-ARO antibodies and Tgaro iKD strain; M. Blackman for the anti-PfAMA1 and anti-PfMSP1 antibodies, pDC2-Cas9-hDHFRyFCU plasmid and p230p DiCre strain; J. Boothroyd for the monoclonal antibody CCL2 anti-TgAMA1; E. Peterson for anti-TgSAG1; R. Waller for the ring-1–GFP plasmid; A. Koshy for the toxofilin-Cre plasmid; and H. Blau’s laboratory for the Cre reporter DsRed cell line. We thank O. Billker for providing us with compound 2. We thank V. Richard and F. Godiard for technical assistance with the electron microscopy service of the University of Montpellier. We are also grateful to E. Jublanc and the imaging facility MRI at the University of Montpellier (part of the national infrastructure France-BioImaging supported by the French National Research Agency (ANR-10-INBS-04; Investments for the future)) and to C. Duperray of the MRI-Cytometry facility at the Institute for Regenerative Medicine and Biotherapy for assistance and technical support. We thank J. Marcetteau for the graphical design of Figs. 1b and 4. We acknowledge the use of instruments at the Electron Microscopy Resource Lab and Beckman Center for Cryo-Electron Microscopy at the University of Pennsylvania. M.L. is an INSERM researcher. This work was supported by the Laboratoire d’Excellence ParaFrap (ANR-11-LABX-0024), ANR (ANR-16-CE15-0010-01), Fondation pour la Recherche Medicale (Equipe FRM DEQ20170336725), European Research Council (ERC advanced grant number 833309 KissAndSpitRhoptry to M.L.), France and Chicago Collaborating in the Sciences (to A.P.T. and M.L.) and the National Institutes of Health (GM105783 to A.P.T.), as well as by a David and Lucile Packard Fellowship for Science and Engineering (2019-69645 to Y.-W.C.) and an EMBO fellowship (number ALTF 58-2018 to A.N.G.).
Author information
Authors and Affiliations
Contributions
E.A. performed most of the work on Toxoplasma. M.M.C. generated the Plasmodium data and completed the work on Toxoplasma during revision. A.G. performed the phylogenetic analyses. E.A., D.S. and R.N. generated the T. thermophila data. C.S. generated the donor plasmid for Pfnd9 iKO. A.N.G., N.D.S.P., R.N. and M.M. contributed to the Toxoplasma phenotypic analyses. D.M.P.-V. performed the immunoprecipitation analyses. S.U. performed the mass spectrometry analyses. E.A., L.B.-S. and J.-F.D. performed the electron microscopy. E.A., P.R.F. and J.-F.D. performed the freeze-fracture analyses. S.K.M. and Y.-W.C. performed the cryo-ET. A.N.G. and S.K.M. prepared the samples for cryo-ET. M.L. designed the study with support from E.A. M.L. supervised the research. E.A. and M.L. wrote the paper with editorial support from M.M.C., A.P.T., D.S., A.N.G., S.K.M., Y.-W.C. and B.S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Microbiology thanks Ke Hu, Geoffrey Ian G. McFadden and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Amino acid sequence alignments and phylogenetic analyses.
a–d, Maximum likelihood phylogenetic trees based on Alveolata non discharge (Nd) Nd6 (a) Nd9 (b) NdP1 (c) and NdP2 (d) protein sequences. Asterisks indicate two homologous genes in the same species. The scale bar represents the branch length values. Amino acids sequences, accession number and alignments used to generate phylogenetic tree of Nd6, Nd9, NdP1 and NdP2 are presented in Source Data Extended Data Fig. 1.
Extended Data Fig. 2 Generation of Tgnd6 iKD and Tgnd9 iKD.
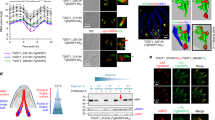
a, Strategy for conditional depletion of TgNd6 (TGGT1_248640) using the Auxin-inducible degron system. b, Integration PCRs showing double homologous recombination at the 5’ (top) and at the 3’ (bottom) of the Tgnd6 locus. Representative of 3 experiments. c, Strategy for conditional depletion of TgNd9 (TGGT1_249730) using the Tet-OFF system. d, Integration PCRs showing double homologous recombination at the 3’ (top) and at the 5’ (bottom) of the Tgnd9 locus. Representative of 3 experiments. e, Immunoblot of Tgnd6 iKD ± IAA (top) and Tgnd9 iKD ± ATc (bottom) showing the timing for protein depletion. ROP5, loading control. Ctrl: ΔKu80-Tir1 (upper panel) and ΔKu80-TaTi (lower panel). Representative of one experiment for Tgnd9 iKD and 2 experiments for Tgnd6 iKD. f, IFA of Tgnd6 iKD shows depletion of TgNd6 after 24 hours IAA treatment (top right), IFA of Tgnd9 iKD shows depletion of TgNd9 after 72 hours ATc treatment (bottom right). Red, anti-ARO. Green, anti-HA. Representative of 3 experiments. g, Detergent extraction profile of TgND6 and TgND9 after solubilization of HA‐tagged TgNds parasites with Tris 1 mM, 1% TritonX‐100 or 1% SDS. Representative of 2 experiments.
Extended Data Fig. 3 Depletion of Nd6 or Nd9 in T. gondii parasites does not affect replication, egress, gliding motility nor attachment.
a, Plaque assay. Left: ΔKu80-Tir1 (Ctrl) ± IAA vs Tgnd6 iKD ± IAA. Right: ΔKu80-TaTi (Ctrl) ± ATc vs Tgnd9 iKD ± ATc. 7 dpi lysis plaque areas were measured. No plaques were observed for Tgnd9 iKD +ATc. Means of plaque area ± SD of 3 biological samples. Representative of 3 independent assays. b, Intracellular replication. Left: ΔKu80-Tir1 (Ctrl) and Tgnd6 iKD ± IAA. Right: ΔKu80-TaTi (Ctrl) and Tgnd9 iKD ± ATc. The percentage of vacuoles containing 2, 4, 8, 16 or 32 parasites was determined for 200 vacuoles. Means ± SD from 3 biological samples. Representative of 3 independent experiments. c, Percentage of egressed vacuoles. Left: ΔKu80-Tir1 (Ctrl) and Tgnd6 iKD ± IAA. Right: ΔKu80-TaTi (Ctrl) and Tgnd9 iKD ± ATc. Egress events were quantified for 200 vacuoles. Means ± SD from 3 biological samples. Representative of 2 independent experiments. d, Conoid protrusion assay. The graph represents the percentage of protruded conoids induced by 5 μM A23187, or DMSO as control. Means ±SD from 3 biological samples. Representative of 3 independent experiments. e, Gliding assays were performed with ΔKu80-Tir1 (Ctrl) and Tgnd6 iKD parasite lines ± IAA 24 h (left) and ΔKu80-TaTi (Ctrl) and Tgnd9 iKD parasite lines ± ATc 72 h (right). Trails were revealed by IFA using anti-SAG1 antibodies. Scale bar=10 µm. Representative of 3 experiments. f, Quantification of parasite motility by time−lapse video microscopy. Data are plotted as a percentage of circular, helical and twirling movements. Values represent mean of 2–4 movies. g, Quantification of attachment on glutaraldehyde fixed host cell. As control of microneme secretion-dependent attachment, parasites were treated with BAPTA-AM. h, Quantification of rhoptry secretion by e-vacuoles assay. Tgnd6 iKD was compared to the background line Δku80-Tir1 ± IAA 24 h and Tgnd9 iKD was compared to the background line Δku80-TATi ± Atc 72 h. (g,h) Means ± SD from 3 biological samples. Representative of one experiment. The significance of the results was assessed using a parametric paired t-test (Student’s two-tailed t-test). ns = non-significant.
Extended Data Fig. 4 Secretory organelles biogenesis and positioning is not affected in Tgnd6-iKD and Tgnd9-iKD.
a, b, IFA using anti-ARO antibodies (rhoptry marker) and anti-MIC2 (microneme marker) on Tgnd6 iKD parasites ± IAA 24 h (a) and on Tgnd9 iKD parasites ± ATc 72 h (b). Scale bar=1 µm. Representative of 3 experiments. c, d, Electron microscopy images showing the apical complex of extracellular tachyzoites of Tgnd6 iKD +IAA 24 h (c) and of Tgnd9-iKD +ATc 72 h. Representative of one experiment. (d). Rh B = rhoptry bulb, Rh N = rhoptry neck, Co = conoid, m = microneme. Bar = 500 nm.
Extended Data Fig. 5 Generation of Pfnd9 iKO.
a, Strategy for conditional depletion of PfNd9 (PF3D7_1232700) using the DiCre system. b, Integration PCRs showing double homologous recombination at the 5’ end (top panel) and at the 3’ end (middle panel) of the Pfnd9 locus. In the bottom panel, PCR demonstrating the absence of the wt locus in the floxed clonal population. Representative of 2 experiments. c, PCRs showing efficient excision of the Pfnd9 locus upon addition of rapamycin. Representative of 3 experiments. d, Immunoblot on parasite lysates from Pfnd9 iKO mutant parasites. We could not detect the protein by IFA and only observed a faint band at the expected size in late schizonts by Western blot, both consistent with the very low transcript level of PfNd9 (Plasmodb.org). Representative of 2 experiments. e, Representative images of 2 experiments from the time lapse video microscopy of merozoites egressing from schizonts in the DMSO control and in the rapamycin-treated Pfnd9 iKO parasites. Relative time shown in minutes.
Extended Data Fig. 6 Generation of Tgnp1 iKD and Tgndp2 iKD.
a, Strategy for conditional depletion of TgNdP1 (TGGT1_222660) using the Tet-OFF system. b, PCRs confirming integration events (double homologous recombination) at the 5’ end (top panel) and at the 3’ end (bottom panel) at the Tgnp1 locus. Representative of 3 experiments. c, Strategy for conditional depletion of TgNdP2 (TGGT1_ 316730) using the Auxin-inducible degron system. d, PCRs confirming integration events (double homologous recombination) at the 5’ end (top panel) and at the 3’ end (bottom panel) at the Tgnp2 locus. Representative of 2 experiments. e, Immunoblot of Tgndp1 iKD +ATc and Tgndp2 iKD +IAA showing proteins depletion. ROP5, loading control. Ctrl: ΔKu80-TATi (left) and ΔKu80-Tir1 (right). Representative of one experiment for Tgndp1 iKD and 2 experiments for Tgndp2 iKD. f, IFA of Tgndp1 iKD shows depletion of TgNdP1 after 72 hours ATc treatment (left), IFA of Tgndp2 iKD shows depletion of TgNdP2 after 24 hours IAA treatment (right). Representative of 3 experiments.
Extended Data Fig. 7 Depletion of NdP1 or NdP2 in T. gondii parasites does not affect replication, egress, gliding motility, nor attachment.
a, Plaque assay. Top: ΔKu80-TaTi (Ctrl) ± ATc vs Tgndp1-iKD ± ATc. Bottom: ΔKu80-Tir1 (Ctrl) ± IAA vs Tgndp2 iKD ± IAA. Means of plaque area ± SD from 3 biological samples. Representative of 3 independent experiments. b, IFA using anti-ROP5 and anti-ARO antibodies (rhoptry markers), and anti-MIC2 (microneme marker) on Tgndp1 iKD +ATc 72 h and Tgndp2 iKD +IAA 24 h. Scale bar=1 µm. Representative of 3 experiments. c, Intracellular replication. ΔKu80-TaTi (Ctrl) vs Tgndp1 iKD ±ATc and ΔKu80-Tir1 (Ctrl) vs Tgndp2 iKD ±IAA. The percentage of vacuoles containing 2, 4, 8, 16 or 32 parasites was determined for 200 vacuoles. Means ± SD from 3 biological samples. Representative of 3 independent experiments. d, Percentage of egressed vacuoles. ΔKu80-TaTi (Ctrl) vs Tgndp1 iKD ± ATc and ΔKu80-Tir1 (Ctrl) vs Tgndp2 iKD ± IAA. Egress events were quantified for 200 vacuoles. Means ± SD from 3 biological samples. Representative of 2 independent experiments. e, Conoid protrusion assay. The graph represents the percentage of protruded conoids induced by 5 μM A23187, and DMSO as control. Means ± SD from 3 biological samples. Representative of 3 independent experiments. f, Gliding assays comparing ΔKu80-TaTi (Ctrl) vs Tgndp1 iKD parasite lines ± ATc 72 h or comparing ΔKu80-Tir1 (Ctrl) vs Tgndp2 iKD ± IAA 24 h. Trails were revealed by IFA using anti-SAG1 antibodies. Scale bar=10 µm. Representative of 3 experiments. g, Quantification of parasite motility by time-lapse video microscopy. Data are plotted as a percentage of circular, helical and twirling movements. Values represent mean of 2–4 movies. h, Quantification of attachment on glutaraldehyde fixed host cell. As control of microneme secretion-dependent attachment, parasites were treated with BAPTA-AM. i, Immunoblot showing microneme secretion assessed by the release of the processed form of AMA1 (arrow=processed/secreted TgAMA1) in Tgndp1 iKD ± ATc 72 h and Tgndp2 iKD ± IAA 24 h. Ctrl = ΔKu80-Tir1 + IAA 24. P = pellet, Sup = supernatant, Sup ind = propanolol-induced supernatant. GRA3, loading control. j, Quantification of rhoptry secretion by e-vacuoles assay in Tgndp1 iKD ±ATc 72 h. (h, j) Means ± SD from 3 biological samples. Representative of one experiment. The significance of the results was assessed using a parametric paired t-test (Student’s two-tailed t-test). ns = non-significant.
Extended Data Fig. 8 Mucocyst biogenesis and positioning is not affected in TtΔndp1 and TtΔndp2.
a, Generation of Ttndp1 and Ttndp2 knockout strains in T. thermophila Cu428. b, cDNA from TtΔndp1 (clones 4 and 9) or TtΔndp2 (Clones 1 and 5) were PCR amplified with primers P13/P14 and P21/P22 respectively, to assay the presence of the corresponding transcripts in the knockout strain. To confirm that equal amounts of cDNA were being amplified, reactions with primers specific for β-tubulin 1 were run in parallel (loading control). Representative of 2 experiments. c, Immunoblot showing proGrl1p processing in wild-type and mutant strains (clones 4 and 9 for TtΔndp1; clones 1 and 5 for TtΔndp2). Representative of one experiment. d, Confocal cross (top) and transverse (bottom) sections, with paired differential interference contrast (DIC) images, of TtΔndp1 (clone 9) and TtΔndp2 (clone 5). Cells were immunostained with mAbs 5E9, which recognize the mucocyst protein Grl3p. The wild-type pattern of the docked mucocysts is maintained in TtΔndp1 and TtΔndp2 clones. Scale bar = 10μm. Representative of 3 experiments.
Extended Data Fig. 9 Cryo-ET of rhoptry secretion system.
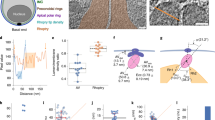
Four examples (a-d) from cryo-ET imaging of apical ends of the rhoptry secretion system. (i) vertical tomogram slices show the arrangement of the rhoptry (orange), tip density (cyan), apical vesicle (AV; magenta), rosette (dark blue) and the plasma membrane (PM; light blue). Original images (right) are annotated with color overlays (left). (ii) Magnified image of the boxed region in (i) showing the side view of the rosette. (iii) Top view of the rosette from a horizontal tomogram section through it and perpendicular to the plane in (ii) showing an 8-fold rotational symmetry. All measurements are made in 3D. Panel (a) shows images from the unfiltered version of the tomogram that is shown in Fig. 4a-c. In each of the four examples, images in (ii) and (iii) are from the same cell. Scale bars: 50 nm. Representative of 20 tomograms produced from 2 independent samples.
Extended Data Fig. 10 Apical area of a Sarcocystis tenella cystozoite.
Similarly to what was observed in literature and experimentally in T. gondii, cystozoites of S. tenella display an apical vesicle in between the apical tip of the rhoptry neck and the plasma membrane, at the site of exocytosis. PCR=posterior preconoidal ring; co = conoid; Rh = rhoptry neck; m = micronemes; AV=apical vesicle. Bar is 0.2 µm. Representative of 2 experiments.
Supplementary information
Supplementary Information
Supplementary Fig. 1, Table 1, Text, Methods and references.
Supplementary Table 2
Mass spectrometry data of Nd9 immunoprecipitation.
Supplementary Table 3
Mass spectrometry data of NdP1 immunoprecipitation.
Supplementary Table 4
List of primers used.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 1
Amino acid sequence alignments.
Source Data Extended Data Fig. 2
Unprocessed gels and western blots.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed gels and western blot.
Source Data Extended Data Fig. 6
Unprocessed gels and western blots.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 7
Unprocessed western blots.
Source Data Extended Data Fig. 8
Unprocessed gels and western blots.
Rights and permissions
About this article
Cite this article
Aquilini, E., Cova, M.M., Mageswaran, S.K. et al. An Alveolata secretory machinery adapted to parasite host cell invasion. Nat Microbiol 6, 425–434 (2021). https://doi.org/10.1038/s41564-020-00854-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-020-00854-z
This article is cited by
-
Genomic insights into the cellular specialization of predation in raptorial protists
BMC Biology (2024)
-
Toxoplasma protein export and effector function
Nature Microbiology (2024)
-
Sustained rhoptry docking and discharge requires Toxoplasma gondii intraconoidal microtubule-associated proteins
Nature Communications (2024)
-
Cryo-tomography reveals rigid-body motion and organization of apicomplexan invasion machinery
Nature Communications (2023)
-
A splitCas9 phenotypic screen in Toxoplasma gondii identifies proteins involved in host cell egress and invasion
Nature Microbiology (2022)