Abstract
Background
Distinguishing between true indolent and potentially life-threatening prostate cancer is challenging in tumours displaying clinicopathologic features associated with low or intermediate risk of relapse. Several somatic DNA copy number alterations (CNAs) have been identified as potential prognostic biomarkers, but the standard cytogenetic method to assess them has a limited multiplexing capability.
Methods
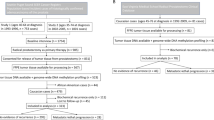
Multiplex ligation-dependent probe amplification (MLPA) targeting 14 genes was optimised to survey 448 tumours of patients with low or intermediate risk (Grade Group 1–3, Gleason score ≤7) who underwent radical prostatectomy. A 6-gene CNA classifier was developed using random survival forest and Cox proportional hazard modelling to predict biochemical recurrence.
Results
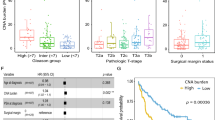
The classifier score was significantly associated with biochemical recurrence after adjusting for standard clinicopathologic variables and the known prognostic index CAPRA-S score with a hazard ratio of 2.17 and 1.80, respectively (n = 406, P < 0.01). The prognostic value of this classifier was externally validated in published CNA data from three radical prostatectomy cohorts and one radiation therapy pre-treatment biopsy cohort.
Conclusion
The 6-gene CNA classifier generated by a single MLPA assay compatible with the small quantities of DNA extracted from formalin-fixed paraffin-embedded (FFPE) tissue specimens has the potential to improve the clinical management of patients with low or intermediate risk disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Data are available from the corresponding author upon reasonable request.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71:209–49.
Fletcher SA, von Landenberg N, Cole AP, Gild P, Choueiri TK, Lipsitz SR, et al. Contemporary national trends in prostate cancer risk profile at diagnosis. Prostate Cancer Prostatic Dis. 2020;23:81–7.
Eastham JA, Auffenberg GB, Barocas DA, Chou R, Crispino T, Davis JW, et al. Clinically localized prostate cancer: AUA/ASTRO guideline, part i: introduction, risk assessment, staging, and risk-based management. J Urol. 2022;208:10–8.
Lapointe J, Li C, Giacomini CP, Salari K, Huang S, Wang P, et al. Genomic profiling reveals alternative genetic pathways of prostate tumorigenesis. Cancer Res. 2007;67:8504–10.
Bramhecha YM, Rouzbeh S, Guerard KP, Scarlata E, Brimo F, Chevalier S, et al. The combination of PTEN deletion and 16p13.3 gain in prostate cancer provides additional prognostic information in patients treated with radical prostatectomy. Mod Pathol. 2019;32:128–38.
Yoshimoto M, Cunha IW, Coudry RA, Fonseca FP, Torres CH, Soares FA, et al. FISH analysis of 107 prostate cancers shows that PTEN genomic deletion is associated with poor clinical outcome. Br J Cancer. 2007;97:678–85.
Hieronymus H, Schultz N, Gopalan A, Carver BS, Chang MT, Xiao Y, et al. Copy number alteration burden predicts prostate cancer relapse. Proc Natl Acad Sci USA. 2014;111:11139–44.
Lalonde E, Ishkanian AS, Sykes J, Fraser M, Ross-Adams H, Erho N, et al. Tumour genomic and microenvironmental heterogeneity for integrated prediction of 5-year biochemical recurrence of prostate cancer: a retrospective cohort study. Lancet Oncol. 2014;15:1521–32.
Zafarana G, Ishkanian AS, Malloff CA, Locke JA, Sykes J, Thoms J, et al. Copy number alterations of c-MYC and PTEN are prognostic factors for relapse after prostate cancer radiotherapy. Cancer. 2012;118:4053–62.
Stuppia L, Antonucci I, Palka G, Gatta V. Use of the MLPA assay in the molecular diagnosis of gene copy number alterations in human genetic diseases. Int J Mol Sci. 2012;13:3245–76.
Ebrahimizadeh W, Guerard KP, Rouzbeh S, Bramhecha YM, Scarlata E, Brimo F, et al. Design and development of a fully synthetic multiplex ligation-dependent probe amplification-based probe mix for detection of copy number alterations in prostate cancer formalin-fixed, paraffin-embedded tissue samples. J Mol Diagn. 2020;22:1246–63.
McShane LM, Altman DG, Sauerbrei W, Taube SE, Gion M, Clark GM, et al. Reporting recommendations for tumour MARKer prognostic studies (REMARK). Br J Cancer. 2005;93:387–91.
Humphrey PA, Moch H, Cubilla AL, Ulbright TM, Reuter VE. The 2016 WHO classification of tumours of the urinary system and male genital organs-part B: prostate and bladder tumours. Eur Urol. 2016;70:106–19.
Bramhecha YM, Guerard KP, Rouzbeh S, Scarlata E, Brimo F, Chevalier S, et al. Genomic gain of 16p13.3 in prostate cancer predicts poor clinical outcome after surgical intervention. Mol Cancer Res. 2018;16:115–23.
Ishwaran H, Kogalur UB, Blackstone EH, Lauer MS. Random survival forests. Annal Appl Stat. 2008;2:841–60.
Ogluszka M, Orzechowska M, Jedroszka D, Witas P, Bednarek AK. Evaluate cutpoints: adaptable continuous data distribution system for determining survival in Kaplan-Meier estimator. Comput Methods Prog Biomed. 2019;177:133–9.
Taylor BS, Schultz N, Hieronymus H, Gopalan A, Xiao Y, Carver BS, et al. Integrative genomic profiling of human prostate cancer. Cancer Cell. 2010;18:11–22.
Ross-Adams H, Lamb A, Dunning M, Halim S, Lindberg J, Massie C, et al. Integration of copy number and transcriptomics provides risk stratification in prostate cancer: a discovery and validation cohort study. EBioMedicine. 2015;2:1133–44.
Espiritu SMG, Liu LY, Rubanova Y, Bhandari V, Holgersen EM, Szyca LM, et al. The evolutionary landscape of localized prostate cancers drives clinical aggression. Cell. 2018;173:1003–13.e15
Punnen S, Freedland SJ, Presti JC, Aronson WJ, Terris MK, Kane CJ, et al. Multi-institutional validation of the CAPRA-S score to predict disease recurrence and mortality after radical prostatectomy. Eur Urol. 2014;65:1171–7.
Cooperberg MR, Pasta DJ, Elkin EP, Litwin MS, Latini DM, Du Chane J, et al. The University of California, San Francisco Cancer of the Prostate Risk Assessment score: a straightforward and reliable preoperative predictor of disease recurrence after radical prostatectomy. J Urol. 2005;173:1938–42.
D’Amico AV, Whittington R, Malkowicz SB, Schultz D, Blank K, Broderick GA, et al. Biochemical outcome after radical prostatectomy, external beam radiation therapy, or interstitial radiation therapy for clinically localized prostate cancer. J Am Med Assoc. 1998;280:969–74.
Mohler JL, Antonarakis ES, Armstrong AJ, D’Amico AV, Davis BJ, Dorff T, et al. Prostate cancer, version 2.2019, NCCN clinical practice guidelines in oncology. J Natl Compr Cancer Netw. 2019;17:479–505.
Sun J, Liu W, Adams TS, Sun J, Li X, Turner AR, et al. DNA copy number alterations in prostate cancers: a combined analysis of published CGH studies. Prostate. 2007;67:692–700.
Wu Y, Tedesco L, Lucia K, Schlitter AM, Garcia JM, Esposito I, et al. RSUME is implicated in tumorigenesis and metastasis of pancreatic neuroendocrine tumors. Oncotarget. 2016;7:57878–93.
Huang J, Yan J, Zhang J, Zhu S, Wang Y, Shi T, et al. SUMO1 modification of PTEN regulates tumorigenesis by controlling its association with the plasma membrane. Nat Commun. 2012;3:911.
Fukasawa S, Kino M, Kobayashi M, Suzuki H, Komiya A, Imamoto T, et al. Genetic changes in pT2 and pT3 prostate cancer detected by comparative genomic hybridization. Prostate Cancer Prostatic Dis. 2008;11:303–10.
van Wietmarschen N, Nathan WJ, Nussenzweig A. The WRN helicase: resolving a new target in microsatellite unstable cancers. Curr Opin Genet Dev. 2021;71:34–8.
Hart SN, Ellingson MS, Schahl K, Vedell PT, Carlson RE, Sinnwell JP, et al. Determining the frequency of pathogenic germline variants from exome sequencing in patients with castrate-resistant prostate cancer. BMJ Open. 2016;6:e010332.
Choucair KA, Guerard KP, Ejdelman J, Chevalier S, Yoshimoto M, Scarlata E, et al. The 16p13.3 (PDPK1) genomic gain in prostate cancer: a potential role in disease progression. Transl Oncol. 2012;5:453–60.
Fromont G, Godet J, Peyret A, Irani J, Celhay O, Rozet F, et al. 8q24 amplification is associated with Myc expression and prostate cancer progression and is an independent predictor of recurrence after radical prostatectomy. Hum Pathol. 2013;44:1617–23.
Kluth M, Harasimowicz S, Burkhardt L, Grupp K, Krohn A, Prien K, et al. Clinical significance of different types of p53 gene alteration in surgically treated prostate cancer. Int J Cancer. 2014;135:1369–80.
Bottcher R, Kweldam CF, Livingstone J, Lalonde E, Yamaguchi TN, Huang V, et al. Cribriform and intraductal prostate cancer are associated with increased genomic instability and distinct genomic alterations. BMC Cancer. 2018;18:8.
Trudel D, Downes MR, Sykes J, Kron KJ, Trachtenberg J, van der Kwast TH. Prognostic impact of intraductal carcinoma and large cribriform carcinoma architecture after prostatectomy in a contemporary cohort. Eur J Cancer. 2014;50:1610–6.
Kweldam CF, Wildhagen MF, Steyerberg EW, Bangma CH, van der Kwast TH, van Leenders GJ. Cribriform growth is highly predictive for postoperative metastasis and disease-specific death in Gleason score 7 prostate cancer. Mod Pathol. 2015;28:457–64.
Tilki D, Chen MH, Wu J, Huland H, Graefen M, Wiegel T, et al. Adjuvant versus early salvage radiation therapy for men at high risk for recurrence following radical prostatectomy for prostate cancer and the risk of death. J Clin Oncol. 2021;39:2284–93.
Boutros PC, Fraser M, Harding NJ, de Borja R, Trudel D, Lalonde E, et al. Spatial genomic heterogeneity within localized, multifocal prostate cancer. Nat Genet. 2015;47:736–45.
Chen Y, Sadasivan SM, She R, Datta I, Taneja K, Chitale D, et al. Breast and prostate cancers harbor common somatic copy number alterations that consistently differ by race and are associated with survival. BMC Med Genomics. 2020;13:116.
Liu W, Hou J, Petkewicz J, Na R, Wang CH, Sun J, et al. Feasibility and performance of a novel probe panel to detect somatic DNA copy number alterations in clinical specimens for predicting prostate cancer progression. Prostate. 2020;80:1253–62.
Wang S, Li H, Song M, Tao Z, Wu T, He Z, et al. Copy number signature analysis tool and its application in prostate cancer reveals distinct mutational processes and clinical outcomes. PLoS Genet. 2021;17:e1009557.
Wang X, Grasso CS, Jordahl KM, Kolb S, Nyame YA, Wright JL, et al. Copy number alterations are associated with metastatic-lethal progression in prostate cancer. Prostate Cancer Prostatic Dis. 2020;23:494–506.
Acknowledgements
Authors thank all the patients who consented to provide prostate tissues for research.
Funding
This study was funded by the Movember Foundation through Prostate Cancer Canada (grant #T2014.01) and carried out as a part of personalised risk stratification for patients with early prostate cancer (PRONTO). WE received a studentship and doctoral training award from the Research Institute of McGill University Health Centre and Fonds de la Recherche du Québec–Santé (FRQS), respectively.
Author information
Authors and Affiliations
Contributions
Conception and design: WE, KPG and JL. Development of methodology: WE, KPG and JL. Acquisition of the data: WE, KPG, SR, ES, FB, PGP, TJ, LH, AGA, AYL, DMB, JMSB, SC and JL. Analysis and interpretation of the data: WE, KPG, AYL and JL. Writing, review, and/or revision of the manuscript: WE, KPG, SR, ES, FB, PGP, TJ, LH, AGA, AYL, DMB, JMSB, SC and JL. Administrative, technical, or material support: ES, LH, TJ and AGA. Study supervision: JL.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The study was approved by the Research Ethics Board of McGill University Health Centre (Quebec, Canada, BDM-10-115) and by the Queen’s University Research Ethics Board (Ontario, Canada, PATH-144-14). Informed consent was obtained from participants and the study was performed in accordance with the Declaration of Helsinki.
Consent for publication
Not applicable.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ebrahimizadeh, W., Guérard, KP., Rouzbeh, S. et al. A DNA copy number alteration classifier as a prognostic tool for prostate cancer patients. Br J Cancer 128, 2165–2174 (2023). https://doi.org/10.1038/s41416-023-02236-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-023-02236-8