Abstract
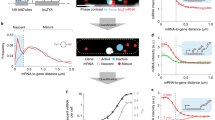
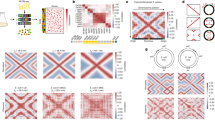
Gene expression is inherently stochastic; precise gene regulation by transcription factors is important for cell-fate determination. Many transcription factors regulate their own expression, suggesting that autoregulation counters intrinsic stochasticity in gene expression. Using a new strategy, cotranslational activation by cleavage (CoTrAC), we probed the stochastic expression dynamics of cI, which encodes the bacteriophage λ repressor CI, a fate-determining transcription factor. CI concentration fluctuations influence both lysogenic stability and induction of bacteriophage λ. We found that the intrinsic stochasticity in cI expression was largely determined by CI expression level irrespective of autoregulation. Furthermore, extrinsic, cell-to-cell variation was primarily responsible for CI concentration fluctuations, and negative autoregulation minimized CI concentration heterogeneity by counteracting extrinsic noise and introducing memory. This quantitative study of transcription factor expression dynamics sheds light on the mechanisms cells use to control noise in gene regulatory networks.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
15 July 2012
In the version of this article initially published online, equations 1 and 3 contained errors. These have been corrected for the print, PDF and HTML versions of this article.
References
Shen-Orr, S.S. et al. Network motifs in the transcriptional regulation network of Escherichia coli. Nat. Genet. 31, 64–68 (2002).
Ptashne, M. A Genetic Switch: Phage Lambda Revisited 3rd edn. (Cold Spring Harbor Laboratory Press, 2004).
Yu, J. et al. Probing gene expression in live cells, one protein molecule at a time. Science 311, 1600–1603 (2006).
Tobias, J.W. et al. The N-end rule in bacteria. Science 254, 1374–1377 (1991).
Pedraza, J.M. & Paulsson, J. Effects of molecular memory and bursting on fluctuations in gene expression. Science 319, 339–343 (2008).
Meyer, B.J., Maurer, R. & Ptashne, M. Gene regulation at the right operator (OR) of bacteriophage λ. II. OR1, OR2, and OR3: their roles in mediating the effects of repressor and cro. J. Mol. Biol. 139, 163–194 (1980).
Sarai, A. & Takeda, Y. λ repressor recognizes the approximately 2-fold symmetric half-operator sequences asymmetrically. Proc. Natl. Acad. Sci. USA 86, 6513–6517 (1989).
Levine, A., Bailone, A. & Devoret, R. Cellular levels of the prophage λ and 434 repressors. J. Mol. Biol. 131, 655–661 (1979).
Reichardt, L. & Kaiser, A.D. Control of λ repressor synthesis. Proc. Natl. Acad. Sci. USA 68, 2185–2189 (1971).
Wang, J. & Wolynes, P. Intermittency of single-molecule reaction dynamics in fluctuating environments. Phys. Rev. Lett. 74, 4317–4320 (1995).
Ozbudak, E.M. et al. Regulation of noise in the expression of a single gene. Nat. Genet. 31, 69–73 (2002).
Golding, I. et al. Real-time kinetics of gene activity in individual bacteria. Cell 123, 1025–1036 (2005).
Huh, D. & Paulsson, J. Nongenetic heterogeneity from stochastic partitioning at cell division. Nat. Genet. 43, 95–100 (2011).
Elowitz, M.B. et al. Stochastic gene expression in a single cell. Science 297, 1183–1186 (2002).
Rosenfeld, N. et al. Gene regulation at the single-cell level. Science 307, 1962–1965 (2005).
Swain, P.S., Elowitz, M.B. & Siggia, E.D. Intrinsic and extrinsic contributions to stochasticity in gene expression. Proc. Natl. Acad. Sci. USA 99, 12795–12800 (2002).
Taniguchi, Y. et al. Quantifying E. coli proteome and transcriptome with single-molecule sensitivity in single cells. Science 329, 533–538 (2010).
Raj, A. et al. Stochastic mRNA synthesis in mammalian cells. PLoS Biol. 4, e309 (2006).
So, L.H. et al. General properties of transcriptional time series in Escherichia coli. Nat. Genet. 43, 554–560 (2011).
Shahrezaei, V. & Swain, P.S. Analytical distributions for stochastic gene expression. Proc. Natl. Acad. Sci. USA 105, 17256–17261 (2008).
Zenklusen, D., Larson, D.R. & Singer, R.H. Single-RNA counting reveals alternative modes of gene expression in yeast. Nat. Struct. Mol. Biol. 15, 1263–1271 (2008).
Suter, D.M. et al. Mammalian genes are transcribed with widely different bursting kinetics. Science 332, 472–474 (2011).
Hornos, J.E. et al. Self-regulating gene: an exact solution. Phys. Rev. E 72, 051907 (2005) (Epub 4 November 2005).
Lepzelter, D., Kim, K.Y. & Wang, J. Dynamics and intrinsic statistical fluctuations of a gene switch. J. Phys. Chem. B 111, 10239–10247 (2007).
Feng, H., Han, B. & Wang, J. Adiabatic and nonadiabatic nonequilibrium stochastic dynamics of single regulating genes. J. Phys. Chem. B 115, 1254–1261 (2011).
Feng, H. & Wang, J. Landscape and global stability of nonadiabatic and adiabatic oscillations in a gene network. Biophys. J. 102, 1001–1010 (2012).
Choi, P.J. et al. A stochastic single-molecule event triggers phenotype switching of a bacterial cell. Science 322, 442–446 (2008).
Singh, A. & Weinberger, L.S. Stochastic gene expression as a molecular switch for viral latency. Curr. Opin. Microbiol. 12, 460–466 (2009).
Kalmar, T. et al. Regulated fluctuations in nanog expression mediate cell-fate decisions in embryonic stem cells. PLoS Biol. 7, e1000149 (2009).
Feng, H. & Wang, J. A new formulation of two-time correlation functions of Markov chains applied to gene networks. Chem. Phys. Lett. 501, 562–566 (2011).
Lu, T., Hasty, J. & Wolynes, P.G. Effective temperature in stochastic kinetics and gene networks. Biophys. J. 91, 84–94 (2006).
Dodd, I.B. et al. Cooperativity in long-range gene regulation by the λ CI repressor. Genes Dev. 18, 344–354 (2004).
Bar-Even, A. et al. Noise in protein expression scales with natural protein abundance. Nat. Genet. 38, 636–643 (2006).
Zong, C. et al. Lysogen stability is determined by the frequency of activity bursts from the fate-determining gene. Mol. Syst. Biol. 6, 440 (2010).
Wang, J. Statistics, pathways and dynamics of single molecule protein folding. J. Chem. Phys. 118, 952 (2003).
Ross, S.M. Stochastic Processes (John Wiley & Sons, 1983).
Cai, L., Friedman, N. & Xie, X.S. Stochastic protein expression in individual cells at the single-molecule level. Nature 440, 358–362 (2006).
Hawley, D.K. & McClure, W.R. Mechanism of activation of transcription initiation from the λ PRM promoter. J. Mol. Biol. 157, 493–525 (1982).
Tobias, J.W. & Varshavsky, A. Cloning and functional analysis of the ubiquitin-specific protease gene UBP1 of Saccharomyces cerevisiae. J. Biol. Chem. 266, 12021–12028 (1991).
Datsenko, K.A. & Wanner, B.L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. USA 97, 6640–6645 (2000).
Acknowledgements
We thank J. Little (University of Arizona), D. Finley (Harvard Medical School), R. Baker (John Curtin School of Medical Research) and the Yale University Coli Genetic Stock Center for strains and reagents. Funding provided by National Science Foundation CAREER Award 0746796 (to J.X.), March of Dimes Research grant 1-FY2011 (to J.X.), March of Dimes Basil O'Connor Starter Scholar Research Award 5-FY20 (to J.X.) and National Science Foundation Emerging Frontiers Award 0926287 (to J.W.).
Author information
Authors and Affiliations
Contributions
Z.H. and J.X. designed experiments. Z.H. and C.H. engineered strains and performed immunoblotting experiments. Z.H. acquired and analyzed fluorescence images. H.F., Z.H., B.H., J.X. and J.W. developed analytical methods. Z.H., H.F. and J.X. analyzed data. Z.H., J.X., H.F. and J.W. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figs 1–4, Supplementary Tables 1–5 and Supplementary Notes 1 and 2 (PDF 3229 kb)
Supplementary Movie 1
Timelapse fluorescence images for strain λWT (MOV 1422 kb)
Supplementary Movie 2
Timelapse fluorescence images for strain λr3 (MOV 2032 kb)
Supplementary Movie 3
Timelapse fluorescence images for strain λb (MOV 1457 kb)
Supplementary Movie 4
Timelapse fluorescence images for strain λ– (MOV 1083 kb)
Rights and permissions
About this article
Cite this article
Hensel, Z., Feng, H., Han, B. et al. Stochastic expression dynamics of a transcription factor revealed by single-molecule noise analysis. Nat Struct Mol Biol 19, 797–802 (2012). https://doi.org/10.1038/nsmb.2336
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2336
This article is cited by
-
From deterministic to fuzzy decision-making in artificial cells
Nature Communications (2020)
-
Cell fate potentials and switching kinetics uncovered in a classic bistable genetic switch
Nature Communications (2018)
-
Rate-limiting steps in transcription dictate sensitivity to variability in cellular components
Scientific Reports (2017)
-
Efficient and flexible implementation of Langevin simulation for gene burst production
Scientific Reports (2017)
-
Mechanical slowing-down of cytoplasmic diffusion allows in vivo counting of proteins in individual cells
Nature Communications (2016)