Abstract
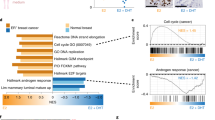
Estrogen receptor alpha gene (ESR1) mutations occur frequently in ER-positive metastatic breast cancer, and confer clinical resistance to aromatase inhibitors. Expression of the ESR1 Y537S mutation induced an epithelial–mesenchymal transition (EMT) with cells exhibiting enhanced migration and invasion potential in vitro. When small subpopulations of Y537S ESR1 mutant cells were injected along with WT parental cells, tumor growth was enhanced with mutant cells becoming the predominant population in distant metastases. Y537S mutant primary xenograft tumors were resistant to the antiestrogen tamoxifen (Tam) as well as to estradiol (E2) withdrawal. Y537S ESR1 mutant primary tumors metastasized efficiently in the absence of E2; however, Tam treatment significantly inhibited metastasis to distant sites. We identified a nine-gene expression signature, which predicted clinical outcomes of ER-positive breast cancer patients, as well as breast cancer metastasis to the lung. Androgen receptor (AR) protein levels were increased in mutant models, and the AR agonist dihydrotestosterone significantly inhibited estrogen-regulated gene expression, EMT, and distant metastasis in vivo, suggesting that AR may play a role in distant metastatic progression of ESR1 mutant tumors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
15 November 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41388-021-02104-w
References
Howlader N, Noone AM, Krapcho M, Miller D, Bishop K, Altekruse SF, et al. SEER cancer statistics review, 1975-2013. Bethesda, MD: National Cancer Institute; 2016. http://seer.cancer.gov/csr/1975_2013/.
Klein CA. Selection and adaptation during metastatic cancer progression. Nature. 2013;501:365–72.
Fuqua SA. The role of estrogen receptors in breast cancer metastasis. J Mammary Gland Biol Neoplasia. 2001;6:407–17.
Herynk MH, Fuqua SA. Estrogen receptor mutations in human disease. Endocr Rev. 2004;25:869–98.
Jeselsohn R, Bergholz JS, Pun M, Cornwell M, Liu W, Nardone A, et al. Allele-specific chromatin recruitment and therapeutic vulnerabilities of ESR1 activating mutations. Cancer Cell. 2018;33:173–86.e5.
Zhang QX, Borg A, Wolf DM, Oesterreich S, Fuqua SA. An estrogen receptor mutant with strong hormone-independent activity from a metastatic breast cancer. Cancer Res. 1997;57:1244–9.
Razavi P, Chang MT, Xu G, Bandlamudi C, Ross DS, Vasan N, et al. The genomic landscape of endocrine-resistant advanced breast cancers. Cancer Cell. 2018;34:427–38.e6.
Fuqua SA, Gu G, Rechoum Y. Estrogen receptor (ER) alpha mutations in breast cancer: hidden in plain sight. Breast Cancer Res Treat. 2014;144:11–9.
Gelsomino L, Gu G, Rechoum Y, Beyer AR, Pejerrey SM, Tsimelzon A, et al. ESR1 mutations affect anti-proliferative responses to tamoxifen through enhanced cross-talk with IGF signaling. Breast Cancer Res Treat. 2016;157:253–65.
Takeshita T, Yamamoto Y, Yamamoto-Ibusuki M, Inao T, Sueta A, Fujiwara S, et al. Droplet digital polymerase chain reaction assay for screening of ESR1 mutations in 325 breast cancer specimens. Transl Res. 2015;166:540–53.e2.
Schiavon G, Hrebien S, Garcia-Murillas I, Cutts RJ, Pearson A, Tarazona N, et al. Analysis of ESR1 mutation in circulating tumor DNA demonstrates evolution during therapy for metastatic breast cancer. Sci Transl Med. 2015;7:313ra182.
O’Leary B, Cutts RJ, Liu Y, Hrebien S, Huang X, Fenwick K, et al. The genetic landscape and clonal evolution of breast cancer resistance to palbociclib plus fulvestrant in the PALOMA-3 trial. Cancer Discov. 2018;8:1390–403.
Chandarlapaty S, Chen D, He W, Sung P, Samoila A, You D, et al. Prevalence of ESR1 mutations in cell-free DNA and outcomes in metastatic breast cancer: a secondary analysis of the BOLERO-2 clinical trial. JAMA Oncol. 2016;2:1310–5.
De Amicis F, Thirugnansampanthan J, Cui Y, Selever J, Beyer A, Parra I, et al. Androgen receptor overexpression induces tamoxifen resistance in human breast cancer cells. Breast Cancer Res Treat. 2010;121:1–11.
Ciupek A, Rechoum Y, Gu G, Gelsomino L, Beyer AR, Brusco L, et al. Androgen receptor promotes tamoxifen agonist activity by activation of EGFR in ERalpha-positive breast cancer. Breast Cancer Res Treat. 2015;154:225–37.
Rechoum Y, Rovito D, Iacopetta D, Barone I, Ando S, Weigel NL, et al. AR collaborates with ERalpha in aromatase inhibitor-resistant breast cancer. Breast Cancer Res Treat. 2014;147:473–85.
Selever J, Gu G, Lewis MT, Beyer A, Herynk MH, Covington KR, et al. Dicer-mediated upregulation of BCRP confers tamoxifen resistance in human breast cancer cells. Clin Cancer Res. 2011;17:6510–21.
Park SH, Cheung LW, Wong AS, Leung PC. Estrogen regulates Snail and Slug in the down-regulation of E-cadherin and induces metastatic potential of ovarian cancer cells through estrogen receptor alpha. Mol Endocrinol. 2008;22:2085–98.
Taube JH, Herschkowitz JI, Komurov K, Zhou AY, Gupta S, Yang J, et al. Core epithelial-to-mesenchymal transition interactome gene-expression signature is associated with claudin-low and metaplastic breast cancer subtypes. Proc Natl Acad Sci USA. 2010;107:15449–54.
Wallden B, Storhoff J, Nielsen T, Dowidar N, Schaper C, Ferree S, et al. Development and verification of the PAM50-based Prosigna breast cancer gene signature assay. BMC Med Genomics. 2015;8:54.
Li S, Shen D, Shao J, Crowder R, Liu W, Prat A, et al. Endocrine-therapy-resistant ESR1 variants revealed by genomic characterization of breast-cancer-derived xenografts. Cell Rep. 2013;4:1116–30.
Harris LN, Ismaila N, McShane LM, Andre F, Collyar DE, Gonzalez-Angulo AM, et al. Use of biomarkers to guide decisions on adjuvant systemic therapy for women with early-stage invasive breast cancer: American Society of Clinical Oncology Clinical Practice Guideline. J Clin Oncol. 2016;34:1134–50.
Mihaly Z, Kormos M, Lanczky A, Dank M, Budczies J, Szasz MA, et al. A meta-analysis of gene expression-based biomarkers predicting outcome after tamoxifen treatment in breast cancer. Breast Cancer Res Treat. 2013;140:219–32.
Pidugu VK, Wu MM, Yen AH, Pidugu HB, Chang KW, Liu CJ, et al. IFIT1 and IFIT3 promote oral squamous cell carcinoma metastasis and contribute to the anti-tumor effect of gefitinib via enhancing p-EGFR recycling. Oncogene. 2019;38:3232–47.
Li C, Wang J, Zhang H, Zhu M, Chen F, Hu Y, et al. Interferon-stimulated gene 15 (ISG15) is a trigger for tumorigenesis and metastasis of hepatocellular carcinoma. Oncotarget. 2014;5:8429–41.
Hiratsuka S, Watanabe A, Sakurai Y, Akashi-Takamura S, Ishibashi S, Miyake K, et al. The S100A8-serum amyloid A3-TLR4 paracrine cascade establishes a pre-metastatic phase. Nat Cell Biol. 2008;10:1349–55.
Chiang KC, Huang ST, Wu RC, Huang SC, Yeh TS, Chen MH, et al. Interferon alpha-inducible protein 27 is an oncogene and highly expressed in cholangiocarcinoma patients with poor survival. Cancer Manag Res. 2019;11:1893–905.
Jiang J, Liu W, Guo X, Zhang R, Zhi Q, Ji J, et al. IRX1 influences peritoneal spreading and metastasis via inhibiting BDKRB2-dependent neovascularization on gastric cancer. Oncogene. 2011;30:4498–508.
Curtis C, Shah SP, Chin SF, Turashvili G, Rueda OM, Dunning MJ, et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature. 2012;486:346–52.
Bos PD, Zhang XH, Nadal C, Shu W, Gomis RR, Nguyen DX, et al. Genes that mediate breast cancer metastasis to the brain. Nature. 2009;459:1005–9.
Ponnusamy S, Asemota S, Schwartzberg LS, Guestini F, McNamara KM, Pierobon M, et al. Androgen receptor is a non-canonical inhibitor of wild-type and mutant estrogen receptors in hormone receptor-positive breast cancers. iScience. 2019;21:341–58.
Martin LA, Ribas R, Simigdala N, Schuster E, Pancholi S, Tenev T, et al. Discovery of naturally occurring ESR1 mutations in breast cancer cell lines modelling endocrine resistance. Nat Commun. 2017;8:1865.
Felipe JL, Nofech-Mozes S, Bayani J, Bartlett JM. EMT in Breast Carcinoma-A Review. J Clin Med. 2016;5:65. https://doi.org/10.3390/jcm5070065.
Dhasarathy A, Kajita M, Wade PA. The transcription factor snail mediates epithelial to mesenchymal transitions by repression of estrogen receptor-alpha. Mol Endocrinol. 2007;21:2907–18.
Li Y, Wu Y, Abbatiello TC, Wu WL, Kim JR, Sarkissyan M, et al. Slug contributes to cancer progression by direct regulation of ERalpha signaling pathway. Int J Oncol. 2015;46:1461–72.
Jin L, Han B, Siegel E, Cui Y, Giuliano A, Cui X. Breast cancer lung metastasis: Molecular biology and therapeutic implications. Cancer Biol Ther. 2018;19:858–68.
Krop I, Abramson V, Colleoni M, Traina T, Holmes F, Estevez L, et al. Results from a randomized placebo-controlled phase 2 trial evaluating exemestane±enzalutamide in patients with hormone receptor–positive breast cancer [abstract]. In: Proceedings of the 2017 San Antonio breast cancer symposium. Philadelphia, PA: AACR; 2018.
Hickey TE, Robinson JL, Carroll JS, Tilley WD. Minireview: the androgen receptor in breast tissues: growth inhibitor, tumor suppressor, oncogene? Mol Endocrinol. 2012;26:1252–67.
Cochrane DR, Bernales S, Jacobsen BM, Cittelly DM, Howe EN, D’Amato NC, et al. Role of the androgen receptor in breast cancer and preclinical analysis of enzalutamide. Breast Cancer Res. 2014;16:R7.
Rangel N, Rondon-Lagos M, Annaratone L, Osella-Abate S, Metovic J, Mano MP, et al. The role of the AR/ER ratio in ER-positive breast cancer patients. Endocr Relat Cancer. 2018;25:163–72.
Narayanan R, Coss CC, Dalton JT. Development of selective androgen receptor modulators (SARMs). Mol Cell Endocrinol. 2018;465:134–42.
Kim JA, Tan Y, Wang X, Cao X, Veeraraghavan J, Liang Y, et al. Comprehensive functional analysis of the tousled-like kinase 2 frequently amplified in aggressive luminal breast cancers. Nat Commun. 2016;7:12991.
Barone I, Brusco L, Gu G, Selever J, Beyer A, Covington KR, et al. Loss of Rho GDIalpha and resistance to tamoxifen via effects on estrogen receptor alpha. J Natl Cancer Inst. 2011;103:538–52.
Gates LA, Gu G, Chen Y, Rohira AD, Lei JT, Hamilton RA, et al. Proteomic profiling identifies key coactivators utilized by mutant ERalpha proteins as potential new therapeutic targets. Oncogene. 2018;37:4581–98.
Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F. Genome engineering using the CRISPR-Cas9 system. Nat Protoc. 2013;8:2281–308.
Kollipara RK, Kittler R. An integrated functional genomic analysis identifies the antitumorigenic mechanism of action for PPARgamma in lung cancer cells. Genom Data. 2015;3:80–6.
Lambert JR, Nordeen SK. Analysis of steroid hormone-induced histone acetylation by chromatin immunoprecipitation assay. In: Lieberman BA, editor. Steroid receptor methods: protocols and assays. Totowa, NJ: Humana Press; 2001. pp. 273–81.
Sinn BV, Fu C, Lau R, Litton J, Tsai TH, Murthy R, et al. SETER/PR: a robust 18-gene predictor for sensitivity to endocrine therapy for metastatic breast cancer. NPJ Breast Cancer. 2019;5:16.
Gyorffy B, Surowiak P, Budczies J, Lanczky A. Online survival analysis software to assess the prognostic value of biomarkers using transcriptomic data in non-small-cell lung cancer. PLoS ONE. 2013;8:e82241.
Acknowledgements
The authors would like to thank Drs. Chandandeep Nagi for his pathology expertise and Sasha Pejerrey for initial edits of the paper. We acknowledge the joint participation with the Adrienne Helis Malvin Medical Research Foundation through its direct engagement in the continuous active conduct of medical research in conjunction with BCM and the Cancer Program. We acknowledge the joint collaboration with WFS through the CPRIT MIRA grant mechanism.
Funding
Breast Cancer Research Foundation BCRF 19-055, NIH R01 CA207270, NIH R01CA072038, Cancer Prevention Institute of Texas CPRIT MIRA (Kelly Hunt, PI) to SAWF; Adrienne Helis Malvin Medical Research Foundation Cancer Program M-2017 to CEF; P30 CA125123-05 Dan L Duncan Cancer Center Cell-based assay screening core; Translational Breast Cancer Research Training Program T32-CA203690-02; Predoctoral Institutional Research Training Grant (T32) Training Grant 5T32GM008231-30; T32 2389301302, Cancer Prevention Institute of Texas (CPRIT) RP170005; NIEHS grants P30 ES030285, P42 ES027725; U01 CA217842, Susan G. Komen Breast Cancer Foundation SAC110052, Breast Cancer Research Foundation BCRF‐19-110. Grant number RP180712/CPRIT-MIRA.
Author information
Authors and Affiliations
Contributions
Conception and design: GG, SAWF. Development of methodology: GG, LT, DL, SKH, CC. Acquisition of data: GG, SKH, HCL, TG, YR, RK, LD, ARB, MG, DD, LG, J-AK, HJH, NMF, JX, AA, CEF, DGE, SLG. Analysis and interpretation of data (e.g., statistical analysis, biostatistics, and computation analysis): GG, SAWF, LT, TG, AT, SGH, CC, SLG, SKH, WFS. Writing, review, and/or revision of the paper: GG, SAWF, TG, ARB. Administrative, technical, or material support: ARB, SA, SL, GBM, FJ, XH-FZ. Study supervision: SAWF, GG.
Corresponding author
Ethics declarations
Conflict of interest
(1) CEF discloses: an equity position in Coactigon, Inc. (2) FJ discloses: (a) Grant/Research Funding (Institutional): Novartis, Genentech, BioMed Valley Discoveries, Plexxikon, Deciphera, Piqur, Symphogen, Bayer, FujiFilm Corporation and Upsher-Smith Laboratories, Astex, Asana, Astellas, Agios, Proximagen, Bristol-Myers Squibb; (b) Scientific Advisory Board: Deciphera, IFM Therapeutics, Synlogic, Guardant Health, Ideaya, PureTech Health; (c) Paid Consultant: Trovagene, Immunomet, Jazz Pharmaceuticals, Sotio; (d) Ownership Interests: Trovagene. (3) GBM discloses: (a) SAB/Consultant: AstraZeneca, Chrysallis Biotechnology, GSK, ImmunoMET, Ionis, Lilly, PDX Pharmaceuticals, Signalchem Lifesciences, Symphogen, Tarveda, Turbine, Zentalis Pharmaceuticals; (b) Stock/Options/Financial: Catena Pharmaceuticals, ImmunoMet, SignalChem, Tarveda; (c) Licensed Technology: HRD assay to Myriad Genetics, DSP patents with Nanostring. (4) SL discloses: (a) The Washington Unversity PDX development and trial center is supported by NIH 3U54CA224083-02S3. (b) SL has received license fee from Envigo. He received research funding from Pfizer, Takeda Oncology, and Zenopharm, outside of this project.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Gu, G., Tian, L., Herzog, S.K. et al. Hormonal modulation of ESR1 mutant metastasis. Oncogene 40, 997–1011 (2021). https://doi.org/10.1038/s41388-020-01563-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-020-01563-x
This article is cited by
-
Proteomic profiling reveals that ESR1 mutations enhance cyclin-dependent kinase signaling
Scientific Reports (2024)
-
Therapeutic resistance to anti-oestrogen therapy in breast cancer
Nature Reviews Cancer (2023)
-
ESR1 mutations and therapeutic resistance in metastatic breast cancer: progress and remaining challenges
British Journal of Cancer (2022)
-
Cluster analyses of the TCGA and a TMA dataset using the coexpression of HSP27 and CRYAB improves alignment with clinical-pathological parameters of breast cancer and suggests different epichaperome influences for each sHSP
Cell Stress and Chaperones (2022)
-
Genome engineering for estrogen receptor mutations reveals differential responses to anti-estrogens and new prognostic gene signatures for breast cancer
Oncogene (2022)