Abstract
Microdeletions in the 10q26.1 region are related to intellectual disability, growth delay, microcephaly, distinctive craniofacial features, cardiac defects, genital abnormalities and inner ear abnormalities. The genes responsible for inner ear abnormalities have been narrowed to fibroblast growth factor receptor 2 gene (FGFR2), H6 family homeobox 2 gene (HMX2) and H6 family homeobox 3 gene (HMX3). An additional patient with distinctive craniofacial features, congenital deafness and balance dysfunctions showed a de novo microdeletion of 10q26.11q26.13, indicating the existence of a gene responsible for inner ear abnormalities in this region.
Similar content being viewed by others
Many patients have been reported to have 10q monosomies.1–4 The 10q26.1 region is proximal to the 10q telomere. Patients with interstitial microdeletions within this region show characteristic findings that differ from those of patients with terminal deletions of 10q. These characteristics include intellectual disability, growth delay, microcephaly, distinctive craniofacial features, cardiac defects, genital abnormalities and inner ear abnormalities, including deafness and balance dysfunction.5,6 By performing a genotype–phenotype correlational study, others have suggested critical regions related to each clinical finding.7 In this study, we identified an additional case of a patient with inner ear abnormalities associated with a 10q26.1 microdeletion. This finding provides further evidence supporting the existence of a gene responsible for deafness in this region.
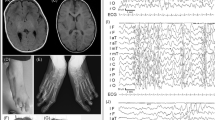
A 22-month-old Japanese girl was born to healthy and non-consanguineous parents following in vitro fertilization and gestation. She was born with a standard stature. Her family history was not remarkable. Patent ductus arteriosus was identified in the patient, and, soon after, it was surgically repaired. She exhibited a mild motor developmental delay: crawling at 12 months and walking alone at 20 months. Congenital deafness at a level of 50–60 dB was confirmed, and a hearing aid was required. A computed tomography scan of the temporal bone was performed, and bilateral middle ear hypoplasia was identified (Figures 1a,b).
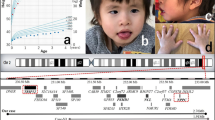
The results of the radiological and genetic examinations. Computed tomography scans focused on the middle ears (a; right side, b; left side) show that bilateral lateral semicircular canals are dilated, shortened and non-circular (white arrows). Bilateral anterior superior semicircular canals and bilateral posterior semicircular canals are hypoplastic (black arrows). Hypoplastic findings are extreme on the right side. (c) Loss of genomic copy number involving 10q26.11q16.13 is shown using Chromosome View by Agilent Genomic Workbench (Agilent Technologies). (d) A 10q26.11q16.13 microdeletion is confirmed by fluorescence in situ hybridization (FISH). Red signals represent markers of 10p15.3 labeled on RP11–387K19, and green signals are targets of 10q26.13 labeled on RP11–57J8. A loss of the green signal (a white arrow) indicates a deletion of this region.
At present, the patient’s height is 79.2 cm (−1.2 s.d.), weight is 9.6 kg (−1.1 s.d.) and occipitofrontal circumference is 46 cm (−0.7 s.d.). She shows distinctive craniofacial features, including facial asymmetry, malformed ears associated with low-set and posteriorly rotated ears, epicanthic folds and a lateral cervical fistula. She can walk alone but with instability; she easily falls down and often suffers injuries. This may indicate difficulties in balance due to hypoplastic semicircular canals identified by computed tomography. Her intelligent quotient was evaluated as 92 according to the Kyoto Scale of Psychological Development.
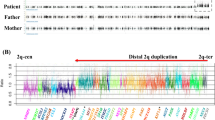
The study was performed in accordance with the principles outlined in the Declaration of Helsinki and was approved by the ethics committee of Tokyo Women’s Medical University. Blood samples were obtained from the patient and her parents after receiving written informed consent. Chromosomal microarray testing using the Agilent 60 K Human Genome CGH Microarray platform (Agilent Technologies, Santa Clara, CA, USA) was performed in accordance with previous descriptions of the method.8 A genomic copy number loss of 10q26.11q26.13 was identified in the patient. The molecular karyotype was arr 10q26.11q26.13(120,807,022–126,581,953)×1 (Figure 1c), indicating a 5.8-Mb deletion. Fluorescence in situ hybridization (FISH) analysis, using the bacterial artificial chromosome RP11–57J8 as a target probe, confirmed the deletion on the homologous chromosome 10q26.13 (Figure 1d). Parental FISH analysis showed no abnormality, determining a de novo occurrence in the patient. In this study, genomic positions were referred to as GRCh37/hg19.
We identified a de novo microdeletion of 10q26.11q26.13 in a patient with distinctive craniofacial features and congenital deafness. Using a genotype–phenotype correlational analysis, we constructed a genome map and depicted the deletions identified in this patient and in previously reported patients (Figure 2). All clinical manifestations of the present patient were common to the previously reported patients with microdeletions in this region. Facial asymmetry, which was observed in this patient, was common to the patient reported as Case 14 by Irving et al.4
The genome map around the 10q26.1 region. Deletion regions of the present patient and previously reported patients are depicted by bars. Deletions in patients with and without inner ear abnormalities are shown in red and blue, respectively. Dots at the end of the bars indicate more expansion of the deletions. A reticulated region indicates an ambiguous region due to the results obtained by conventional karyotyping. The genes discussed in the text are indicted by black rectangles. Proposed critical regions are designated by arrows.
Extra genital abnormality is one of the characteristic findings of male patients with 10q26.1 deletions.9,10 The critical region for genital abnormality was suggested to be in the 10q25.3-q26.11 region, where some of the possible candidate genes, such as the empty spiracles homeobox 2 gene (EMX2), are located.5,7,11 Because the present patient was female, there was no extra genital abnormality, and the critical region for genital abnormalities could not be discussed.
Instead, we focused on the critical region for inner ear abnormalities. Miller et al.6 identified overlapping deletions of this region in patients with deafness and balance dysfunction; the commonly deleted region was suggested to be a critical region for such inner ear abnormalities (Figure 2). They suggested two contiguous genes, HMX2 and HMX3, as possible candidate genes because Hmx2/3 double knockout mice had altered vestibular dysfunction.12 Consequently, haploinsufficiency of HMX2 and HMX3 may be related to inner ear abnormalities, although mice with heterozygous mutations in the genes showed no such findings. Such a discrepancy is commonly observed and could be due to the difference in species.
FGFR2 was also considered a possible candidate. It is well known that gain-of-function mutations of this gene are responsible for craniosynostosis. Although the functional relevance of FGFR2 haploinsufficiency is unknown, it may be implicated in the development of distinctive craniofacial features in affected patients.7
As shown in Figure 2, three patients whose deletion regions were within the critical region did not show deafness. As a result, the genotype–phenotype correlation for inner ear abnormalities was unclear. The lack of deafness in these patients may be due to incomplete penetrance.
In conclusion, the present patient with inner ear abnormalities has a microdeletion in 10q26.11q26.13, suggesting that the gene responsible for these abnormalities may be located in the suggested critical region within 10q26.13.
References
References
Courtens W, Wuyts W, Rooms L, Pera SB, Wauters J . A subterminal deletion of the long arm of chromosome 10: a clinical report and review. Am J Med Genet A 2006; 140: 402–409.
Vera-Carbonell A, Lopez-Gonzalez V, Bafalliu JA, Ballesta-Martinez MJ, Fernandez A, Guillen-Navarro E et al. Clinical comparison of 10q26 overlapping deletions: delineating the critical region for urogenital anomalies. Am J Med Genet A 2015; 167A: 786–790.
Plaisancie J, Bouneau L, Cances C, Garnier C, Benesteau J, Leonard S et al. Distal 10q monosomy: new evidence for a neurobehavioral condition? Eur J Med Genet 2014; 57: 47–53.
Irving M, Hanson H, Turnpenny P, Brewer C, Ogilvie CM, Davies A et al. Deletion of the distal long arm of chromosome 10; is there a characteristic phenotype? A report of 15 de novo and familial cases. Am J Med Genet A 2003; 123A: 153–163.
Yatsenko SA, Kruer MC, Bader PI, Corzo D, Schuette J, Keegan CE et al. Identification of critical regions for clinical features of distal 10q deletion syndrome. Clin Genet 2009; 76: 54–62.
Miller ND, Nance MA, Wohler ES, Hoover-Fong JE, Lisi E, Thomas GH et al. Molecular (SNP) analyses of overlapping hemizygous deletions of 10q25.3 to 10qter in four patients: evidence for HMX2 and HMX3 as candidate genes in hearing and vestibular function. Am J Med Genet A 2009; 149A: 669–680.
Choucair N, Abou Ghoch J, Fawaz A, Megarbane A, Chouery E . 10q26.1 microdeletion: redefining the critical regions for microcephaly and genital anomalies. Am J Med Genet A 2015; 167A: 2707–2713.
Shimojima K, Okamoto N, Tamasaki A, Sangu N, Shimada S, Yamamoto T . An association of 19p13.2 microdeletions with Malan syndrome and Chiari malformation. Am J Med Genet A 2015; 167A: 724–730.
Mardo V, Squibb EE, Braverman N, Hoover-Fong JE, Migeon C, Batista DA et al. Molecular cytogenetic analysis of a de novo interstitial deletion of chromosome 10q (q25.3q26.13) in a male child with ambiguous genitalia: evidence for a new critical region for genital development. Am J Med Genet A 2008; 146A: 2293–2297.
Tosur M, Geary CA, Matalon R, Radhakrishnan RS, Swischuk LE, Tarry WF et al. Persistence of mullerian duct structures in a genetic male with distal monosomy 10q. Am J Med Genet A 2015; 167A: 791–796.
Piard J, Mignot B, Arbez-Gindre F, Aubert D, Morel Y, Roze V et al. Severe sex differentiation disorder in a boy with a 3.8 Mb 10q25.3-q26.12 microdeletion encompassing EMX2. Am J Med Genet A 2014; 164A: 2618–2622.
Wang W, Lufkin T . Hmx homeobox gene function in inner ear and nervous system cell-type specification and development. Exp Cell Res 2005; 306: 373–379.
Data Citations
Yamamoto, Toshiyuki HGV Database (2016) http://dx.doi.org/10.6084/m9.figshare.hgv.790
Yamamoto, Toshiyuki HGV Database (2016) http://dx.doi.org/10.6084/m9.figshare.hgv.793
Yamamoto, Toshiyuki HGV Database (2016) http://dx.doi.org/10.6084/m9.figshare.hgv.796
Acknowledgements
We would like to express our gratitude to the patient and her parents for their cooperation. This research was supported by the Practical Research Project for Rare/Intractable Diseases from the Japan Agency for Medical Research and Development (AMED) and the Japan Society for the Promotion of Science (JSPS) KAKENHI Grant Number 15K09631 (TY).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Sangu, N., Okamoto, N., Shimojima, K. et al. A de novo microdeletion in a patient with inner ear abnormalities suggests that the 10q26.13 region contains the responsible gene. Hum Genome Var 3, 16008 (2016). https://doi.org/10.1038/hgv.2016.8
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/hgv.2016.8
This article is cited by
-
Intellectual disability: dendritic anomalies and emerging genetic perspectives
Acta Neuropathologica (2021)
-
A 7q31.33q32.1 microdeletion including LRRC4 and GRM8 is associated with severe intellectual disability and characteristics of autism
Human Genome Variation (2017)
-
Loss-of-function mutations and global rearrangements in GPC3 in patients with Simpson–Golabi–Behmel syndrome
Human Genome Variation (2016)