Abstract
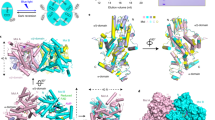
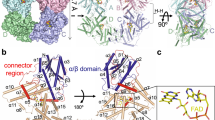
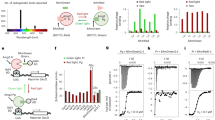
Arabidopsis thaliana cryptochrome 2 (AtCRY2), a light-sensitive photosensory protein, was previously adapted for use in controlling protein–protein interactions through light-dependent binding to a partner protein, CIB1. While the existing CRY2–CIB dimerization system has been used extensively for optogenetic applications, some limitations exist. Here, we set out to optimize function of the CRY2–CIB system by identifying versions of CRY2–CIB that are smaller, show reduced dark interaction, and maintain longer or shorter signaling states in response to a pulse of light. We describe minimal functional CRY2 and CIB1 domains maintaining light-dependent interaction and new signaling mutations affecting AtCRY2 photocycle kinetics. The latter work implicates an α13–α14 turn motif within plant CRYs whose perturbation alters signaling-state lifetime. Using a long-lived L348F photocycle mutant, we engineered a second-generation photoactivatable Cre recombinase, PA-Cre2.0, that shows five-fold improved dynamic range, allowing robust recombination following exposure to a single, brief pulse of light.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Shimizu-Sato, S., Huq, E., Tepperman, J.M. & Quail, P.H. A light-switchable gene promoter system. Nat. Biotechnol. 20, 1041–1044 (2002).
Levskaya, A., Weiner, O.D., Lim, W.A. & Voigt, C.A. Spatiotemporal control of cell signalling using a light-switchable protein interaction. Nature 461, 997–1001 (2009).
Yazawa, M., Sadaghiani, A.M., Hsueh, B. & Dolmetsch, R.E. Induction of protein-protein interactions in live cells using light. Nat. Biotechnol. 27, 941–945 (2009).
Kennedy, M.J. et al. Rapid blue-light-mediated induction of protein interactions in living cells. Nat. Methods 7, 973–975 (2010).
Strickland, D. et al. TULIPs: tunable, light-controlled interacting protein tags for cell biology. Nat. Methods 9, 379–384 (2012).
Chen, D., Gibson, E.S. & Kennedy, M.J. A light-triggered protein secretion system. J. Cell Biol. 201, 631–640 (2013).
Crefcoeur, R.P., Yin, R., Ulm, R. & Halazonetis, T.D. Ultraviolet-B-mediated induction of protein-protein interactions in mammalian cells. Nat. Commun. 4, 1779 (2013).
Müller, K. et al. Multi-chromatic control of mammalian gene expression and signaling. Nucleic Acids Res. 41, e124 (2013).
Guntas, G. et al. Engineering an improved light-induced dimer (iLID) for controlling the localization and activity of signaling proteins. Proc. Natl. Acad. Sci. USA 112, 112–117 (2015).
Kawano, F., Suzuki, H., Furuya, A. & Sato, M. Engineered pairs of distinct photoswitches for optogenetic control of cellular proteins. Nat. Commun. 6, 6256 (2015).
Hughes, R.M., Bolger, S., Tapadia, H. & Tucker, C.L. Light-mediated control of DNA transcription in yeast. Methods 58, 385–391 (2012).
Konermann, S. et al. Optical control of mammalian endogenous transcription and epigenetic states. Nature 500, 472–476 (2013).
Polstein, L.R. & Gersbach, C.A. A light-inducible CRISPR-Cas9 system for control of endogenous gene activation. Nat. Chem. Biol. 11, 198–200 (2015).
Nihongaki, Y., Yamamoto, S., Kawano, F., Suzuki, H. & Sato, M. CRISPR-Cas9-based photoactivatable transcription system. Chem. Biol. 22, 169–174 (2015).
Boulina, M., Samarajeewa, H., Baker, J.D., Kim, M.D. & Chiba, A. Live imaging of multicolor-labeled cells in Drosophila. Development 140, 1605–1613 (2013).
Idevall-Hagren, O., Dickson, E.J., Hille, B., Toomre, D.K. & De Camilli, P. Optogenetic control of phosphoinositide metabolism. Proc. Natl. Acad. Sci. USA 109, E2316–E2323 (2012).
Giordano, F. et al. PI(4,5)P(2)-dependent and Ca(2+)-regulated ER-PM interactions mediated by the extended synaptotagmins. Cell 153, 1494–1509 (2013).
Kakumoto, T. & Nakata, T. Optogenetic control of PIP3: PIP3 is sufficient to induce the actin-based active part of growth cones and is regulated via endocytosis. PLoS One 8, e70861 (2013).
Aoki, K. et al. Stochastic ERK activation induced by noise and cell-to-cell propagation regulates cell density-dependent proliferation. Mol. Cell 52, 529–540 (2013).
O'Neill, P.R. & Gautam, N. Subcellular optogenetic inhibition of G proteins generates signaling gradients and cell migration. Mol. Biol. Cell 25, 2305–2314 (2014).
Maiuri, P. et al. Actin flows mediate a universal coupling between cell speed and cell persistence. Cell 161, 374–386 (2015).
Duan, L. et al. Optogenetic control of molecular motors and organelle distributions in cells. Chem. Biol. 22, 671–682 (2015).
Bugaj, L.J., Choksi, A.T., Mesuda, C.K., Kane, R.S. & Schaffer, D.V. Optogenetic protein clustering and signaling activation in mammalian cells. Nat. Methods 10, 249–252 (2013).
Lee, S. et al. Reversible protein inactivation by optogenetic trapping in cells. Nat. Methods 11, 633–636 (2014).
Taslimi, A. et al. An optimized optogenetic clustering tool for probing protein interaction and function. Nat. Commun. 5, 4925 (2014).
Pathak, G.P., Strickland, D., Vrana, J.D. & Tucker, C.L. Benchmarking of optical dimerizer systems. ACS Synth. Biol. 3, 832–838 (2014).
Liu, Y., Li, X., Li, K., Liu, H. & Lin, C. Multiple bHLH proteins form heterodimers to mediate CRY2-dependent regulation of flowering-time in Arabidopsis. PLoS Genet. 9, e1003861 (2013).
Liu, H. et al. Photoexcited CRY2 interacts with CIB1 to regulate transcription and floral initiation in Arabidopsis. Science 322, 1535–1539 (2008).
Brautigam, C.A. et al. Structure of the photolyase-like domain of cryptochrome 1 from Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 101, 12142–12147 (2004).
Müller, P. et al. ATP binding turns plant cryptochrome into an efficient natural photoswitch. Sci. Rep. 4, 5175 (2014).
Yang, H.Q. et al. The C termini of Arabidopsis cryptochromes mediate a constitutive light response. Cell 103, 815–827 (2000).
Zeugner, A. et al. Light-induced electron transfer in Arabidopsis cryptochrome-1 correlates with in vivo function. J. Biol. Chem. 280, 19437–19440 (2005).
Zoltowski, B.D. et al. Structure of full-length Drosophila cryptochrome. Nature 480, 396–399 (2011).
Partch, C.L. & Sancar, A. Photochemistry and photobiology of cryptochrome blue-light photopigments: the search for a photocycle. Photochem. Photobiol. 81, 1291–1304 (2005).
Engelhard, C. et al. Cellular metabolites enhance the light sensitivity of Arabidopsis cryptochrome through alternate electron transfer pathways. Plant Cell 26, 4519–4531 (2014).
El-Esawi, M. et al. Cellular metabolites modulate in vivo signaling of Arabidopsis cryptochrome-1. Plant Signal. Behav. 10, e1063758 (2015).
Gao, J. et al. Trp triad-dependent rapid photoreduction is not required for the function of Arabidopsis CRY1. Proc. Natl. Acad. Sci. USA 112, 9135–9140 (2015).
Zoltowski, B.D. Resolving cryptic aspects of cryptochrome signaling. Proc. Natl. Acad. Sci. USA 112, 8811–8812 (2015).
Vaidya, A.T. et al. Flavin reduction activates Drosophila cryptochrome. Proc. Natl. Acad. Sci. USA 110, 20455–20460 (2013).
Müller, P. & Bouly, J.-P. Searching for the mechanism of signalling by plant photoreceptor cryptochrome. FEBS Lett. 589, 189–192 (2015).
Stanewsky, R. et al. The cryb mutation identifies cryptochrome as a circadian photoreceptor in Drosophila. Cell 95, 681–692 (1998).
Oztürk, N., Song, S.-H., Selby, C.P. & Sancar, A. Animal type 1 cryptochromes. Analysis of the redox state of the flavin cofactor by site-directed mutagenesis. J. Biol. Chem. 283, 3256–3263 (2008).
Schindler, S.E. et al. Photo-activatable Cre recombinase regulates gene expression in vivo. Sci. Rep. 5, 13627 (2015).
Jullien, N., Sampieri, F., Enjalbert, A. & Herman, J.P. Regulation of Cre recombinase by ligand-induced complementation of inactive fragments. Nucleic Acids Res. 31, e131 (2003).
Li, X. et al. Arabidopsis cryptochrome 2 (CRY2) functions by the photoactivation mechanism distinct from the tryptophan (trp) triad-dependent photoreduction. Proc. Natl. Acad. Sci. USA 108, 20844–20849 (2011).
Thöing, C., Oldemeyer, S. & Kottke, T. Microsecond Deprotonation of Aspartic Acid and Response of the α/β Subdomain Precede C-Terminal Signaling in the Blue Light Sensor Plant Cryptochrome. J. Am. Chem. Soc. 137, 5990–5999 (2015).
Solov'yov, I.A., Domratcheva, T., Moughal Shahi, A.R. & Schulten, K. Decrypting cryptochrome: revealing the molecular identity of the photoactivation reaction. J. Am. Chem. Soc. 134, 18046–18052 (2012).
Herbel, V. et al. Lifetimes of Arabidopsis cryptochrome signaling states in vivo. Plant J. 74, 583–592 (2013).
Bouly, J.P. et al. Cryptochrome blue light photoreceptors are activated through interconversion of flavin redox states. J. Biol. Chem. 282, 9383–9391 (2007).
Banerjee, R. et al. The signaling state of Arabidopsis cryptochrome 2 contains flavin semiquinone. J. Biol. Chem. 282, 14916–14922 (2007).
Matsuda, T. & Cepko, C.L. Controlled expression of transgenes introduced by in vivo electroporation. Proc. Natl. Acad. Sci. USA 104, 1027–1032 (2007).
Cadwell, R.C. & Joyce, G.F. Randomization of genes by PCR mutagenesis. PCR Methods Appl. 2, 28–33 (1992).
Acknowledgements
We thank Dr. Constance Cepko for the pCALVL-dsRed Cre reporter (13769), obtained through Addgene, Dr. Matthew Kennedy for critical reading of the manuscript, and Jessica Spiltoir and Qi Liu for experimental assistance. This work was supported by grants from the National Institutes of Health (GM100225) and the McKnight Endowment Fund for Neuroscience (Technological Innovations in Neuroscience Award) to C.L.T.
Author information
Authors and Affiliations
Contributions
A.T., B.Z., J.G.M., G.P.P., R.M.H., and C.L.T. carried out experiments. B.Z. carried out protein structural characterization and in vitro studies with OtCPF1. A.T., B.Z., and C.L.T. analyzed data and wrote the manuscript. C.L.T. conceived the project and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–7. (PDF 2393 kb)
Supplementary Table 1
Sequences of constructs used in studies (XLS 247 kb)
Rights and permissions
About this article
Cite this article
Taslimi, A., Zoltowski, B., Miranda, J. et al. Optimized second-generation CRY2–CIB dimerizers and photoactivatable Cre recombinase. Nat Chem Biol 12, 425–430 (2016). https://doi.org/10.1038/nchembio.2063
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2063
This article is cited by
-
Tuning CARs: recent advances in modulating chimeric antigen receptor (CAR) T cell activity for improved safety, efficacy, and flexibility
Journal of Translational Medicine (2023)
-
A doxycycline- and light-inducible Cre recombinase mouse model for optogenetic genome editing
Nature Communications (2022)
-
Rapid and reversible optogenetic silencing of synaptic transmission by clustering of synaptic vesicles
Nature Communications (2022)
-
Efficient spatially targeted gene editing using a near-infrared activatable protein-conjugated nanoparticle for brain applications
Nature Communications (2022)
-
A red light–responsive photoswitch for deep tissue optogenetics
Nature Biotechnology (2022)