Abstract
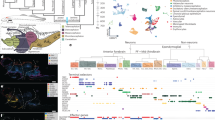
The origin and evolution of cell types has emerged as a key topic in evolutionary biology. Driven by rapidly accumulating single-cell datasets, recent attempts to infer cell type evolution have largely been limited to pairwise comparisons because we lack approaches to build cell phylogenies using model-based approaches. Here we approach the challenges of applying explicit phylogenetic methods to single-cell data by using principal components as phylogenetic characters. We infer a cell phylogeny from a large, comparative single-cell dataset of eye cells from five distantly related mammals. Robust cell type clades enable us to provide a phylogenetic, rather than phenetic, definition of cell type, allowing us to forgo marker genes and phylogenetically classify cells by topology. We further observe evolutionary relationships between diverse vessel endothelia and identify the myelinating and non-myelinating Schwann cells as sister cell types. Finally, we examine principal component loadings and describe the gene expression dynamics underlying the function and identity of cell type clades that have been conserved across the five species. A cell phylogeny provides a rigorous framework towards investigating the evolutionary history of cells and will be critical to interpret comparative single-cell datasets that aim to ask fundamental evolutionary questions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
A GitHub repository (https://github.com/dunnlab/cellphylo) is provided containing select input, intermediate and output files (‘cellphylo/analysis/’) sufficient to reproduce the analyses.
Code availability
All custom code is available in our GitHub repository: https://github.com/dunnlab/cellphylo.
References
Sulston, J. E., Schierenberg, E., White, J. G. & Thomson, J. N. The embryonic cell lineage of the nematode Caenorhabditis elegans. Dev. Biol. 100, 64–119 (1983).
Martindale, M. Q. & Henry, J. Q. Intracellular fate mapping in a basal metazoan, the ctenophore Mnemiopsis leidyi, reveals the origins of mesoderm and the existence of indeterminate cell lineages. Dev. Biol. 214, 243–257 (1999).
Spencer Chapman, M. et al. Lineage tracing of human development through somatic mutations. Nature 595, 85–90 (2021).
Tanay, A. & Sebé-Pedrós, A. Evolutionary cell type mapping with single-cell genomics. Trends Genet. 37, 919–932 (2021).
Shekhar, K. et al. Comprehensive classification of retinal bipolar neurons by single-cell transcriptomics. Cell 166, 1308–1323 (2016).
Gilbert, E. et al. Molecular and cellular architecture of the larval sensory organ in the cnidarian Nematostella vectensis. Development 149, dev200833 (2022).
Kin, K., Nnamani, M. C., Lynch, V. J., Michaelides, E. & Wagner, G. P. Cell-type phylogenetics and the origin of endometrial stromal cells. Cell Rep. 10, 1398–1409 (2015).
Liang, C., Forrest, A. R. R. & Wagner, G. P. The statistical geometry of transcriptome divergence in cell-type evolution and cancer. Nat. Commun. 6, 6066 (2015).
Musser, J. M. et al. Profiling cellular diversity in sponges informs animal cell type and nervous system evolution. Science 374, 717–723 (2021).
Hughes, A. L. & Friedman, R. A phylogenetic approach to gene expression data: evidence for the evolutionary origin of mammalian leukocyte phenotypes. Evol. Dev. 11, 382–390 (2009).
Wagner, G. P. Homology, Genes, and Evolutionary Innovation (Princeton Univ. Press, 2014).
Arendt, D. et al. The origin and evolution of cell types. Nat. Rev. Genet. 17, 744–757 (2016).
Arendt, D. The evolution of cell types in animals: emerging principles from molecular studies. Nat. Rev. Genet. 9, 868–882 (2008).
Arendt, D. Evolution of eyes and photoreceptor cell types. Int. J. Dev. Biol. 47, 563–571 (2003).
Arendt, D., Bertucci, P. Y., Achim, K. & Musser, J. M. Evolution of neuronal types and families. Curr. Opin. Neurobiol. 56, 144–152 (2019).
Serb, J. M. & Oakley, T. H. Hierarchical phylogenetics as a quantitative analytical framework for evolutionary developmental biology. Bioessays 27, 1158–1166 (2005).
Packer, J. S. et al. A lineage-resolved molecular atlas of C. elegans embryogenesis at single-cell resolution. Science 365, eaax1971 (2019).
Wagner, D. E. & Klein, A. M. Lineage tracing meets single-cell omics: opportunities and challenges. Nat. Rev. Genet. 21, 410–427 (2020).
Whitman, C. O. The embryology of Clepsine. J. Cell Sci. s2-18, 215–315 (1878).
Raj, B. et al. Simultaneous single-cell profiling of lineages and cell types in the vertebrate brain. Nat. Biotechnol. 36, 442–450 (2018).
Nowell, P. C. The clonal evolution of tumor cell populations. Science 194, 23–28 (1976).
Navin, N. et al. Tumour evolution inferred by single-cell sequencing. Nature 472, 90–94 (2011).
Yang, D. et al. Lineage tracing reveals the phylodynamics, plasticity, and paths of tumor evolution. Cell 185, 1905–1923 (2022).
Levy, S. et al. A stony coral cell atlas illuminates the molecular and cellular basis of coral symbiosis, calcification, and immunity. Cell 184, 2973–2987 (2021).
Seidel, S. & Stadler, T. TiDeTree: a Bayesian phylogenetic framework to estimate single-cell trees and population dynamic parameters from genetic lineage tracing data. Proc. R. Soc. B 289, 20221844 (2022).
Zhao, Z.-M. et al. Early and multiple origins of metastatic lineages within primary tumors. Proc. Natl Acad. Sci. USA 113, 2140–2145 (2016).
Moravec, J. C., Lanfear, R., Spector, D. L., Diermeier, S. D. & Gavryushkin, A. Testing for phylogenetic signal in single-cell RNA-seq data. J. Comput. Biol. 30, 518–537 (2023).
Saitou, N. & Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425 (1987).
Paganos, P., Voronov, D., Musser, J. M., Arendt, D. & Arnone, M. I. Single-cell RNA sequencing of the Strongylocentrotus purpuratus larva reveals the blueprint of major cell types and nervous system of a non-chordate deuterostome. Elife 10, e70416 (2021).
Wang, Y. et al. Clonal evolution in breast cancer revealed by single nucleus genome sequencing. Nature 512, 155–160 (2014).
Frumkin, D., Wasserstrom, A., Kaplan, S., Feige, U. & Shapiro, E. Genomic variability within an organism exposes its cell lineage tree. PLoS Comput. Biol. 1, e50 (2005).
Tarashansky, A. J. et al. Mapping single-cell atlases throughout Metazoa unravels cell type evolution. Elife 10, e66747 (2021).
van Zyl, T. et al. Cell atlas of aqueous humor outflow pathways in eyes of humans and four model species provides insight into glaucoma pathogenesis. Proc. Natl Acad. Sci. USA 117, 10339–10349 (2020).
Sebé-Pedrós, A. et al. Early metazoan cell type diversity and the evolution of multicellular gene regulation. Nat. Ecol. Evol. 2, 1176–1188 (2018).
Wang, R. et al. Construction of a cross-species cell landscape at single-cell level. Nucleic Acids Res. 51, 501–516 (2023).
Chen, D. et al. Single cell atlas for 11 non-model mammals, reptiles and birds. Nat. Commun. 12, 7083 (2021).
Dunn, C. W., Zapata, F., Munro, C., Siebert, S. & Hejnol, A. Pairwise comparisons across species are problematic when analyzing functional genomic data. Proc. Natl Acad. Sci. USA 115, E409–E417 (2018).
Felsenstein, J. Phylogenies and the comparative method. Am. Nat. 125, 1–15 (1985).
Grafen, A. The phylogenetic regression. Phil. Trans. R. Soc. Lond. B 326, 119–157 (1997).
Sebé-Pedrós, A., Degnan, B. M. & Ruiz-Trillo, I. The origin of Metazoa: a unicellular perspective. Nat. Rev. Genet. 18, 498–512 (2017).
Mah, J. L., Christensen-Dalsgaard, K. K. & Leys, S. P. Choanoflagellate and choanocyte collar-flagellar systems and the assumption of homology. Evol. Dev. 16, 25–37 (2014).
Laundon, D., Larson, B. T., McDonald, K., King, N. & Burkhardt, P. The architecture of cell differentiation in choanoflagellates and sponge choanocytes. PLoS Biol. 17, e3000226 (2019).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
McInnes, L., Healy, J. & Melville, J. UMAP: Uniform Manifold Approximation and Projection for dimension reduction. Preprint at http://arxiv.org/abs/1802.03426 (2020).
Felsenstein, J. Maximum-likelihood estimation of evolutionary trees from continuous characters. Am. J. Hum. Genet. 25, 471–492 (1973).
Felsenstein, J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39, 783–791 (1985).
Lemoine, F. et al. Renewing Felsenstein’s phylogenetic bootstrap in the era of big data. Nature 556, 452–456 (2018).
Thorley, J. L. & Wilkinson, M. Testing the phylogenetic stability of early tetrapods. J. Theor. Biol. 200, 343–344 (1999).
Levy, C., Khaled, M. & Fisher, D. E. MITF: master regulator of melanocyte development and melanoma oncogene. Trends Mol. Med. 12, 406–414 (2006).
van Zyl, T. et al. Cell atlas of the human ocular anterior segment: tissue-specific and shared cell types. Proc. Natl Acad. Sci. USA 119, e2200914119 (2022).
Sokal, R. R. et al. Principles of Numerical Taxonomy (WH Freeman & Co, 1963).
Sokal, R. R. & Michener, C. D. A statistical method for evaluating systematic relationships. Univ. Kans. Sci. Bull. 38, 1409–1438 (1958).
Iwasa, M. A. & Suzuki, H. Evolutionary significance of chromosome changes in northeastern Asiatic red-backed voles inferred with the aid of intron 1 sequences of the G6pd gene. Chromosome Res. 10, 419–428 (2002).
Leclaire, S., Menard, S. & Berry, A. Molecular characterization of Babesia and Cytauxzoon species in wild South-African meerkats. Parasitology 142, 543–548 (2015).
Dimayacyac, J. R., Wu, S. & Pennell, M. Evaluating the performance of widely used phylogenetic models for gene expression evolution. Prepint at bioRxiv https://doi.org/10.1101/2023.02.09.527893 (2023).
Rohlfs, R. V., Harrigan, P. & Nielsen, R. Modeling gene expression evolution with an extended Ornstein–Uhlenbeck process accounting for within-species variation. Mol. Biol. Evol. 31, 201–211 (2014).
Bertram, J. et al. CAGEE: computational analysis of gene expression evolution. Mol. Biol. Evol. 40, msad106 (2023).
Harmon, L. J. et al. Early bursts of body size and shape evolution are rare in comparative data. Evolution 64, 2385–2396 (2010).
Wagner, G. P. Homologues, natural kinds and the evolution of modularity. Integr. Comp. Biol. 36, 36–43 (1996).
Liang, C., Musser, J. M., Cloutier, A., Prum, R. O. & Wagner, G. P. Pervasive correlated evolution in gene expression shapes cell and tissue type transcriptomes. Genome Biol. Evol. 10, 538–552 (2018).
Graf, T. & Enver, T. Forcing cells to change lineages. Nature 462, 587–594 (2009).
Hobert, O. Regulatory logic of neuronal diversity: terminal selector genes and selector motifs. Proc. Natl Acad. Sci. USA 105, 20067–20071 (2008).
Yin, W., Mendoza, L., Monzon-Sandoval, J., Urrutia, A. O. & Gutierrez, H. Emergence of co-expression in gene regulatory networks. PLoS ONE 16, e0247671 (2021).
Spellman, P. T. et al. Comprehensive identification of cell cycle-regulated genes of the yeast Saccharomyces cerevisiae by microarray hybridization. Mol. Biol. Cell 9, 3273–3297 (1998).
Wang, J. et al. Single-cell co-expression analysis reveals distinct functional modules, co-regulation mechanisms and clinical outcomes. PLoS Comput. Biol. 12, e1004892 (2016).
Hall, B. K. Germ layers, the neural crest and emergent organization in development and evolution. Genesis 56, e23103 (2018).
Hashimshony, T., Feder, M., Levin, M., Hall, B. K. & Yanai, I. Spatiotemporal transcriptomics reveals the evolutionary history of the endoderm germ layer. Nature 519, 219–222 (2015).
Steinmetz, P. R. H., Aman, A., Kraus, J. E. M. & Technau, U. Gut-like ectodermal tissue in a sea anemone challenges germ layer homology. Nat. Ecol. Evol. 1, 1535–1542 (2017).
Gage, P. J., Rhoades, W., Prucka, S. K. & Hjalt, T. Fate maps of neural crest and mesoderm in the mammalian eye. Invest. Ophthalmol. Vis. Sci. 46, 4200–4208 (2005).
Williams, A. L. & Bohnsack, B. L. Neural crest derivatives in ocular development: discerning the eye of the storm. Birth Defects Res. C Embryo Today 105, 87–95 (2015).
Rodrigues, M. M., Katz, S. I., Foidart, J. M. & Spaeth, G. L. Collagen, factor VIII antigen, and immunoglobulins in the human aqueous drainage channels. Ophthalmology 87, 337–345 (1980).
Pandolfi, M. Coagulation factor VIII: localization in the aqueous outflow pathways. Arch. Ophthalmol. 94, 656–658 (1976).
Kizhatil, K., Ryan, M., Marchant, J. K., Henrich, S. & John, S. W. M. Schlemm’s canal is a unique vessel with a combination of blood vascular and lymphatic phenotypes that forms by a novel developmental process. PLoS Biol. 12, e1001912 (2014).
Ramírez, J. M. et al. Schlemm’s canal and the collector channels at different developmental stages in the human eye. Cells Tissues Organs 178, 180–185 (2004).
Ashton, N. Anatomical study of Schlemm’s canal and aqueous veins by means of neoprene casts. Part I. Aqueous veins. Br. J. Ophthalmol. 35, 291–303 (1951).
Smelser, G. K. & Ozanics, V. The development of the trabecular meshwork in primate eyes. Am. J. Ophthalmol. 71, 366–385 (1971).
Krohn, J. Expression of factor VIII-related antigen in human aqueous drainage channels. Acta Ophthalmol. Scand. 77, 9–12 (1999).
Francois, M., Harvey, N. L. & Hogan, B. M. The transcriptional control of lymphatic vascular development. Physiology 26, 146–155 (2011).
Aspelund, A. et al. The Schlemm’s canal is a VEGF-C/VEGFR-3-responsive lymphatic-like vessel. J. Clin. Invest. 124, 3975–3986 (2014).
Trost, A. et al. Brain and retinal pericytes: origin, function and role. Front. Cell. Neurosci. 10, 20 (2016).
Alarcon-Martinez, L. et al. Capillary pericytes express α-smooth muscle actin, which requires prevention of filamentous-actin depolymerization for detection. Elife 7, e34861 (2018).
Etchevers, H. C., Vincent, C., Le Douarin, N. M. & Couly, G. F. The cephalic neural crest provides pericytes and smooth muscle cells to all blood vessels of the face and forebrain. Development 128, 1059–1068 (2001).
Ignarro, L. J. et al. Mechanism of vascular smooth muscle relaxation by organic nitrates, nitrites, nitroprusside and nitric oxide: evidence for the involvement of S-nitrosothiols as active intermediates. J. Pharmacol. Exp. Ther. 218, 739–749 (1981).
Bouallegue, A., Daou, G. B. & Srivastava, A. K. Endothelin-1-induced signaling pathways in vascular smooth muscle cells. Curr. Vasc. Pharmacol. 5, 45–52 (2007).
Kamikawatoko, S. et al. Nitric oxide relaxes bovine ciliary muscle contracted by carbachol through elevation of cyclic GMP. Exp. Eye Res. 66, 1–7 (1998).
Lepple-Wienhues, A., Stahl, F., Willner, U., Schäfer, R. & Wiederholt, M. Endothelin-evoked contractions in bovine ciliary muscle and trabecular meshwork: interaction with calcium, nifedipine and nickel. Curr. Eye Res. 10, 983–989 (1991).
Rucker, H. K., Wynder, H. J. & Thomas, W. E. Cellular mechanisms of CNS pericytes. Brain Res. Bull. 51, 363–369 (2000).
Cunningham, F. et al. Ensembl 2022. Nucleic Acids Res. 50, D988–D995 (2022).
Vilella, A. J. et al. EnsemblCompara GeneTrees: complete, duplication-aware phylogenetic trees in vertebrates. Genome Res. 19, 327–335 (2009).
Felsenstein, J. Evolutionary trees from gene frequencies and quantitative characters: finding maximum likelihood estimates. Evolution 35, 1229–1242 (1981).
Parins-Fukuchi, T. Use of continuous traits can improve morphological phylogenetics. Syst. Biol. 67, 328–339 (2018).
Parins‐Fukuchi, T. Bayesian placement of fossils on phylogenies using quantitative morphometric data. Evolution 72, 1801–1814 (2018).
Caumul, R. & Polly, P. D. Phylogenetic and environmental components of morphological variation: skull, mandible, and molar shape in marmots (Marmota, Rodentia). Evolution 59, 2460–2472 (2005).
Schliep, K. P. phangorn: phylogenetic analysis in R. Bioinformatics 27, 592–593 (2011).
Robinson, D. F. & Foulds, L. R. Comparison of phylogenetic trees. Math. Biosci. 53, 131–147 (1981).
Kuhner, M. K. & Felsenstein, J. A simulation comparison of phylogeny algorithms under equal and unequal evolutionary rates. Mol. Biol. Evol. 11, 459–468 (1994).
Smith, S. A. & Dunn, C. W. Phyutility: a phyloinformatics tool for trees, alignments and molecular data. Bioinformatics 24, 715–716 (2008).
Young, M. D., Wakefield, M. J., Smyth, G. K. & Oshlack, A. Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol. 11, R14 (2010).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. 57, 289–300 (1995).
Paradis, E. & Schliep, K. ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35, 526–528 (2019).
Acknowledgements
We express our gratitude to members of the Dunn lab, including S. Church, for their invaluable feedback throughout this project. We would also like to thank J. Musser, G. Wagner, D. Stadtmauer, A. Chavan, D. Adams and L. Revell for insightful comments that greatly improved our manuscript and analyses. We thank the Yale Center for Research Computing for guidance and the cloud resources provided as a part of the AWS Cloud Credit for Research Program at Yale. J.L.M. acknowledges funding from the Gruber Foundation (Gruber Science Fellowship) and the Natural Sciences and Engineering Research Council of Canada (NSERC PGS-D).
Author information
Authors and Affiliations
Contributions
J.L.M. and C.W.D. conceptualized this study and developed its methodology. J.L.M. designed and conducted the formal analyses, investigation and visualization, wrote the manuscript and performed revisions. C.W.D. additionally supervised the project and contributed text, feedback and edits.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Ecology & Evolution thanks Xavier Grau-Bové and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Matrices created at each step of the workflow.
Each grey box represents a matrix, with the number of cells indicated. Matrices were processed separately for each species (‘Within-Species Analysis’ - steps 1-4), then combined into a single multi-species matrix (‘Cross-Species Analysis’ - steps 5-6) and rotated (step 7) to produce a 919 cell 919 principal component matrix. The matrices used for Figures 2–5 were subsequently built by subsetting from a 919 cell 919 PC matrix. Note that the step 7 matrix is re-calculated for each figure. The PCA (via the R function ‘prcomp’) is reproducible, but small differences are possible due to machine rounding error. Because of this, all input files for each figure are provided in the git repo (https://github.com/dunnlab/cellphylo) to ensure reproducibility. A step-by-step walk through to produce the results described by Figures 2, 4 and 5 are provided in the repo. The animal silhouettes were from PhyloPic (https://www.phylopic.org).
Extended Data Fig. 2 Cells cluster into multi-species cell type clusters after cross-species integration.
A UMAP plot of cells, colored by A. species identity and B. cell type identity. macF, Macaca fascicularis, macM, Macaca mulatta, CollectorChn, collector channel cell, CollectorChnAqVein, collector channel aqueous vein cell, Endo, endothelium, Endo-Corneal, corneal endothelium, Endo-Schlemms, Schlemm’s canal endothelium, Endo-Vasc, vascular endothelium, Epi-CiliaryNonPigment, non-pigmented ciliary epithelium, Epi-CiliaryPigment, pigmented ciliary epithelium, Epi-Corneal, corneal epithelium, Epi-Pigment, pigmented epithelium, JCT, juxtacanalicular tissue, NKT, natural killer T cell, my, myelinating, nmy, non-myelinating.
Extended Data Fig. 3 A cell phylogeny of 92 aqueous humor cells.
This phylogeny was created using the same methods as the 54 cell phylogeny of Figure 4, but was inferred from the 92 cell 20 PC matrix (Extended Data Fig. 1, step C1.1) that included one cell per cell type group per species for all 92 cell type groups. This matrix is more encompassing than the 54 cell 20 PC matrix (Extended Data Fig. 1, step C1.2), as it includes the unstable cell type groups that were excluded from the final analysis (Methods). Jumble scores are plotted at the nodes. The scale bar indicates units of expected evolutionary change. H. sap., Homo sapiens, M. fas., Macaca fascicularis, M. mul., Macaca mulatta, M. mus., Mus musculus, Sus. scr., Sus scrofa, my, myelinating, nmy, non-myelinating, JCT, juxtacanalicular tissue, NKT, natural killer T cell.
Extended Data Fig. 4 Felsenstein’s bootstrap is not a suitable measure of biological repeatability for a cell phylogeny.
The 54 cell phylogeny (Fig. 4) is annotated with Felsenstein’s bootstrap scores calculated from scjackknife trees. While scjackknife trees still produced cell type clades, cell clades among scjackknife trees frequently varied by a few cells. This subtle variability is not well captured by traditional bootstrap scores, which mark clades as present/absent based on the presence of all tips, without acknowledging the degree of similarity. Species and cell type groups are labeled at the tips; numbers refer to the cluster number of the cell type group. Cell type clades are indicated by vertical bars. The scale bar indicates units of expected evolutionary change. H. sap., Homo sapiens, M. fas., Macaca fascicularis, M. mul., Macaca mulatta, M. mus., Mus musculus, Sus. scr., Sus scrofa, JCT, juxtacanalicular tissue, my, myelinating, nmy, non-myelinating, NKT, natural killer T cell.
Extended Data Fig. 5 Leaf stability index by cell type.
Leaf stability indices (LSI) calculated from the Fig. 4 phylogeny are highly concordant with negligible spread within each cell type clade, with the exception of immune cells (NKT and macrophages). Mean LSI for each cell type is plotted and vertical lines indicate standard deviation. The number of tips per cell type label is indicated along the bottom as ‘n tips’. The standard deviation for n tips < 3 is not shown. Cell type labels are labeled along the x-axis. Superclades are indicated with horizontal lines. Endo-Vasc, vascular endothelium, CollectorChn, collector channel cell, NKT, natural killer T cell, JCT, juxtacanalicular tissue, Endo-Corneal, corneal endothelium, my, myelinating, nmy, non-myelinating.
Extended Data Fig. 6 The averaged tree exhibits high tip stability.
The leaf stability index (LSI) is plotted as tip color onto the Fig. 5 tree. Most tips exhibit an LSI close to 1, the maximum score. Species and cell type groups are labeled at the tips, with numbers referring to the cluster number of the cell type group. Cell type clades are indicated by vertical bars. The scale bar indicates units of expected evolutionary change. Inset: Mean LSI for each cell type is plotted and vertical lines indicate standard deviation. The number of tips per cell type label is indicated along the bottom as ‘n tips’. The standard deviation for n tips < 3 is not shown. Cell type labels are labeled along the x-axis. Superclades are indicated with horizontal lines. H. sap., Homo sapiens, M. fas., Macaca fascicularis, M. mul., Macaca mulatta, M. mus., Mus musculus, Sus. scr., Sus scrofa, my, myelinating, nmy, non-myelinating, JCT, juxtacanalicular tissue, CC, collector channel, NKT, natural killer T cell, Endo-Vasc, vascular endothelium, CollectorChn, collector channel cell, Endo-Corneal, corneal endothelium.
Extended Data Fig. 7 Neighbour joining tree calculated from averaged expression levels.
The neighbour joining method was used to calculate a phenetic tree from the averaged matrix (Extended Data Fig. 1, step D1) used to infer the Fig. 5 cell phylogeny. Cell type clades are indicated with vertical bars. Species and cell type groups are labeled at the tips, with numbers indicating the cluster number of the cell type group. H. sap., Homo sapiens, M. fas., Macaca fascicularis, M. mul., Macaca mulatta, M. mus., Mus musculus, Sus. scr., Sus scrofa, my, myelinating, nmy, non-myelinating, JCT, juxtacanalicular tissue, CC, collector channel, VE, vascular endothelium, NKT, natural killer T cell.
Extended Data Fig. 8 UPGMA tree calculated from averaged expression levels.
UPGMA was used to calculate a phenetic tree from the averaged matrix (Extended Data Fig. 1, step D1) used to infer the Fig. 5 cell phylogeny. Species and cell type groups are labeled at the tips, with numbers indicating the cluster number of the cell type group. Cell type clades are indicated with vertical bars. Red dots highlight cells that failed to group with their corresponding cell type clade. The Schwann cells and immune cells superclades are paraphyletic, as indicated with dashed lines. H. sap., Homo sapiens, M. fas., Macaca fascicularis, M. mul., Macaca mulatta, M. mus., Mus musculus, Sus. scr., Sus scrofa, my, myelinating, nmy, non-myelinating, JCT, juxtacanalicular tissue, CC, collector channel, VE, vascular endothelium, NKT, natural killer T cell.
Extended Data Fig. 9 WPGMA tree calculated from averaged expression levels.
WPGMA was used to calculate a phenetic tree from the averaged matrix (Extended Data Fig. 1, step D1) used to infer the Fig. 5 cell phylogeny. Species and cell type groups are labeled at the tips, with numbers indicating the cluster number of the cell type group. Cell type clades are indicated with vertical bars. Red dots highlight cells that failed to group with their corresponding cell type clade. The vessel endothelia superclade is polyphyletic, as indicated with dashed lines. H. sap., Homo sapiens, M. fas., Macaca fascicularis, M. mul., Macaca mulatta, M. mus., Mus musculus, Sus. scr., Sus scrofa, my, myelinating, nmy, non-myelinating, JCT, juxtacanalicular tissue, CC, collector channel, VE, vascular endothelium, NKT, natural killer T cell.
Extended Data Fig. 10 The topologies of the cell phylogeny and neighbour joining tree are broadly similar.
Neighbour joining was used to create scjackknife trees. The resulting scjackknife scores were plotted onto the Fig. 5 cell phylogeny. There is broad agreement at nodes defining cell type clades and the relationships between them. H. sap., Homo sapiens, M. fas., Macaca fascicularis, M. mul., Macaca mulatta, M. mus., Mus musculus, Sus. scr., Sus scrofa, my, myelinating, nmy, non-myelinating, JCT, juxtacanalicular tissue, CC, collector channel, NKT, natural killer T cell.
Supplementary information
Supplementary Tables
Supplementary Tables 1–6.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mah, J.L., Dunn, C.W. Cell type evolution reconstruction across species through cell phylogenies of single-cell RNA sequencing data. Nat Ecol Evol 8, 325–338 (2024). https://doi.org/10.1038/s41559-023-02281-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-023-02281-9