Abstract
The detrimental effects of global warming on crop productivity threaten to reduce the world's food supply1,2,3. Although plant responses to changes in temperature have been studied4, genetic modification of crops to improve thermotolerance has had little success to date. Here we demonstrate that overexpression of the Arabidopsis thaliana receptor-like kinase ERECTA (ER) in Arabidopsis, rice and tomato confers thermotolerance independent of water loss and that Arabidopsis er mutants are hypersensitive to heat. A loss-of-function mutation of a rice ER homolog and reduced expression of a tomato ER allele decreased thermotolerance of both species. Transgenic tomato and rice lines overexpressing Arabidopsis ER showed improved heat tolerance in the greenhouse and in field tests at multiple locations in China during several seasons. Moreover, ER-overexpressing transgenic Arabidopsis, tomato and rice plants had increased biomass. Our findings could contribute to engineering or breeding thermotolerant crops with no growth penalty.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Peng, S. et al. Rice yields decline with higher night temperature from global warming. Proc. Natl. Acad. Sci. USA 101, 9971–9975 (2004).
Lobell, D.B. & Field, C.B. Global scale climate-crop yield relationships and the impacts of recent warming. Environ. Res. Lett. 2, 014002 (2007).
Long, S.P. & Ort, D.R. More than taking the heat: crops and global change. Curr. Opin. Plant Biol. 13, 241–248 (2010).
Penfield, S. Temperature perception and signal transduction in plants. New Phytol. 179, 615–628 (2008).
Fitter, A.H. & Fitter, R.S. Rapid changes in flowering time in British plants. Science 296, 1689–1691 (2002).
Samach, A. & Wigge, P.A. Ambient temperature perception in plants. Curr. Opin. Plant Biol. 8, 483–486 (2005).
Kumar, S.V. & Wigge, P.A. H2A.Z-containing nucleosomes mediate the thermosensory response in Arabidopsis. Cell 140, 136–147 (2010).
Posé, D.S.H. et al. Temperature-dependent regulation of flowering by antagonistic FLM variants. Nature 503, 414–417 (2013).
Queitsch, C., Hong, S.W., Vierling, E. & Lindquist, S. Heat shock protein 101 plays a crucial role in thermotolerance in Arabidopsis. Plant Cell 12, 479–492 (2000).
Finka, A., Cuendet, A.F.H., Maathuis, F.J.M. & Saidi, Y. Plasma membrane cyclic nucleotide gated calcium channels control land plant thermal sensing and acquired thermotolerance. Plant Cell 24, 3333–3348 (2012).
Iba, K. Acclimative response to temperature stress in higher plants: approaches of gene engineering for temperature tolerance. Annu. Rev. Plant Biol. 53, 225–245 (2002).
Kim, M., Lee, U., Small, I., des Francs-Small, C.C. & Vierling, E. Mutations in an Arabidopsis mitochondrial transcription termination factor-related protein enhance thermotolerance in the absence of the major molecular chaperone HSP101. Plant Cell 24, 3349–3365 (2012).
Guan, Q., Yue, X., Zeng, H. & Zhu, J. The protein phosphatase RCF2 and its interacting partner NAC019 are critical for heat stress–responsive gene regulation and thermotolerance in Arabidopsis. Plant Cell 26, 438–453 (2014).
Burke, J.J. & Chen, J. Enhancement of reproductive heat tolerance in plants. PLoS ONE 10, e0122933 (2015).
Li, X. et al. Natural alleles of a proteasome α2 subunit gene contribute to thermotolerance and adaptation of African rice. Nat. Genet. 47, 827–833 (2015).
Grover, A., Mittal, D., Negi, M. & Lavania, D. Generating high temperature tolerant transgenic plants: Achievements and challenges. Plant Sci. 205-206, 38–47 (2013).
Hancock, A.M. et al. Adaptation to climate across Arabidopsis thaliana genome. Science 334, 83–86 (2011).
Fournier-Level, A. et al. A map of local adaptation in Arabidopsis thaliana. Science 334, 86–89 (2011).
Yan, C., Shen, H., Li, Q. & He, Z. A novel ABA-hypersensitive mutant in Arabidopsis defines a genetic locus that confers tolerance to xerothermic stress. Planta 224, 889–899 (2006).
Lin, L., Zhong, S.H., Cui, X.F., Li, J. & He, Z.H. Characterization of temperature- sensitive mutants reveals a role for receptor-like kinase CRAMBLED/STRUBBELIG in coordinating cell proliferation and differentiation during Arabidopsis leaf development. Plant J. 72, 707–720 (2012).
Zhong, S. et al. Warm temperatures induce transgenerational epigenetic release of RNA silencing by inhibiting siRNA biogenesis in Arabidopsis. Proc. Natl. Acad. Sci. USA 110, 9171–9176 (2013).
Torii, K.U. et al. The Arabidopsis ERECTA gene encodes a putative receptorprotein kinase with extracellular leucine-rich repeats. Plant Cell 8, 735–746 (1996).
Shpak, E.D., Berthiaume, C.T., Hill, E.J. & Torii, K.U. Synergistic interaction of three ERECTA-family receptor-like kinases controls Arabidopsis organ growth and flower development by promoting cell proliferation. Development 131, 1491–1501 (2004).
Shpak, E.D., McAbee, J.M., Pillitteri, L.J. & Torii, K.U. Stomatal patterning and differentiation by synergistic interactions of receptor kinases. Science 309, 290–293 (2005).
van Zanten, M., Snoek, L.B., Proveniers, M.C. & Peeters, A.J. The many functions of ERECTA. Trends Plant Sci. 14, 214–218 (2009).
Lee, J.S. et al. Direct interaction of ligand-receptor pairs specifying stomatal patterning. Genes Dev. 26, 126–136 (2012).
Uchida, N. et al. Regulation of inflorescence architecture by intertissue layer ligand-receptor communication between endodermis and phloem. Proc. Natl. Acad. Sci. USA 109, 6337–6342 (2012).
Tisné, S. et al. Combined genetic and modeling approaches reveal that epidermal cell area and number in leaves are controlled by leaf and plant developmental processes in Arabidopsis. Plant Physiol. 148, 1117–1127 (2008).
Qi, Y., Sun, Y., Xu, L., Xu, Y. & Huang, H. ERECTA is required for protection against heat-stress in the AS1/AS2 pathway to regulate adaxial-abaxial leaf polarity in Arabidopsis. Planta 219, 270–276 (2004).
Masle, J., Gilmore, S.R. & Farquha, G.D. The ERECTA gene regulates plant transpiration efficiency in Arabidopsis. Nature 436, 866–870 (2005).
Li, S. et al. HEAT-INDUCED TAS1 TARGET1 mediates thermotolerance via HEAT STRESS TRANSCRIPTION FACTOR A1a–directed pathways in Arabidopsis. Plant Cell 26, 1764–1780 (2014).
Ishikawa, T., Uchimiya, H. & Kawai-Yamada, M. The role of plant Bax inhibitor-1 in suppressing H2O2-induced cell death. Methods Enzymol. 527, 239–256 (2013).
Villagarcia, H., Morin, A.C., Shpaket, E.D. & Khodakovskaya, M.V. Modification of tomato growth by expression of truncated ERECTA protein from Arabidopsis thaliana. J. Exp. Bot. 63, 6493–6504 (2012).
Mickelbart, M.V., Hasegawa, P.M. & Bailey-Serres, J. Genetic mechanisms of abiotic stress tolerance that translate to crop yield stability. Nat. Rev. Genet. 16, 237–251 (2015).
Xing, H.T., Guo, P., Xia, X.L. & Yin, W.L. PdERECTA, a leucine-rich repeat receptor-like kinase of poplar, confers enhanced water use efficiency in Arabidopsis. Planta 234, 229–241 (2011).
Shiu, S.H. et al. Comparative analysis of the receptor-like kinase family in Arabidopsis and rice. Plant Cell 16, 1220–1234 (2004).
Jagadish, S.V. et al. Physiological and proteomic approaches to address heat tolerance during anthesis in rice (Oryza sativa L.). J. Exp. Bot. 61, 143–156 (2010).
Lister, C. & Dean, C. Recombinant inbred lines for mapping RFLP and phenotypic markers in Arabidopsis thaliana. Plant J. 4, 745–750 (1993).
Clough, S.J. & Bent, A.F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Wang, E.T. et al. Control of rice grain-filling and yield by a gene with a potential signature of domestication. Nat. Genet. 40, 1370–1374 (2008).
The Tomato Genome Consortium. The tomato genome sequence provides insights into fleshy fruit evolution. Nature 485, 635–641 (2012).
Pennycooke, J.C. et al. The low temperature-responsive, Solanum CBF1 genes maintain high identity in their upstream regions in a genomic environment undergoing gene duplications, deletions, and rearrangements. Plant Mol. Biol. 67, 483–497 (2008).
Zhang, H., Zhang, X., Mao, B., Li, Q. & He, Z. Alpha-picolinic acid, a fungal toxin and mammal apoptosis-inducing agent, elicits hypersensitive-like response and enhances disease resistance in rice. Cell Res. 14, 27–33 (2004).
Parida, A.K., Dasgaonkar, V.S., Phalak, M.S., Umalkar, G.V. & Aurangabadkar, L.P. Alterations in photosynthetic pigments, protein, and osmotic components in cotton genotypes subjected to short-term drought stress followed by recovery. Plant Biotechnol. Rep. 1, 37–48 (2007).
Karaba, A. et al. Improvement of water use efficiency in rice by expression of HARDY, an Arabidopsis drought and salt tolerance gene. Proc. Natl. Acad. Sci. USA 104, 15270–15275 (2007).
Charng, Y.-Y. et al. A heat-inducible transcription factor, HsfA2, is required for extension of acquired thermotolerance in Arabidopsis. Plant Physiol. 143, 251–262 (2007).
Weng, L. et al. The zinc finger transcription factor SlZFP2negatively regulates abscisic acid biosynthesis and fruit ripening in tomato. Plant Physiol. 167, 931–949 (2015).
Acknowledgements
We thank J.-S. Jeon and C.Y. Wu for the rice T-DNA mutants, X.L. Wang for the er-105 line, X.G. Zhu, Y.J. Zhang and M.Z. Lv for help in statistical analysis, D.Y. Sun and H.X. Lin for helpful discussions, and L. Lin for help in experiments. This work was supported by the National Key Basic Research and Development Program (2011CB100700 to H.Z.), the National Natural Science Foundation of China (3130061 to H.Z.), the Ministry of Science and Technology of China (2012AA10A302 to Z.H.), the National GMO project (2013ZX08009-003-001 to Z.H.), the National High Technology Research and Development Program of China (2012AA100104-6 to H.X.), the Chinese Academy of Sciences (KSCX2-EW-N-01 to H.Z. and 2009OHTP07 to H.X.), the Shanghai Committee of Science and Technology (11PJ1410900 to H.X.), and the National Key Basic Research and Development Program (2015CB150104 to G.C.).
Author information
Authors and Affiliations
Contributions
H.S., X.Z., F.Z., Y.W., G.C., Y.H., H.X., J.L. and Z.H. conceived the research project, designed experiments and analyzed the data. H.S., X.Z., F.Z., Y.W., B.Y., Q.L., G.C., B.M., J.W., Y.L. and G.X. conducted the experiments. Z.H. and J.L. oversaw the entire study and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 QTL mapping for heat tolerance
In the preliminary QTL mapping, the RILs derived from the cross of Col-0 and Ler were classified into 3 groups according to their heat tolerance phenotypes after 40°C treatment for 2 d: lines with high tolerance similar to Col-0 (survival rate 30.8-42.4%), lines with high sensitivity similar to Ler (survival rate 7.5-12.4%), and lines with intermediate properties (survival rate 14.7-28.4%). Values are means ± SD (n = 3). (b) QTL distribution in 5 chromosomes. Note that chromosome 2 has a major contribution peak. The horizontal lines plotted for chromosomes 2 and 4 indicated the QTL LOD threshold. (c) Construction of CSSLs that cover chromosome 2. (d) Fine mapping of qHat2-1, which is located at CSSL2-3 that contains the mutated ERECTA (er) gene in BAC T1D16 in the Ler mutant (red arrow). Survival rates were measured for each CSSL2 lines, shown as dots of 3 replicates (30 plants each). Different letters at the top of dots indicate a significant difference at P < 0.05, by Welch’s ANOVA and Bonferroni correction for multiple (6) tests (h,i).
Supplementary Figure 2 Genetic complementation of the ER gene
(a) Heat tolerance of 2-week-old wild-type Col-0, er-105, and complementation pER::ER/er-105 plants treated at 40°C for 48 h. (b) Survival rates of the Col-0, er-105, and complementation plants as shown in dots of 3 replicates (30 plants each). (c) Heat tolerance of two-week-old wild-type Lan, Ler, and complementation pER::ER/Ler plants treated with 40°C for 48 h. (d) Survival rates of Lan, Ler mutants, and complementation plants as shown in dots of 3 replicates (30 plants each). Asterisks indicate a significant difference in comparison with wild-type Col-0 at 40°C, at P < 0.01 (**), by Student’s t-test and Bonferroni correction for multiple (2) tests in the independent temperature experiments (b,d). Bar = 1 cm (a,c).
Supplementary Figure 3 Expression levels of ER and ER homologous genes and leaf sizes in 35S::ER-overexpressing plants
(a) Expression levels of ER in rosette leaves of 2-week-old grown at 22°C wild-type Col-0 and ER-OE lines (T3) and in two continuous homozygous generations (T3 and T4, line L7-1). Values are means ± SD (n = 3). (b) Expression levels of ER in rosette, cauline leaves and inflorescence organs of 6-week-old grown at 22°C wild-type Col-0 and the ER-OE line (L7-1, T3) detected by qPCR. ACTIN2 was used as control to normalise expression levels. Note that ER was stably overexpressed in different tissues of the transgenic plants. Number (2.79) above L7-1 (flower) indicates fold relative to expression level (1) in the control Col-0. Values are means ± SD (n = 3). (c) Ten rosette leaf sizes of 4-week-old ER-OE, Col-0 plants and er-105 plants grown at 22°C. Values are means ± SD (n = 10). (d) Rosette leaf numbers of ER-OE, Col-0 and er-105 plants at rosette (4-week-old) and flowering (5-week-old) stages, grown at 22°C. Values are means ± SD (n = 30). (e) The expression of the homologous ERL1 and ERL2 in rosette, cauline leaves and inflorescence organs of 2- and 6-week-old wild-type Col-0 and ER-OE line (line 7-1, T3) grown at 22°C was not affected by ER overexpression as detected by qPCR. ACTIN2 was used as control to normalise expression levels. Values are means ± SD (n = 3). Asterisks indicate statistically significant differences as determined by Student’s t-test (*, P< 0.05; **, P < 0.01) (c,d).
Supplementary Figure 4 Development of 35S::ER-MH/er105 plants
(a) Transcription levels of ER-MH detected by qPCR, ACTIN2 was used as control to normalise expression levels. Values are means ± SD (n = 3). Number (2.64) above L13 indicates fold relative to expression level (1) in the control Col-0. (b) Plants and leaves of 2-week-old 35S::ER-MH/er105 (LM10 and LM17) lines grown at 22°C in comparison with wild-type Col-0 and er-105, which accumulated high levels of the ER-MH fusion protein and showed increased thermotolerance (Fig. 1d,j). Bar = 1 cm.
Supplementary Figure 5 Cell morphology of 2-week-old Col-0, er-105 and ER-OE plants grown at 22°C
(a) Leaf cells of Col-0 and ER-OE plants (L7-1). Note that the mesophyll cells are larger in L7-1 than in Col-0. (b) SEM images of leaf epidermal cells and stomata of Col-0, er-105 and 35S::ER plants. (c) Palisade cell width of Col-0, er-105 and 35S::ER plants, as shown with boxes (n = 30). Asterisks indicate statistically significant differences as determined by Student’s t-test (*, P < 0.05). (d) Palisade cell length of Col-0, er-105 and ER-OE plants, shown as boxes (n = 30). Asterisks indicate statistically significant differences as determined by Student’s t-test (*, P < 0.05). (e) Stomatal density was significantly increased and decreased in er-105 and ER-OE plants compared with Col-0 plants, respectively. As shown with boxes (n = 25). Asterisks indicate a significant difference in comparison with wild-type Col-0, at P < 0.01, by Student’s t-test and Bonferroni correction for multiple (3) tests. (f) Stomatal index was not changed in er-105 and ER-OE plants, as shown with boxes (n = 25). Bar = 50 μm (a,b).
Supplementary Figure 6 Increased drought tolerance of ER-OE plants
(a) Four-week-old Col-0, er-105, and ER-OE (L7-1) plants grown at 22°C were subjected to drought treatment until the er-105 plants died and then were re-watered to recovery for 2 d. (b) Survival rates of Col-0, er-105 mutant, and L7-1plants with or without (control) drought treatment, as shown in dots of 3 replicates (30 plants each). Asterisks indicate a significant difference in comparison with wild-type Col-0, at P < 0.05 (*) or P < 0.01(**), by Student’s t-test and Bonferroni correction for multiple (2) tests for drought treatment.
Supplementary Figure 7 Leaf epidermal morphology of two-week-old Col-0 and er-105 under heat stress
Leaf epidermal cell morphology of Col-0 and er-105 plants under normal (40°C, 0 h) and heat (40°C, 12 h) treatments. Severely wrinkled leaf epiderm was observed in er-105 under heat stress. Bar = 50 μm.
Supplementary Figure 8 Subcellular observation by TEM during heat treatment
(a) TEM observation of leaf cell collapse of the two-week-old er-105 and Col-0 plants treated by 40ºC for 0 and 24 h. Red arrow indicates disrupted plasma membrane in er-105, and arrowhead indicates plasma membrane collapsing in Col-0. Bar = 10 μm. (b-e) An earlier and more severe collapse of the chloroplast (b,d) and mitochondria (c,e), with formation of spherical bodies, occurred in the er-105 cells than in the Col-0 cells under heat, which was less or delayed in the L7-1 cells compared with Col-0 cells. Asterisks indicate a significant difference in comparison with wild-type Col-0, at P < 0.05 (*) or P < 0.01 (**), by Student’s t-test and Bonferroni correction for multiple (2) tests, as shown in dots of 3 replicates (d,e). Bar = 1 μm (b) or 100 nm (c). cw, cell wall; v, vacuole; sp, spherical bodies; c, chloroplast; m, mitochondria; s, starch granule.
Supplementary Figure 9 Expression inhibition of ER at high temperature (40°C)
Relative expression levels of ER were determined by qPCR in two-week-old Col-0 plants treated by heat for 0h (control) to 24 h. ACTIN2 was used as control to normalise expression levels. Values are means ± SD (n = 3).
Supplementary Figure 10 Leaf size and cell morphology of transgenic tomato plants
(a) Three leaf sizes of 6-week-old 35S::ER transgenic tomato lines and the empty-vector transgenic control grown at 22°C. Values are means ± SD (n = 10). Asterisks indicate statistically significant differences as determined by Student’s t-test (**, P < 0.01). (b) Enlarged epidermal leaf cells of the 35S::ER transgenic tomato plants in comparison with the vector control grown at 22°C. Bar = 20 μm. (c) Decreased stomatal density in the transgenic tomato plants, as shown with boxes (n = 25). Asterisks indicate a significant difference in comparison with vector control, at P < 0.01, by Student’s t-test and Bonferroni correction for multiple (3) tests. (d) No change in the stomatal index was observed in the transgenic tomato plants, as shown with boxes (n = 25).
Supplementary Figure 11 Stomatal conductance, transpiration efficiency and cell death of transgenic tomato plants
(a) Stomatal conductance of 35S::ER transgenic tomato and the empty-vector control plants (6-week-old) under normal growth conditions (22°C), shown as dots (n ≥ 8). (b) Increased transpiration efficiency (instantaneous WUE) in 35S::ER transgenic tomato relative to the empty-vector transgenic control under normal growth conditions (22°C), shown as dots (n ≥ 8). (c) Decreased cell death in 35S::ER transgenic tomato relative to the empty-vector transgenic control under heat treatment. Bar = 1mm. Asterisks indicate a significant difference in comparison with vector control, at P < 0.05 (*) or P < 0.01 (**), by Student’s t-test and Bonferroni correction for multiple (2) tests (a,b).
Supplementary Figure 12 Field tests of transgenic tomato in different locations and seasons
Survival tests in Shanghai (summer, 2012) (a), Wuhan (summer, 2013) (b), and in Hainan (early summer, 2013) (c). Fifty plants of each line were directly compared for survival/death rates. Tomato plants were also grown in early summer (early June-early July) of 2014 in the Shanghai station as normal growth conditions with 3 biological replicates, no statistical difference in survival rate was observed in the test, as shown with dots of 3 replicates (d). The local temperature and humidity conditions for these field tests were shown in Data Set 1.
Supplementary Figure 13 Flower number and fruit weight of tomato
Plants were grown at the greenhouse (22°C) for 10 weeks, flower number was counted (3 inflorescences each plant, 10 plants) (a), and fruit weight (b) was measured for fruits produced by hand-pollination (c), shown as boxes (n = 30 for a, n = 20 for b). Bar = 1 cm (c).
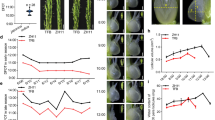
Supplementary Figure 14 Agronomic traits of transgenic rice plants under normal temperature conditions
(a) Field-grown mature plants of transgenic rice plants compared with the empty-vector control line (T1 generation). (b) No difference was observed in plant height of adult transgenic plants relative to the control transgenic line after heading, as shown with boxes (n = 20). (c,d) One-week-old transgenic rice seedlings were significantly larger than the control, with a larger leaf width (c) and whole plant height (from root tip to leaf tip) (d), as shown with boxes (n = 20); asterisks indicate a significant difference in comparison with vector control, at P < 0.05 (*) or P < 0.01 (**), by Student’s t-test and Bonferroni correction for multiple (2) tests (c,d).
Supplementary Figure 15 Yield traits in the field tests of rice in different locations and seasons
(a-c) Yield performance of the transgenic lines grown to flower in the autumn of 2013 in the Shanghai station as the normal temperature growth conditions as shown in boxes (n ≥ 30), showing no difference in seed setting (a) and panicle number (c). Lines 36 and 38 displayed significant higher grain yield potential in comparison with the vector control (b), asterisks indicate a significant difference in comparison with vector control, at P < 0.05, by Student’s t-test. (d) Panicle numbers per plant flowering in the summer of 2013 at the Shanghai, Wuhan and Changsha stations, shown as boxes (n ≥ 30). Asterisks indicate a significant difference in comparison with vector control, at P < 0.05 (*) or P < 0.01 (**), by Student’s t-test and Bonferroni correction for multiple (3) tests in the Shanghai (2013, 2014) station, or at P < 0.05, by Student’s t-test in the Changsha station. (e) Average grain number per panicle in the summers of 2013 and 2014, shown as boxes (n ≥ 30). (f) Grain weight measurements in the field tests of the 2013 and 2014 summers as shown with boxes (n = 5).
Supplementary Figure 16 Characterization of the rice T-DNA insertion mutants Oser1 and Oser2
(a) Identification of the T-DNA insertion mutant Oser1 that contains a T-DNA insert between exon 11 and 12, abolishing the expression of the OsER1gene detected by qPCR. Values are means ± SD (n = 3). (b) The T-DNA inserted into exon 24 of the rice ER homolog gene OsER2, resulting into a null mutant that abolishes OsER2 expression detected by qPCR. Values are means ± SD (n = 3). The rice ACTIN1 gene was used as control to normalise expression levels (a,b). (c) No difference was observed in thermotolerance in the mutant in comparison with the wild-type control during heat treatment (42°C). Pictures were taken after a one-week recovery period following 10-d treatment at 42°C/35°C (day/night) in growth chamber.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–16; Supplementary Tables 1–2 (PDF 2069 kb)
Supplementary Dataset 1
Local temperature and humidity records of field tests (XLSX 182 kb)
Rights and permissions
About this article
Cite this article
Shen, H., Zhong, X., Zhao, F. et al. Overexpression of receptor-like kinase ERECTA improves thermotolerance in rice and tomato. Nat Biotechnol 33, 996–1003 (2015). https://doi.org/10.1038/nbt.3321
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3321
This article is cited by
-
The OsSGS3-tasiRNA-OsARF3 module orchestrates abiotic-biotic stress response trade-off in rice
Nature Communications (2023)
-
Regulatory network of rice in response to heat stress and its potential application in breeding strategy
Molecular Breeding (2023)
-
Secondary Metabolites Mediated Reproductive Tolerance Under Heat Stress in Plants
Journal of Plant Growth Regulation (2023)
-
Insights into genomic variations in rice Hsp100 genes across diverse rice accessions
Planta (2023)
-
Overexpression of Zostera japonica heat shock protein gene ZjHsp70 enhances the thermotolerance of transgenic Arabidopsis
Molecular Biology Reports (2022)