Abstract
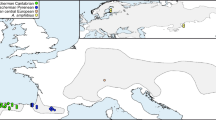
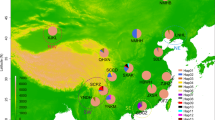
An extensive survey of mitochondrial DNA (mtDNA) restriction polymorphism in 156 isofemale lines from 29 different geographic populations of Drosophila subobscura distributed throughout the Old World was carried out. Ten restriction enzymes were used, five of which revealed restriction site polymorphism. Of the 31 restriction sites detected, 13 were found to be polymorphic. Comparisons with the mtDNA map of Drosophila yakuba indicate that the variable sites are mainly concentrated in protein genes, especially those corresponding to the NADH complex. A total of 13 different haplotypes were observed, two of which (haplotypes I and II) are quite frequent and widely distributed throughout the populations, whereas the other 11 with the exception of VIII, which deserves special attention, are each restricted to one population only and occur at low frequencies. The observed distribution of haplotypes, corroborated by a parsimonious unrooted tree, suggests an ancient origin of haplotypes I and II in the continent.
In order to compare genetic structure according to mtDNA and allozymes, the 10 populations with higher population sizes were studied for 10 polymorphic allozymes also. One striking result is the high degree of population structure of the mtDNA when compared to that obtained for allozymes. If an island model is assumed, estimates of gene flow give values of 0.013 and 1.89 migrants per generation for mtDNA and allozymes, respectively. What is apparent from these estimates is that Drosophila subobscura populations are effectively subdivided for mtDNA genes at migration rates at which nuclear genes (allozymes) are almost panmictic.
Similar content being viewed by others
Article PDF
References
Afonso, J M, Volz, A, Hernandez, M, Ruttkay, H, Gonzalez, A M, Larruga, J M, Cabrera, V, and Sperlich, D. 1990. Mitochondrial DNA variation and genetic structure in Old-World populations of Drosophila subobscura. Mol Biol Evol, 7, 123–142.
Ashley, M V, and Wills, C. 1989. Mitochondrial and allozyme divergence patterns are correlated among island deer mice. Evolution, 43, 646–650.
Avise, J C, Arnold, J, Ball, R M, Bermingham, E, Lamb, T, Neigel, J E, Reeb, C A, and Saunders, N. 1987. Intraspecific phylogeography: the mitochondrial DNA bridge between population genetics and systematics. Ann Rev Ecol Syst, 18, 489–522.
Avise, J C, and Lansman, R C. 1983. Polymorphism of mitochondrial DNA in populations of higher animals. In: Nei, M. and Koehn, R. K. (eds) Evolution of Genes and Proteins, Sinauer, Sunderland, Massachusetts, pp. 147–164.
Avise, J C, Lansman, R A, and Shade, R D. 1979. The Use of restriction endonucleases to measure mitochondrial DNA sequence relatedness in natural populations. I. Population structure and evolution in the genus Peromyscus. Genetics, 92, 279–295.
Ayala, F J, Powell, J R, Tracey, M L, Mourao, C A, and Perezsalas, S. 1972. Enzyme variability in the Drosophila willistoni group. III. Genie variation in natural populations of Drosophila willistoni. Genetics, 70, 113–139.
Ayala, F J, Serra, L, and Prevosti, A. 1989. A grand experiment in evolution: the Drosophila subobscura colonization of the Americas. Drosophila suboscura Genome, 31, 246–255.
Baba-Aissa, F, Sollgnac, M, Dennebouy, N, and David, J R. 1988. Mitochondrial DNA variability in Drosophila simulans: quasi-absence of polymorphism within each of the three cytoplasmic races. Heredity, 61, 419–426.
Birky, C W, Fuerst, P, and Maruyama, T. 1989. Organelle gene diversity under migration, mutation, and drift: equilibrium expectations, approach to equilibrium, effects of heteroplasmic cells, and comparison to nuclear genes. Genetics, 121, 613–627.
Birky, C W, Maruyama, T, and Fuerst, P. 1983. An approach to population and evolutionary genetic theory for genes in mitochondria and chloroplasts, and some results. Genetics, 103, 513–527.
Brewer, G J. 1970. An Introduction to Isozymes Techniques. Academic Press, New York.
Cabrera, V M, Gonzalez, A M, and Gullon, A. 1980. Enzymatic polymorphism in Drosophila subobscura populations from the Canary Islands. Evolution, 34, 875–887.
Clary, D O, and Wolstenholme, D R. 1985. The mitochondrial DNA molecular of Drosophila yakuba: nucleotide sequence, gene organization, and genetic code. J Mol Evol, 22, 252–271.
Crease, T J, Lynch, M, and Spitze, K. 1990. Hierarchical analysis of population genetic variation in mitochondrial and nuclear genes of Daphnia pulex. Mol Biol Evol, 7, 444–458.
Desalle, R, and Hunt, J H. 1987. Molecular evolution in Hawaiian drosophilids. Trends in Ecology and Evolution, 2, 212–216.
Desalle, R, Templeton, A, Mori, I, Pletscher, S, and Johnston, J S. 1987. Temporal and spational heterogeneity of mtDNA polymorphism in natural populations of Drosophila mercatorum. Genetics, 116, 215–223.
Engels, W R. 1981. Estimating genetic divergence and genetic variability with restriction endonucleases. Proc Natl Acad Sci USA, 78, 6329–6333.
Fos, M, Dominguez, M A, Latorre, A, and Moya, A. 1990. Mitochondrial DNA evolution in experimental populations of Drosophila subobscura. Proc Natl Acad Sci USA, 87, 4198–4201.
Hale, L R, and Singh, R S. 1987. Mitochondrial DNA variation and genetic structure in populations of Drosophila melanogaster. Mol Biol Evol, 4, 622–637.
Harrison, R G. 1989. Animal mitochondrial DNA as a genetic marker in population and evolutionary biology. Trends in Ecology and Evolution, 4, 6–11.
Krimbas, C B, and Loukas, M. 1980. The inversion polymorphism of Drosophila suboscura. Evol Biol, 12, 163–234.
Lakovaara, S, and Saura, A. 1971. Genie variation in marginal populations of Drosophila subobscura. Hereditas, 69, 77–82.
Lakovaara, S, and Saura, A. 1973. Genie variation in central and marginal populations of Drosophila subobscura. Hereditas, 75, 33–46.
Latorre, A, Moya, A, and Ayala, F J. 1986. Evolution of mitochondrial DNA in Drosophila subobscura. Proc Natl Acad Sci USA, 83, 8649–8653.
Latorre, A, Barrio, E, Moya, A, and Ayala, F J. 1988. Mitochondrial DNA evolution in the Drosophila obscura group. Mol Biol Evol, 5, 717–728.
Lynch, M, and Crease, T J. 1990. The analysis of population survey data on DNA sequence variation. Mol Biol Evol, 7, 377–394.
Marinkovic, D, Ayala, F J, and Andjelkovic, M. 1978. Genetic polymorphism and phylogeny of Drosophila subobscura. Evolution, 32, 164–173.
Nei, M. 1973. Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA, 70, 3321–3323.
Nei, M. 1987. Molecular Evolutionary Genetics. Columbia University Press, New York.
Nigro, L. 1988. Natural populations of Drosophila simulans show great uniformity of the mitochondrial DNA restriction map. Genetica, 77, 133–137.
Pinsker, W, Lankinen, P, and Sperlich, D. 1978. Allozyme and inversion polymorphism in a central European population of Drosophila subobscura. Genetica, 48, 207–214.
Prevosti, A, De Frutos, R, Alonso, G, Latorre, A, Monclus, M, and Martinez, M J. 1984. Genetic differentiation between natural populations of Drosophila subobscura in the western mediterranean area with respect to chromosomal variation. Genet Sci Evol, 16, 143–156.
Prevosti, A, Ribo, G, Serra, L, Aguade, M, Balaña, J, Monclus, M, and Mestres, F. 1988. Colonization of America by Drosophila subobscura: experiment in natural populations that supports the adaptive role of chromosomal-inversion polymorphism. Proc Natl Acad Sci USA, 85, 5597–5600.
Rand, D M, and Harrison, R G. 1989. Molecular population genetics of mtDNA size variation in crickets. Genetics, 121, 551–569.
Saura, A, Lakovaara, S, Lokki, J, and Lankinen, P. 1973. Genic variation in central and marginal populations of Drosophila subobscura. Hereditas, 75, 33–46.
Singh, R S, and Rhomberg, L R. 1987. A comprehensive study of genic variation in natural populations of Drosophila melanogaster. I. Estimates of gene flow from rare alleles. Genetics, 115, 313–322.
Slatkin, M. 1989. Population structure and evolutionary progress. Genome, 31, 196–202.
Solignac, M, Monnerot, M, and Mounolou, J C. 1986. Mitochondrial DNA evolution in the melanogaster species subgroup of Drosophila. J Mol Evol, 23, 31–40.
Takahata, N, and Palumbi, S. 1985. Extranuclear differentation and gene flow in the finite island model. Genetics, 109, 441–457.
Weir, B S, and Cockerham, C C. 1984. Estimating F-statistics for the analysis of population structure. Evolution, 38, 1358–1370.
Zouros, E, Krimbas, C B, Tsakas, S, and Loukas, M. 1974. Genic versus chromosomal variation in natural populations of Drosophila subobscura. Genetics, 78, 1223–1244.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Latorre, A., Hernández, C., Martínez, D. et al. Population structure and mitochondrial DNA gene flow in Old World populations of Drosophila subobscura. Heredity 68, 15–24 (1992). https://doi.org/10.1038/hdy.1992.2
Received:
Issue Date:
DOI: https://doi.org/10.1038/hdy.1992.2
Keywords
This article is cited by
-
Rate of change for the thermal adapted inversions in Drosophila subobscura
Genetica (2019)
-
Negative frequency dependent selection on sympatric mtDNA haplotypes in Drosophila subobscura
Hereditas (2016)
-
Gene flow and gene flux shape evolutionary patterns of variation in Drosophila subobscura
Heredity (2013)
-
Mitochondrial DNA effects on fitness in Drosophila subobscura
Heredity (2011)
-
Molecular evidence to suggest the origin of a colonization: Drosophila subobscura in America
Genetica (2011)