Abstract
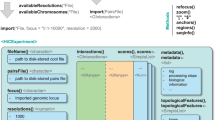
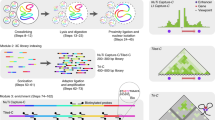
The shape of the genome is thought to play an important part in the coordination of transcription and other DNA-metabolic processes. Chromosome conformation capture (3C) technology allows us to analyze the folding of chromatin in the native cellular state at a resolution beyond that provided by current microscopy techniques. It has been used, for example, to demonstrate that regulatory DNA elements communicate with distant target genes through direct physical interactions that loop out the intervening chromatin fiber. Here we discuss the intricacies of 3C and new 3C-based methods including the 4C, 5C and ChIP-loop assay.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Capturing chromosome conformation. Science 295, 1306–1311 (2002).
Tolhuis, B., Palstra, R.J., Splinter, E., Grosveld, F. & de Laat, W. Looping and interaction between hypersensitive sites in the active β-globin locus. Mol. Cell 10, 1453–1465 (2002).
Splinter, E., Grosveld, F. & de Laat, W. 3C technology: analyzing the spatial organization of genomic loci in vivo. Methods Enzymol. 375, 493–507 (2004).
Miele, A., Gheldof, N., Tabuchi, T.M., Dostie, J. & Dekker, J. Mapping chromatin interactions by chromosome conformation capture (3C). in Current Protocols in Molecular Biology (eds. Ausubel, F.M. et al.) 21.11.1–21.11.20 (John Wiley & Sons, Hoboken, New Jersey, USA, 2006).
Hagege, H. et al. Quantitative analysis of chromosome conformation capture assays (3C-qPCR). Nat. Protoc. 2, 1722–1733 (2007).
Dostie, J. & Dekker, J. Mapping networks of physical interactions between genomic elements using 5C technology. Nat. Protoc. 2, 988–1002 (2007).
Dekker, J. The three 'C' s of chromosome conformation capture: controls, controls, controls. Nat. Methods 3, 17–21 (2006).
Palstra, R.J. et al. The β-globin nuclear compartment in development and erythroid differentiation. Nat. Genet. 35, 190–194 (2003).
Murrell, A., Heeson, S. & Reik, W. Interaction between differentially methylated regions partitions the imprinted genes Igf2 and H19 into parent-specific chromatin loops. Nat. Genet. 36, 889–893 (2004).
Spilianakis, C.G. & Flavell, R.A. Long-range intrachromosomal interactions in the T helper type 2 cytokine locus. Nat. Immunol. 5, 1017–1027 (2004).
Liu, Z. & Garrard, W.T. Long-range interactions between three transcriptional enhancers, active Vκ gene promoters, and a 3′ boundary sequence spanning 46 kilobases. Mol. Cell. Biol. 25, 3220–3231 (2005).
O'Sullivan, J.M. et al. Gene loops juxtapose promoters and terminators in yeast. Nat. Genet. 36, 1014–1018 (2004).
Skok, J.A. et al. Reversible contraction by looping of the Tcra and Tcrb loci in rearranging thymocytes. Nat. Immunol. 8, 378–387 (2007).
Simonis, M. et al. Nuclear organization of active and inactive chromatin domains uncovered by chromosome conformation capture-on-chip (4C). Nat. Genet. 38, 1348–1354 (2006).
de Laat, W. & Grosveld, F. Inter-chromosomal gene regulation in the mammalian cell nucleus. Curr. Opin. Genet. Dev., published online 18 September 2007 (doi:10.1016/j.gde.2007.07.009).
Solomon, M.J. & Varshavsky, A. Formaldehyde-mediated DNA-protein crosslinking: a probe for in vivo chromatin structures. Proc. Natl. Acad. Sci. USA 82, 6470–6474 (1985).
Orlando, V., Strutt, H. & Paro, R. Analysis of chromatin structure by in vivo formaldehyde cross-linking. Methods 11, 205–214 (1997).
Jackson, V. Formaldehyde cross-linking for studying nucleosomal dynamics. Methods 17, 125–139 (1999).
Hogan, G.J., Lee, C.K. & Lieb, J.D. Cell cycle-specified fluctuation of nucleosome occupancy at gene promoters. PLoS Genet. 2, e158 (2006).
Giresi, P.G., Kim, J., McDaniell, R.M., Iyer, V.R. & Lieb, J.D. FAIRE (Formaldehyde-Assisted Isolation of Regulatory Elements) isolates active regulatory elements from human chromatin. Genome Res. 17, 877–885 (2007).
Zhao, Z. et al. Circular chromosome conformation capture (4C) uncovers extensive networks of epigenetically regulated intra- and interchromosomal interactions. Nat. Genet. 38, 1341–1347 (2006).
Splinter, E. et al. CTCF mediates long-range chromatin looping and local histone modification in the β-globin locus. Genes Dev. 20, 2349–2354 (2006).
Wurtele, H. & Chartrand, P. Genome-wide scanning of HoxB1-associated loci in mouse ES cells using an open-ended Chromosome Conformation Capture methodology. Chromosome Res. 14, 477–495 (2006).
Horike, S., Cai, S., Miyano, M., Cheng, J.F. & Kohwi-Shigematsu, T. Loss of silent-chromatin looping and impaired imprinting of DLX5 in Rett syndrome. Nat. Genet. 37, 31–40 (2005).
Cai, S., Lee, C.C. & Kohwi-Shigematsu, T. SATB1 packages densely looped, transcriptionally active chromatin for coordinated expression of cytokine genes. Nat. Genet. 38, 1278–1288 (2006).
Kumar, P.P. et al. Functional interaction between PML and SATB1 regulates chromatin-loop architecture and transcription of the MHC class I locus. Nat. Cell Biol. 9, 45–56 (2007).
Dostie, J. et al. Chromosome Conformation Capture Carbon Copy (5C): a massively parallel solution for mapping interactions between genomic elements. Genome Res. 16, 1299–1309 (2006).
Lomvardas, S. et al. Interchromosomal interactions and olfactory receptor choice. Cell 126, 403–413 (2006).
Branco, M.R. & Pombo, A. Intermingling of chromosome territories in interphase suggests role in translocations and transcription-dependent associations. PLoS Biol. 4, e138 (2006).
Acknowledgements
We thank F. Grosveld and the members of our laboratory for discussion. This work was supported by grants from the Dutch Scientific Organization (NWO) (016-006-026) and (912-04-082) to W.d.L.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
W.d.L. holds a patent application no. PCT/IB2006/002268 on 4C technology.
Supplementary information
Supplementary Text and Figures
Supplementary Protocol 4C technology (PDF 899 kb)
Rights and permissions
About this article
Cite this article
Simonis, M., Kooren, J. & de Laat, W. An evaluation of 3C-based methods to capture DNA interactions. Nat Methods 4, 895–901 (2007). https://doi.org/10.1038/nmeth1114
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth1114
This article is cited by
-
SOX-5 activates a novel RORγt enhancer to facilitate experimental autoimmune encephalomyelitis by promoting Th17 cell differentiation
Nature Communications (2021)
-
Epigenetic regulation of epithelial-mesenchymal transition: focusing on hypoxia and TGF-β signaling
Journal of Biomedical Science (2020)
-
3C and 3C-based techniques: the powerful tools for spatial genome organization deciphering
Molecular Cytogenetics (2018)
-
Insights about genome function from spatial organization of the genome
Human Genomics (2018)
-
Optical High Content Nanoscopy of Epigenetic Marks Decodes Phenotypic Divergence in Stem Cells
Scientific Reports (2017)