Abstract
The structure and diversity at the 3′ ends of mRNA transcripts have been extensively characterized using several tag-based techniques in eukaryotes. However, the 5′ ends of mRNA transcripts are not well understood, owing to a lack of efficient experimental approaches. We developed a new gene expression profiling method, called robust analysis of 5′-transcript ends (5′ RATE), to rapidly isolate the 5′ ends of mRNA transcripts. After ligating RNA oligo linkers to the 5′ regions of decapped mRNA, cDNA is synthesized and digested with the restriction enzyme NlaIII. Ditags are formed by ligating two individual NlaIII tags, and are then PCR-amplified, purified and sequenced using a pyrosequencing approach. The 5′-RATE procedure is simple, fast and cost-effective because the complicated steps in comparative methods such as serial analysis of gene expression (including the formation of concatemers and their subsequent cloning and sequencing) have been eliminated. The longer 5′-RATE tags (>80 bp) provide more accurate matching to reference sequences for gene annotation and allow in-depth analysis of sequence diversity at the 5′ regions of mRNA transcripts. Using our procedure, a 5′-RATE library with about 180,000 end sequences can be generated within a week. We have successfully applied the 5′-RATE method to characterize the transcriptome of various plant species including maize, rice and soybean. This method can be easily adapted to other eukaryotic organisms using the detailed procedures described in this protocol.
Similar content being viewed by others
Introduction
Expression profiling is a powerful genomic approach for identifying novel genes and cataloging expressed genes in a target tissue. Since 1990, various gene expression profiling methods have been developed to characterize mRNA transcripts in eukaryotic organisms. The expressed sequence tag (EST)1 method is the first method used for expression analysis and gene discovery in many eukaryotic organisms. For example, over eight million ESTs have been generated from different human tissues (http://www.ncbi.nlm.nih.gov/dbEST/dbEST_summary.html). The major limitation of this approach is its low rate of novel or rare transcript discovery (<2%) (see refs. 2,3). To obtain the complete transcription units, a full-length (FL) cDNA cloning method4,5 has been developed and applied in several model eukaryotic organisms. Both EST and FL-cDNA approaches are mostly suitable for profiling highly expressed transcripts (see Table 1) and require sequencing of thousands of cDNA clones to recover rare or low abundantly expressed transcripts. Since over 60% of the transcripts in eukaryotic cells are expressed at a low level2,6, high sequencing costs prohibit sampling of several thousands of cDNA clones in most of the EST and FL-cDNA projects. In addition, the EST approach has mostly characterized the transcripts at the 3′ region (see Table 1). Therefore, it is difficult to predict the transcription start sites (TSSs) and promoters when 3′-EST sequences are used for gene annotation. The extensive characterization of 5′ and 3′ regions of over 100,000 FL-cDNAs from mouse revealed a complexity of transcript architectural variation due to alternative promoter usage, splicing and polyadenylation7. The detailed analysis of mammalian FL-cDNAs revealed an average of three promoters per gene based on TSS analysis, suggesting that the 5′ region of transcripts is structurally diverse and complex in nature8.
In parallel, other tag-based methods such as serial analysis of gene expression (SAGE) and massively parallel signature sequencing (MPSS) have been developed to profile transcriptomes in greater depth (see Table 1 for a comparison of tag-based methods of transcriptome analysis). The SAGE method was first developed in 1995 in humans9. So far, about 1.3 million distinct SAGE tags out of 19.3 million total tags from 327 libraries have been generated for the human transcriptome (http://www.ncbi.nlm.nih.gov/projects/SAGE). The conventional SAGE methodology encounters many technical problems such as short inserts of concatamers and low cloning efficiency. To overcome these problems, we previously developed an improved LongSAGE protocol called robust-SAGE (RL-SAGE)10,11. Using this method, we have generated over a million RL-SAGE tags from plants (rice and maize) and fungus (Magnaporthe grisea)12,13. These modifications have also been successfully adopted in a human transcriptome analysis project in which over 30 million tags are sequenced from over 250 tissues14. Recently, the SAGE method was also modified to recover transcript information at 5′ ends of mRNA15,16,17. MPSS was also developed for in-depth transcriptome analysis using a novel hybridization-based sequencing method17. MPSS tags are 17–21 bp in length, and over a million tags per library can be generated. MPSS library construction is, however, complex and only performed by experienced technicians at Illumina Inc. (formerly Solexa and Lynxgen). Both SAGE and MPSS methods can characterize only a short signature (15–26 bp) from each mRNA, which may be problematic for in-depth analysis of a target genome. For example, some transcripts could be missed because of the lack of an anchoring restriction site during tag isolation18. Second, about 5–15% of tag sequences cannot be mapped to the gene level owing to multiple hits in the target genes19. Finally, short tags may not be appropriate to recover splice forms and study sequence diversity across the transcription units. Therefore, additional cDNA cloning strategies are required to confirm the short tag expression3.
Conventional Sanger DNA sequencing has been widely used in gene expression profiling and genome sequencing projects. For large-scale transcriptome sequencing, however, the Sanger sequencing method is too time consuming and expensive following the cloning of cDNA fragments and selection and purification of clones. Recently, 454 Life Sciences (http://www.454.com) has adopted pyrosequencing technology to develop a scalable DNA sequencing method that eliminates both individual colony picking and purification20. The pyrosequencing method is 100 times faster than the Sanger sequencing method and is capable of sequencing more than 200,000 DNA fragments within 5 h20. Initially, the sequence read size was about 100 bp using the Genome Sequencer 20 but now the improved version called Genome Sequencer FLX System is capable of sequencing DNA reads from 200 to 300 bp.
We have successfully incorporated pyrosequencing into our newly developed method called 5′ RATE21. This method is being used for investigating the transcript diversity at the 5′ region and identifying putative TSSs and promoter regions of expressed genes in maize, soybean and rice in our laboratory. The following detailed protocol can also be easily applied to analyze the 5′ regions of transcripts in other eukaryotic organisms. In most eukaryotes, a substantial amount of information is available for the 3′ region of transcripts in EST databases. Identification of the 5′ region using approaches like 5′-RATE method would significantly enhance our understanding of the structure and diversity of the expressed genes. It is noteworthy to mention that the 5′-RATE method can also be easily modified for the characterization of the 3′ region of transcripts: after removing the 3′-polyA tail of mRNAs, the oligo linkers can be ligated to the 3′ region of the treated mRNAs and the subsequent 5′-RATE procedures followed to make a 3′-RATE library.
Experimental design
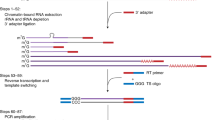
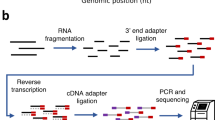
The entire 5′-RATE procedure is shown in Figure 1. This method involves 5′-oligo capping of mRNAs4, isolating 5′-NlaIII tags and ditag formation19, and sequencing of 5′-NlaIII ditags using the pyrosequencing method19. In general, mRNA populations consist of mature (5′-G-capped and 3′-polyA tailed) and premature (5′-phosphate and 3′-polyA tail; 5′-G-capped and 3′ OH; and/or 5′ phosphate and 3′ OH) RNAs. Using the Oligotex direct mRNA purification kit (Qiagen; see Steps 17 and 18), the 3′-polyadenylated mRNAs are first purified from total RNA extracted from a tissue of interest. The 5′ phosphates from premature mRNAs are then removed using phosphatase enzyme and the 5′-G-caps from mature mRNAs are removed using acid pyrophosphatase enzyme. These treated mRNAs are divided into two pools to allow subsequent ditag formation. RNA oligo linkers 1 and 2, for the two pools respectively, are ligated to the 5′ ends of mature mRNAs only (those that have 5′ phosphates following the decapping procedure) using T4 RNA ligase; T4 RNA ligase specifically joins the RNA strands when the donor molecule contains a 5′-phosphate group (PO4) and the acceptor molecule contains a 3′-hydroxyl group (OH). Single-stranded cDNA is synthesized from each pool using a random adapter primer. The double-stranded cDNA is PCR-amplified using a biotinylated primer complementary to the 5′-RNA oligo linker and a non-biotinylated primer specific to the 3′-random adapter primer sequence. The RNA linkers, random adapter primers and PCR primers are designed as described by Hashimoto et al.16 and the same sequences can be used for all 5′-RATE assays. Using streptavidin magnetic beads, PCR amplicons are captured and then digested with NlaIII, a type II restriction enzyme that recognizes CATG and cuts DNA at every 250 bp. This enzyme has been widely used as a tagging enzyme to generate tags in many SAGE libraries9,10. After washing of the magnetic beads, the 5′-NlaIII tags from two pools are mixed together and ligated to generate ditags using T4 DNA ligase. This ditag-based PCR amplification strategy is used because it provides a more faithful representation of the tags' expression frequencies in comparison to the use of individual or mono tags, which was previously tested in SAGE methodology9,10. The ligated NlaIII ditags are PCR-amplified using primers specific to the 5′-RNA oligo linkers. Finally, linkers are removed from the PCR amplicons by digesting with XhoI. Ditags are purified from an agarose gel and are blunt-ended using Lambda exonuclease. Ditags are then ligated with 454 sequencing primers (double-stranded oligonucleotides comprising 20 bases of PCR amplification primer and 20 bases of sequencing primer, previously described in ref. 20). These 454 sequencing primers can be used for any 5′-RATE library construction. The ligation product is purified and subjected to 454 pyrosequencing20. A raw NlaIII ditag sequence contains a forward and a reverse tag. Finally, the two individual 5′-end tag sequences (the forward and reverse NlaIII tags) are isolated from ditag sequence reads and matched to genomic DNA and ESTs/FL-cDNA sequences. Unlike other cDNA cloning approaches such as EST and SAGE, 5′ RATE eliminates insert cloning, colony picking and plasmid purification before sequencing. Since the ligation and transformation of DNA into bacteria are omitted, the biased amplification of some transcripts in the cDNA population may be reduced. The 5′-RATE method may, however, miss the transcripts without an NlaIII site. This can be overcome by constructing another 5′-RATE library using other restriction enzymes such as DpnII, TaqI, MseI or Sau3AI.
The mRNA population consists of mature (5′-G-capped (represented by Gppp) and 3′-polyA tailed (represented by AA(A)n) and premature (5′-phosphate (represented by p) and 3′-polyA tail; 5′-G-capped and 3′-OH; and 5′-phosphate and 3′-OH) RNAs. First, polyadenylated mRNA molecules are isolated from total RNA for use in 5′-RATE library construction. The 5′ phosphates from premature mRNAs are removed using phosphatase enzyme and the 5′-G-caps from mature mRNAs are removed using acid pyrophosphatase enzyme. Then mRNAs treated with acid pyrophosphatase are divided into two pools to allow subsequent ditag formation. The 5′ ends of mature mRNAs only are ligated with RNA oligo linker 1 and RNA oligo linker 2, for the two pools, respectively, using T4 RNA ligase, as this enzyme specifically joins RNA strands when the donor molecule contains a 5′-phosphate group (PO4) and the acceptor molecule contains a 3′-hydroxyl group (OH). Single-stranded cDNA is synthesized from each pool using a random adapter primer. The double-stranded cDNA is PCR-amplified using a biotinylated primer complementary to the 5′-RNA oligo linker and a non-biotinylated primer specific to the 3′-random adapter primer sequence. Using streptavidin magnetic beads, PCR amplicons are captured and are then digested with NlaIII enzyme. After washing the beads, the 5′-NlaIII tags from two pools are mixed together and ligated to generate ditags using T4 DNA ligase. The ligated ditags are PCR-amplified using primers specific to the 5′ linkers. Finally, the linkers are removed from the PCR amplicons by digesting with XhoI. Ditags are purified from an agarose gel and are blunt-ended using lambda exonuclease. Ditags are ligated with 454 sequencing primers and sequenced using pyrosequencing method. Finally, the individual tags are isolated from ditag sequence reads and matched to genomic DNA and ESTs/FL-cDNAs.
Materials
Reagents
-
RNasin (Promega, cat. no. N2111)
-
Tobacco acid pyrophosphatase (Epicentre, cat. no. T19500)
-
T4 RNA ligase (TaKaRa, cat. no. TAK2050A)
-
Bacterial alkaline phosphatase (TaKaRa, cat. no. TAK2120B)
-
DNase I (Invitrogen, cat. no. 18068-015)
-
High-fidelity platinum Taq DNA polymerase (Invitrogen, cat. no. 11304-011)
Critical
The cDNA amplification can be performed using Pfu Turbo DNA polymerase (Stratagene, cat. no. 600250-52), but the PCR amplification was less intense than high-fidelity Taq DNA polymerase (Invitrogen, cat. no. 11304-011).
-
NlaIII (NEB, cat. no. R0125S)
-
Xho1 (NEB, cat. no. R0146S)
-
100 bp ladder (Invitrogen, cat. no. 15628-019)
-
Phenol:chloroform:isoamyl alcohol (Invitrogen, cat. no. 15593-031)
-
Ribonuclease H (Invitrogen, cat. no. 18021-014)
-
Diethyl pyrocarbonate (DEPC; Sigma-Aldrich, cat. no. D5758)
Caution
DEPC is toxic: avoid contacting with body parts.
-
Glycogen (Ambion, cat. no. 9510)
-
Polyethylene glycol 8000 (Sigma-Aldrich, cat. no. 81268)
-
Reverse transcriptase kit (Promega, cat. no. A3500)
-
M-280 streptavidin beads slurry (Dynal, cat. no. 18090-019)
-
Trizol (Invitrogen, cat. no. 18068-015)
-
Chloroform (Sigma, cat. no. C2432)
-
NaOH (Sigma, cat. no. 71689)
Caution
NaOH is toxic: avoid contacting with body parts.
-
Tris-HCl (Sigma, cat. no. T5941)
-
EDTA (Sigma, cat. no. E5134)
-
NaCl (Sigma, cat. no. S7653)
-
SDS (Sigma, cat. no. L4390)
Caution
SDS is toxic: avoid contacting with body parts.
-
BSA (Sigma, cat. no. A9647)
-
LiCl (Sigma, cat. no. L9650)
-
Oligotex direct mRNA Midi/Maxi Kit (Qiagen, cat. no. 72041)
-
QIAquick Gel Extraction Kit (Qiagen, cat. no. 28704)
-
All the following oligonucleotides were obtained from Integrated DNA Technology
-
5′-RNA oligo linker 1: 5′-UUU GGA UUU GCU GGU GCA GUA CAA CUA GGC UUA AUA CUC GAG UCC GAC G-3′
-
5′-RNA oligo linker 2: 5′-UUU CUG CUC GAA UUC AAG CUU CUA ACG AUG UAC GCU CGA GUC CGA CG-3′
-
Random adapter primer: 5′-GCG GCT GAA GAC GGC CTA TGT GGC CNN NNC-3′
-
5′ primer PCR 1: 5′Bio/GGA TTT GCT GGT GCA GTA CAA CTA GGC-3′
-
5′ primer PCR 2: 5′Bio/CTG CTC GAA TTC AAG CTT CTA ACG ATG-3′
-
3′-PCR primer: 5′-GCG GCT GAA GAC GGC CTA TGT-3′
-
454 adaptor A: 5′-CCA TCT CAT CCC TGC GTG TCC CAT CTG TTC CCT CCC TGT CTC AG-3′
-
454 adaptor B: 5′BioTEG/CCT ATC CCC TGT GTG CCT TGC CTA TCC CCT GTT GCG TGT CTC AG-3′
Equipment
-
Water bath
-
Magnetic stand
-
Spectrophotometer or NanoDrop
-
Freezers (−80, −20 °C)
-
4 °C centrifuge
-
Agarose gel electrophoresis unit
Reagent setup
-
0.2 N NaOH
Critical
Should be freshly prepared.
-
1% (v/v) DEPC H2O solution Stir at 37 °C for 12 h.
Critical
Should be freshly prepared.
-
Wash buffer A 10 mM Tris, pH 7.5, 0.5 mM EDTA, 150 mM LiCl and 10 μg ml−1 glycogen.
Critical
Should be freshly prepared.
-
Wash buffer B 5 mM Tris, pH 7.5, 0.5 mM EDTA, 1 M NaCl, 1% SDS (wt/vol) and 10 μg ml−1 glycogen.
Critical
Should be freshly prepared.
-
Wash buffer C 5 mM Tris, pH 7.5, 0.5 mM EDTA, 1 M NaCl and 10 μg ml−1 BSA.
Critical
Should be freshly prepared.
Procedure
Creating an RNase-free environment
Timing 1–2 days
-
1
Wipe work bench with ethanol and RNase-free DEPC H2O.
-
2
Soak tubes, pestle and mortar with 0.2 N NaOH for 30–60 min.
-
3
Soak tubes, pestle and mortar in 1% DEPC solution at 37 °C for 12 h.
-
4
Autoclave tubes and pestle and mortar for 60 min.
Critical Step
Work in a clean and aerosol-free space at all times to avoid RNase contamination.
Isolation of total RNA
Timing 3–5 h
-
5
Grind 1–2 g of tissue of interest in a pestle and mortar using liquid nitrogen and transfer to 50 ml tube.
Caution
Liquid nitrogen is hazardous; avoid contacting body parts.
Critical Step
Minimize the tissue exposure to room temperature. RNA is susceptible to degradation at room temperature.
-
6
Add 20 ml of Trizol solution to the tube immediately.
-
7
Incubate at room temperature (27 °C) for 10 min.
-
8
Add 5 ml of chloroform and incubate at room temperature for 5 min and then centrifuge for 20 min at 4 °C at 9,000g.
Caution
Trizol and chloroform are toxic chemicals; avoid inhaling or body contact.
-
9
Transfer the supernatant (containing RNA) into 15 ml of ice-cold isopropanol, mix well, incubate on ice for 10 min and centrifuge at 9,000g for 15 min at 4 °C.
Critical Step
Look for RNA pellet (white) at the bottom of the tube.
-
10
Remove the supernatant carefully without disturbing the RNA pellet.
Critical Step
Look for the RNA pellet while removing isopropanol.
-
11
Add 20 ml of 70% ethanol (70:30, absolute ethanol:DEPC H2O) and rotate tubes slowly.
Critical Step
Avoid disturbing the RNA pellet.
-
12
Centrifuge at 9,000g for 15 min at 4 °C and remove the supernatant carefully.
Critical Step
Avoid disturbing the RNA pellet.
-
13
Dry RNA in the laminar flow at room temperature for 10 min.
-
14
Dissolve total RNA in 500 μl of RNase-free DEPC H2O.
-
15
Estimate RNA concentration in 1 μl using a spectrophotometer or NanoDrop.
-
16
Confirm RNA integrity by electrophoresis on a 1.2% (w/v) agarose gel with 1 μl of RNA sample according to Sambrook et al.22.
Critical Step
Typically 0.5–1 mg of total RNA can be expected.
Isolation of mRNA
Timing 5–10 h
-
17
Take 1.0 mg of total RNA (from Step 14) in RNase-free 1.5 ml tube and adjust the volume to 500 μl with RNase-free DEPC H2O.
-
18
Follow the mRNA purification procedure using the Oligotex direct mRNA Midi/Maxi Kit.
-
19
After mRNA purification, make up RNA solution to 300 μl using DEPC H2O.
-
20
Add 133 μl of 5 M ammonium acetate, 4 μl of glycogen and 1 ml of absolute ethanol. Mix well and incubate at −80 °C for 3 h to overnight.
Pause point
mRNA can be stored at −80 °C for several days.
-
21
Centrifuge mRNA solution at 9,000g for 30 min at 4 °C and carefully remove the supernatant.
Critical Step
Look for mRNA pellet while removing isopropanol.
-
22
Add 1 ml of 70% ethanol (70:30, absolute ethanol:DEPC H2O) and rotate tubes slowly.
Critical Step
Avoid disturbing the mRNA pellet.
-
23
Centrifuge at 9,000g for 15 min at 4 °C and remove the supernatant carefully.
Critical Step
Avoid disturbing the mRNA pellet.
-
24
Dry mRNA in the laminar flow at room temperature for 10 min.
-
25
Dissolve mRNA in 50 μl of RNase-free DEPC H2O.
-
26
Estimate mRNA concentration using a spectrophotometer or a NanoDrop.
-
27
Confirm mRNA integrity on a 1.5% (w/v) agarose gel according to Sambrook et al.22.
Critical Step
Typically 0.5–1 μg of mRNA can be obtained.
Dephosphorylation of mRNAs
Timing 3–5 h
-
28
Add the following reagents to an RNase-free tube (total volume 100 μl), 50 μl of mRNA (500 ng), 10 μl of 10 × buffer, 3 μl of RNasin (40 U μl−1), 5 μl of bacterial alkaline phosphatase (150 U μl−1) and 32 μl of DEPC H2O.
-
29
Incubate the dephosphorylation reaction at 50 °C for 60 min.
-
30
After the incubation period, add 200 μl of RNase-free DEPC H2O and 300 μl of phenol:chloroform:isoamyl alcohol mixture, gently mix the contents and centrifuge at 9,000g for 10 min at 4 °C.
-
31
Transfer the aqueous layer to another RNase-free tube.
-
32
Add 133 μl of 5 M ammonium acetate, 4 μl of glycogen and 1 ml of 100% ethanol. Incubate at −80 °C for 3 h to overnight.
Pause point
mRNA can be stored at −80 °C for several days.
-
33
Centrifuge the tubes at 9,000g for 30 min at 4 °C and remove the supernatant.
-
34
Add 1 ml of 70% ethanol, mix gently and centrifuge at 9,000g for 10 min at 4 °C.
-
35
Remove the supernatant and dry mRNA pellet in a laminar flow for 10 min.
-
36
Dissolve mRNA in 50 μl of RNase-free DEPC H2O.
Decapping of 5′ regions of mRNA
Timing 3–5 h
-
37
Add the following reagents to an RNase-free tube (total volume 100 μl): 50 μl of dephosphorylated mRNA (from Step 36), 10 μl of 10 × buffer, 3 μl of RNasin (40 U μl−1), 5 μl of tobacco acid pyrophosphatase (10 U μl−1) and 32 μl of RNase-free DEPC H2O.
-
38
Incubate the above contents at 37 °C for 2 h.
-
39
After the incubation period, add 200 μl of DEPC H2O and treat twice with 300 μl of phenol:chloroform:isoamyl alcohol mixture by following Steps 20–25 two times.
-
40
Dissolve mRNA in 5 μl of RNase-free DEPC H2O.
-
41
Divide mRNA equally into two tubes for ligation with two different 5′ RNA oligos (linkers).
Ligation of 5′ mRNAs to RNA oligo linker
Timing 3–5 h
-
42
Add the following reagents to RNase-free tubes (total volume 50 μl).
Pool 1: add 5 μl of decapped mRNA (from Step 41), 5 μl of 10 × T4 RNA ligase buffer, 5 μl of 5′-RNA oligo linker 1 (100 ng μl−1), 3.5 μl of RNasin (40 U μl−1), 5 μl of T4 RNA ligase (40 U μl−1), 3 μl of 0.1% BSA and 23.5 μl of polyethylene glycol 8000 (50%, w/v).
Pool 2: this is same as pool 1 but use 5′-RNA oligo linker 2 (100 ng μl−1).
-
43
Mix the reactions well and incubate at 16 °C overnight.
-
44
To each pool, add 250 μl of RNase-free DEPC H2O (to make up to 300 μl), and treat twice with 300 μl of phenol:chloroform:isoamyl alcohol mixture by following Steps 20–25 two times.
-
45
Dissolve mRNA in 50 μl of RNase-free DEPC H2O.
Digestion of genomic DNA using DNase I
Timing 3–5 h
-
46
To remove residual DNA contamination in mRNA, add the following reagents to each pool: 50 μl of oligo-capped mRNA (from Step 45), 10 μl of 10 × buffer, 3 μl of RNasin (40 U μl−1), 3 μl of DNase I (1 U μl−1) and 34 μl of RNase-free DEPC H2O.
-
47
Incubate reactions at 37 °C for 30 min, then add 50 μl of 0.5 M EDTA and heat-inactivate DNase I at 65 °C for 10 min.
-
48
To each pool, add 200 μl of DEPC H2O and treat twice with 300 μl of phenol:chloroform:isoamyl alcohol mixture by following Steps 20–25 two times.
-
49
Dissolve mRNA in 500 μl of RNase-free DEPC H2O.
Purification of mRNAs
Timing 3–5 h
-
50
Repeat Steps 17–24 for each pool.
-
51
Dissolve mRNA in 10 μl of RNase-free DEPC H2O.
Synthesis of single-strand cDNA
Timing 3–5 h
-
52
Add the following reagents to each mRNA pool (100 μl volume): 10 μl of mRNA (from Step 51), 10 μl of dNTPs (10 mM), 5 μl of random adapter primer (100 ng μl−1), 10 μl of 10 × reverse transcriptase buffer, 3 μl of RNasin, 5 μl of MgCl2, 5 μl of reverse transcriptase and 52 μl of RNase-free DEPC H2O.
-
53
Incubate the reactions at 12 °C for 1 h and then incubate at 42 °C for 4 h.
Digestion of mRNAs using RNase H
Timing 3–5 h
-
54
Increase the volume of each pool to 150 μl by adding 15 μl of 10 × buffer and 5 μl of Escherichia coli RNase H (2 U μl−1) and DNase-free water.
-
55
Incubate the reactions for 1 h at 37 °C.
-
56
To each pool, add 150 μl of DNase-free H2O and treat twice with 300 μl of phenol:chloroform:isoamyl alcohol mixture by following Steps 20–25 two times.
-
57
Dissolve cDNA in 100 μl of DNase-free H2O.
Synthesis of double-strand cDNA using PCR
Timing 3–5 h
-
58
Perform PCRs for pool 1 using the following reagents: 15.0 μl of pool 1 single-stranded cDNA (from Step 57), 2 μl of dNTPs (5 mM), 5 μl of biotinylated 5′ primer PCR 1 (10 pmol μl−1), 5 μl of 3′-specific anchoring primer (10 pmol μl−1), 5 μl of 10 × PCR buffer, 2 μl of MgSO4 (50 mM), 0.5 μl of high-fidelity platinum Taq DNA polymerase (5 U μl−1) and 15.5 μl of DNase-free H2O.
-
59
Set up another PCR as described above by using biotinylated 5′-primer PCR 2 for pool 2 single-stranded DNA (from Step 57). The rest of the steps (below) are the same for the two pools.
-
60
Perform PCR at 94 °C, 5 min followed by 15 PCR cycles at 94 °C for 1 min, 55 °C for 1 min and 72 °C for 2 min and finally extension cycle at 72 °C for 15 min.
-
61
Increase the volume to 300 μl by adding DNase-free H2O and treat twice with 300 μl of phenol:chloroform:isoamyl alcohol mixture by following Steps 20–25 two times.
-
62
Dissolve the DNA in 25 μl of DNase-free H2O.
Generation of 5′ tags from cDNA
Timing 3–5 h
-
63
Mix the following reagents for each pool (100 μl): 25 μl of cDNA (from Step 62), 10 μl of 10 × NlaIII buffer, 1.5 μl of 100 × BSA, 5 μl of NlaIII (10 U μl−1) and 58.5 μl of DNase-free H2O.
-
64
Mix well and incubate at 37 °C for 3 h.
-
65
Increase the volume to 300 μl by adding DNase-free H2O and treat twice with 300 μl of phenol:chloroform:isoamyl alcohol mixture by following Steps 20–25 two times.
-
66
Dissolve the DNA in 25 μl of DNase-free H2O.
Capture of 5′ tags on streptavidin beads
Timing 3–5 h
-
67
Aliquot 200 μl of streptavidin magnetic beads into a 1.5 ml siliconized tube (non-sticky tubes) for each cDNA pool.
-
68
Keep streptavidin magnetic bead-containing tubes on the magnetic stand for 2 min.
-
69
Remove the supernatant and discard the solution.
-
70
Wash the beads three times. For each wash, add 300 μl of wash buffer A to streptavidin magnetic beads, mix well by vortexing, place the tube on the magnetic stand for 2 min and remove the supernatant.
-
71
Add NlaIII-digested cDNA (from Step 66) to the streptavidin magnetic beads and mix well for 30 min at room temperature on a shaker at 100 r.p.m.
-
72
Wash streptavidin magnetic beads twice with prewarmed (50 °C) 300 μl of wash buffer B, as described in Step 70.
-
73
Wash streptavidin magnetic beads four times with 300 μl of wash buffer C, as described in Step 70.
-
74
Wash streptavidin magnetic beads three times with 200 μl of 1 × ligase buffer, as described in Step 70.
-
75
Finally, combine cDNA from pools 1 and 2.
Ditag formation by ligating 5′ tags
Timing 3–5 h
-
76
Add 2.5 μl of 10 × ligase buffer, 2.5 μl of T4 ligase (5 U μl−1) and 20 μl of DNase-free H2O to the above mixture of ditag cDNA pools 1 and 2 (from Step 75).
-
77
Incubate at 16 °C overnight.
Critical Step
Streptavidin magnetic beads settle at the bottom of the tube. Mix beads by flicking tubes at every 20 min for initial 5 h.
PCR amplification of 5′ ditags
Timing 3–5 h
-
78
Dilute the ditags cDNA (ligated product) from Step 77 (1:100 or 1:50) with DNase-free H2O.
Critical Step
In general, 1:100 to 1:50 (ditag:H2O) dilutions give proper results. If faint amplification or overamplification occurs, optimization of ditag cDNA dilution is required depending on the DNA smear intensity on an agarose gel.
-
79
Set up bulk PCRs in 50 μl volume. For each PCR, add 1 μl of diluted ditag cDNA (from Step 78), 2 μl of dNTPs (5 mM), 5 μl of biotinylated 5′-primer PCR 1 (10 pmol μl−1), 5 μl of biotinylated 5′-primer PCR 2 (10 pmol μl−1), 5 μl of 10 × PCR buffer, 2 μl of MgSO4 (50 mM), 0.5 μl of platinum Taq DNA polymerase high fidelity (5 U μl−1) and 29.5 μl of DNase-free H2O.
Critical Step
Generally ten PCRs are enough to get required amount of template DNA for 454 sequencing. If needed, up to 50 PCRs can be followed.
-
80
Perform PCR at 94 °C, 5 min followed by 27 PCR cycles at 94 °C for 1 min, 58 °C for 1 min and 72 °C for 2 min and finally extension cycle at 72 °C for 15 min.
-
81
Confirm PCR products by electrophoresis on a 1.5% (w/v) agarose gel at 120 V for 30 min using 0.5 × TBE buffer22.
Critical Step
DNA smear should be seen from 100 bp to 3 kb (more DNA smear around 250–500 bp region).
-
82
Increase volume to 300 μl by adding DNase-free H2O and treat twice with 300 μl of phenol:chloroform:isoamyl alcohol mixture by following Steps 20–25 two times.
-
83
Dissolve the DNA in 10 μl of DNase-free H2O.
Gel purification of 5′-ditags
Timing 3–5 h
-
84
Prepare a 3.5% (w/v) agarose gel using 0.5 × TBE buffer22 and load entire PCR.
-
85
Perform electrophoresis at 120 V for 60 min.
-
86
Visualize NlaIII ditag DNA under UV lamp and excise bands from 100 bp to 3 kb.
Critical Step
Minimize exposure of DNA to UV.
Caution
Direct viewing of UV rays can damage eyes.
Caution
Ethidium bromide is highly carcinogenic. Always handle gel by wearing gloves and avoid contact with skin.
-
87
Purify cDNA bands from 100 bp to 3 kb using the Qiagen gel purification kit according to the manufacturer's instructions.
Critical Step
DNA-containing gel cannot be incubated at 50 °C for more than 5 min in QG buffer. QG buffer is a potent denaturing agent; therefore, follow the manufacturers' instructions strictly.
-
88
Increase volume to 300 μl by adding DNase-free H2O and treat twice with 300 μl of phenol:chloroform:isoamyl alcohol mixture by following Steps 20–25 two times.
-
89
Dissolve the DNA in 25 μl of DNase-free H2O.
Removal of linkers from 5′ ditags by digesting with XhoI
Timing 3–5 h
-
90
Prepare the following reaction mixture (100 μl) by adding 25 μl of ditag DNA (from Step 89), 10 μl of 10 × XhoI buffer, 1.5 μl of 100 × BSA, 5 μl of XhoI (10 U μl−1) and 59.5 μl of DNase-free H2O.
-
91
Mix well and incubate the reaction at 37 °C for 3 h.
-
92
Increase volume to 300 μl by adding DNase-free H2O and treat twice with 300 μl of phenol:chloroform:isoamyl alcohol mixture by following Steps 20–25 two times.
-
93
Dissolve DNA in 10 μl of H2O.
Gel purification of XhoI-digested 5′ ditags
Timing 3–5 h
-
94
Repeat Steps 84–88.
-
95
Dissolve the DNA in 100 μl of DNase-free H2O.
Removal of linkers and undigested ditag DNA
Timing 3–5 h
-
96
Repeat Steps 67–70.
-
97
Add XhoI-digested 5′-ditag cDNA (from Step 95) to streptavidin magnetic beads (from Step 96) and shake at 100 r.p.m. for 30 min at room temperature.
-
98
Collect the supernatant (cDNAs) into a separate tube.
Critical Step
Ditag DNA is in the supernatant, handle with care and do not discard the supernatant.
-
99
Increase volume to 300 μl by adding DNase-free H2O and treat twice with 300 μl of phenol:chloroform:isoamyl alcohol mixture by following Steps 20–25 two times.
-
100
Dissolve cDNA in 10 μl of DNase-free H2O and estimate DNA concentration.
Pyrosequencing of 5′ ditags, tags isolation and sequence analysis
Timing 1 week
-
101
The 5′-ditag fragments (from Step 100) are polished using T4 DNA polymerase in a 50 μl volume as described in ref. 20. The detailed 454 protocol is available at http://www.nature.com/nature/journal/v437/n7057/extref/nature03959-s3.doc.
-
102
Ligate 454 adapters to 5 μg of blunt-ended 5′-RATE ditag DNA (from Step 101) in a 40 μl volume as described in ref. 20.
-
103
Perform pyrosequencing reaction according to the protocol described in ref. 20, using a 454 Life Sciences Genome Sequencer at your research facility or send the DNA samples to 454 Life Sciences (http://www.454.com) for custom sequencing.
Critical Step
Typically, about 150,000–200,000 sequence reads (21–150 bp) can be obtained.
RATE tag extraction
Timing 1–2 days
-
104
Extract the forward tags from the pyrosequencing data obtained as follows: find sequences with forward signature (TCGAGT), trim off the forward signature at the beginning and find the end marker CATG (NlaIII site). If a CATG signature does not exist in the ditag, isolate the tag sequence until 100 bp.
-
105
Extract the reverse tags by reverse complementation of raw sequence reads (from Step 103) and repeat Step 104 to get tags in reverse direction.
-
106
Combine tags from Steps 104 and 105, and then tags are clustered using a software program to obtain distinct tags per library.
Critical Step
Generally, high sequence diversity exists at the 5′ region of many expressed genes. Reduce the clustering criteria as needed.
Mapping of NlaIII tags
Timing 1–2 days
-
107
Use standlone local BLAST 2.0 program to map distinct NlaIII tags to the reference databases including genomic, EST and FL-cDNA sequences.
-
108
Use these BLAST search criteria as a reference: 90% identity and an E-value of e−5.
Critical Step
Sequence variation may exist among the transcripts of expressed genes. Adjust the BLAST search criteria when mapping tags to target sequences.
Troubleshooting
Troubleshooting advice can be found in Table 2.
Timing
Steps 1–4, creating an RNase-free environment: 1–2 days
Steps 5–16, isolation of total RNA: 3–5 h
Steps 17–27, isolation of mRNA: 5–10 h
Steps 28–36, dephosphorylation of mRNAs: 3–5 h
Steps 37–41, decapping of 5′ regions of mRNAs: 3–5 h
Steps 42–45, ligation of 5′ mRNAs to RNA oligo linker: 3–5 h
Steps 46–49, digestion of genomic DNA using DNase I: 3–5 h
Steps 50 and 51, purification of mRNAs: 3–5 h
Steps 52 and 53, synthesis of single-strand cDNA: 3–5 h
Steps 54–57, digestion of mRNAs using RNase H: 3–5 h
Steps 58–62, synthesis of double-strand cDNA using PCR: 3–5 h
Steps 63–66, generation of 5′ tags from cDNA: 3–5 h
Steps 67–75, capture of 5′ tags on streptavidin beads: 3–5 h
Steps 76 and 77, ditag formation by ligating 5′ tags: 3–5 h
Steps 78–83, PCR amplification of 5′ ditags: 3–5 h
Steps 84–89, gel purification of 5′ ditags: 3–5 h
Steps 90–93, removal of linkers from 5′ ditags by digesting with XhoI: 3–5 h
Steps 94 and 95, gel purification of XhoI-digested 5′ ditags: 3–5 h
Steps 96–100, removal of linkers and undigested ditag DNA: 3–5 h
Steps 101–103, pyrosequencing of 5′ ditags, tags isolation and sequence analysis: 1 week
Steps 104–106, RATE tag extraction: 1–2 days
Steps 107 and 108, mapping of NlaIII tags: 1–2 days
Anticipated results
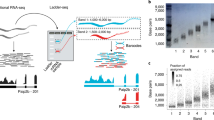
If the above steps are followed strictly, a 5′-RATE library can be made within 1 week. About 160,000–180,000 reads could be obtained from a 454 run. The number of distinct tags from each library may vary depending on the complexity of the transcripts at the 5′ region of mRNA. A substantial variation at the TSS of the transcripts of the same gene is expected, as shown in Figure 2, for the photosystem I complex PsaH subunit gene and also for other genes in maize20. This method can also possibly reveal an unusual 5′-polyA tail structure at the beginning of the transcripts, as previously reported in maize21 (Fig. 2) and soybean (data not shown) 5′-RATE libraries. The 5′-polyA tail structure has not been reported previously in any organism except for a late gene in poxvirus23. Similar 5′-polyA tail structures can be found in FL cDNAs derived from plants, animals and viruses in the NCBI GenBank databases21. Further investigation of the formation and function of 5′-polyA tails may reveal a novel mechanism of post-transcriptional processing and gene regulation in eukaryotic cells24. Information of the TSSs of the expressed genes from the 5′-RATE libraries will be helpful for gene annotation of sequenced genomes.
The 5′ region of the FL cDNA (ZM_BFb0290I10.r; see the second line of sequence alignment) and the genomic DNA (CG727954; see the first line of sequence alignment) is matched to 5′-RATE tags (subsequent lines of sequence alignment). The capital letters represent the perfectly matched bases to FL cDNA and genomic DNA, whereas lowercase letters with underlines represent the mismatched or deletion/addition of bases.
References
Adams, M.D. et al. Initial assessment of human gene diversity and expression patterns based upon 83 million nucleotides of cDNA sequence. Nature 377, 3–174 (1995).
Sun, M. et al. SAGE is far more sensitive than EST for detecting low-abundance transcripts. BMC Genomics 5, 1 (2004).
Chen, J. et al. Identifying novel transcripts and novel genes in the human genome by using novel SAGE tags. Proc. Natl. Acad. Sci. USA 99, 12257–12262 (2002).
Suzuki, Y., Yoshitomo-Nakagawa, K., Maruyama, K., Suyama, A. & Sugano, S. Construction and characterization of a full length-enriched and a 5-end-enriched cDNA library. Gene 200, 149–156 (997).
Suzuki, Y. et al. Diverse transcriptional initiation revealed by fine, large-scale mapping of mRNA start sites. EMBO Rep. 2, 388–393 (2001).
Poroyko, V. et al. The maize root transcriptome by serial analysis of gene expression. Plant Physiol. 138, 1700–1710 (2005).
Carninci, P. et al. The transcriptional landscape of the mammalian genome. Science 309, 1559–1563 (2005).
Kimura, K. et al. Diversification of transcriptional modulation: large-scale identification and characterization of putative alternative promoters of human genes. Genome Res. 16, 55–65 (2006).
Velculescu, V.E., Zhang, L., Vogelstein, B. & Kinzler, K.W. Serial analysis of gene expression. Science 270, 484–487 (1995).
Gowda, M., Jantasuriyarat, C., Dean, R.A. & Wang, G.L. Robust-LongSAGE (RL-SAGE): a substantially improved LongSAGE method for gene discovery and transcriptome analysis. Plant Physiol. 134, 890–897 (2004).
Gowda, M. & Wang, G.L. Robust-LongSAGE (RL-SAGE): an improved LongSAGE method for high-throughput transcriptome analysis. Humana Press, Totowa, NJ. Methods Mol. Biol. (in the press).
Gowda, M. et al. Deep and comparative analysis of the mycelium and appressorium transcriptomes of Magnaporthe grisea using MPSS, RL-SAGE, and oligoarray methods. BMC Genomics 7, 310 (2006).
Gowda, M. et al. Magnaporthe grisea infection triggers RNA variation and antisense transcript expression in rice. Plant Physiol. 144, 524–533 (2007).
Khattra, J. et al. Large-scale production of SAGE libraries from microdissected tissues, flow-sorted cells, and cell lines. Genome Res. 17, 108–116 (2007).
Wei, C.L. et al. 5′ Long serial analysis of gene expression (LongSAGE) and 3′ LongSAGE for transcriptome characterization and genome annotation. Proc. Natl. Acad. Sci. USA 101, 11701–11706 (2004).
Hashimoto, S. et al. 5′-End SAGE for the analysis of transcriptional start sites. Nat. Biotechnol. 22, 1146–1149 (2004).
Kodzius, R. et al. CAGE: cap analysis of gene expression. Nat. Methods 3, 211–222 (2006).
Saha, S. et al. Using the transcriptome to annotate the genome. Nat. Biotechnol. 20, 508–512 (2002).
Pleasance, E.D, Marra, M.A & Jones, S.J. Assessment of SAGE in transcript identification. Genome Res. 13, 1203–1215 (2003).
Margulies, M. et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature 437, 376–380 (2005).
Gowda, M. et al. Robust analysis of 5′-transcript ends (5′-RATE): a novel method for transcriptome profiling and genome annotation. Nucleic Acids Res. 34, e126 (2006).
Sambrook, J., Fritsch, D.F. & Maniatis, T. Molecular Cloning: A Laboratory Manual. 2nd edn. (Cold Spring Harbor Press, Cold Spring Harbor, NY, 1989).
Ahn, B.Y. & Moss, B. Capped poly(A) leader of variable lengths at the 50 ends of vaccinia virus late mRNAs. J. Virol. 63, 226–232 (1989).
Gudkov, A.T., Ozerova, M.V., Shiryaev, V.M. & Spirin, A.S. 5-Poly(A) sequence as an effective leader for translation in eukaryotic cell-free systems. Biotechnol. Bioeng. 91, 468–473 (2005).
Brenner, S. et al. Gene expression analysis by massively parallel signature sequencing (MPSS) on microbead arrays. Nat. Biotechnol. 18, 630–634 (2000).
Acknowledgements
We are grateful to Dr. Feng Chen for his generous help in sequencing 5′-RATE libraries. We are also thankful to Dr. Baba Fakrudin and Dr. Abdelaty Saleh for reading the manuscript. This work was supported by The Ohio Agricultural Research and Development Center (OARDC) Research Enhancement Grant Program and the Plant Genome Research Program of the National Science Foundation (#0321437).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Gowda, M., Li, H. & Wang, GL. Robust analysis of 5′-transcript ends: a high-throughput protocol for characterization of sequence diversity of transcription start sites. Nat Protoc 2, 1622–1632 (2007). https://doi.org/10.1038/nprot.2007.242
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2007.242
This article is cited by
-
Transcriptome analysis of the filamentous fungus Aspergillus nidulans directed to the global identification of promoters
BMC Genomics (2013)
-
Diverse and tissue-enriched small RNAs in the plant pathogenic fungus, Magnaporthe oryzae
BMC Genomics (2011)
-
Analysis of 5′ transcript heterogeneity by high-throughput sequencing of cDNA
Nature Methods (2009)
-
Analysis of 5′ transcript heterogeneity by high-throughput sequencing of cDNA
Nature Methods (2009)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.