Abstract
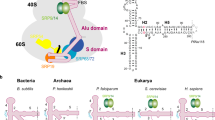
The signal recognition particle (SRP) recognizes and binds the signal sequence of nascent proteins as they emerge from the ribosome. We present here the 3.0-Å structure of a signal sequence bound to the Methanococcus jannaschii SRP core. Structural comparison with the free SRP core shows that signal-sequence binding induces formation of the GM-linker helix and a 180° flip of the NG domain—structural changes that ensure a hierarchical succession of events during protein targeting.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Egea, P.F., Stroud, R.M. & Walter, P. Curr. Opin. Struct. Biol. 15, 213–220 (2005).
Batey, R.T., Rambo, R.P., Lucast, L., Rha, B. & Doudna, J.A. Science 287, 1232–1239 (2000).
Freymann, D.M., Keenan, R.J., Stroud, R.M. & Walter, P. Nature 385, 361–364 (1997).
Keenan, R.J., Freymann, D.M., Walter, P. & Stroud, R.M. Cell 94, 181–191 (1998).
Egea, P.F. et al. Nature 427, 215–221 (2004).
Focia, P.J., Shepotinovskaya, I.V., Seidler, J.A. & Freymann, D.M. Science 303, 373–377 (2004).
Peluso, P. et al. Science 288, 1640–1643 (2000).
Siu, F.Y., Spanggord, R.J. & Doudna, J.A. RNA 13, 240–250 (2007).
Lee, H.C. & Bernstein, H.D. Proc. Natl. Acad. Sci. USA 98, 3471–3476 (2001).
Bradshaw, N., Neher, S.B., Booth, D.S. & Walter, P. Science 323, 127–130 (2009).
Halic, M. et al. Nature 427, 808–814 (2004).
Schaffitzel, C. et al. Nature 444, 503–506 (2006).
Janda, C.Y. et al. Nature 465, 507–510 (2010).
Hainzl, T., Huang, S. & Sauer-Eriksson, A.E. Nature 417, 767–771 (2002).
Hainzl, T., Huang, S. & Sauer-Eriksson, A.E. Proc. Natl. Acad. Sci. USA 104, 14911–14916 (2007).
Kuglstatter, A., Oubridge, C. & Nagai, K. Nat. Struct. Biol. 9, 740–744 (2002).
Jagath, J.R. et al. RNA 7, 293–301 (2001).
Zhang, X., Kung, S. & Shan, S.O. J. Mol. Biol. 381, 581–593 (2008).
Acknowledgements
We thank the European Synchrotron Radiation Facility (Grenoble, France) for provision of synchrotron radiation facilities, and T. Bergfors and U. Sauer for critical reading of the manuscript. This work was supported by the Swedish Research Council (A.E.S.-E.) and Kempe Foundation (A.E.S.-E.).
Author information
Authors and Affiliations
Contributions
T.H., S.H. and A.E.S.-E. contributed to every aspect of this work. G.M. contributed to cloning and protein production and K.B. to Biacore experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Table 1 and Supplementary Methods (PDF 793 kb)
Rights and permissions
About this article
Cite this article
Hainzl, T., Huang, S., Meriläinen, G. et al. Structural basis of signal-sequence recognition by the signal recognition particle. Nat Struct Mol Biol 18, 389–391 (2011). https://doi.org/10.1038/nsmb.1994
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1994
This article is cited by
-
Hydrophobicity, rather than secondary structure, is essential for the SRP dependent targeting of GPR35 to the ER membrane
Journal of Bioenergetics and Biomembranes (2019)
-
Cotranslational protein targeting to the membrane: Nascent-chain transfer in a quaternary complex formed at the translocon
Scientific Reports (2018)
-
“Multiple partial recognitions in dynamic equilibrium” in the binding sites of proteins form the molecular basis of promiscuous recognition of structurally diverse ligands
Biophysical Reviews (2018)
-
Protein export through the bacterial Sec pathway
Nature Reviews Microbiology (2017)
-
The signal recognition particle contacts uL23 and scans substrate translation inside the ribosomal tunnel
Nature Microbiology (2017)