Abstract
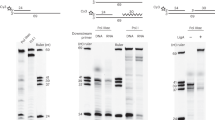
Extensive work on the maturation of lagging strands during the replication of simian virus 40 DNA suggests that the initiator RNA primers of Okazaki fragments are removed by the combined action of two nucleases, RNase HI and Fen1, before the Okazaki fragments join1,2,3,4,5. Despite the well established in vitro roles of these two enzymes6, genetic analyses in yeast revealed that null mutants of RNase HI and/or Fen1 are not lethal7,8,9, suggesting that an additional enzymatic activity may be required for the removal of RNA. One such enzyme is the Saccharomyces cerevisiae Dna2 helicase10,11,12/endonuclease12, which is essential for cell viability13,14 and is well suited to removing RNA primers of Okazaki fragments15. In addition, Dna2 interacts genetically and physically with several proteins involved in the elongation or maturation of Okazaki fragments10,16. Here we show that the endonucleases Dna2 and Fen1 act sequentially to facilitate the complete removal of the primer RNA. The sequential action of these enzymes is governed by a single-stranded DNA-binding protein, replication protein-A (RPA). Our results demonstrate that the processing of Okazaki fragments in eukaryotes differs significantly from, and is more complicated than, that occurring in prokaryotes. We propose a novel biochemical mechanism for the maturation of eukaryotic Okazaki fragments.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Challberg, M. D. & Kelly, T. J. Animal virus DNA replication. Annu. Rev. Biochem. 58, 671–717 (1989).
Hurwitz, J., Dean, F. B., Kwong, A. D. & Lee, S. H. The in vitro replication of DNA containing the SV40 origin. J. Biol. Chem. 265, 18043–18046 (1990).
Waga, S. & Stillman, B. The DNA replication fork in eukaryotic cells. Annu. Rev. Biochem. 67, 721–751 (1998).
Lieber, M. R. The FEN1 family of structure-specific nucleases in eukaryotic DNA replication, recombination and repair. Bioessays 19, 233–240 (1997).
Bambara, R. A., Murante, R. S. & Hendericksen, L. A. Enzymes and reactions at the eukaryotic DNA replication forks. J. Biol. Chem. 272, 4647–4650 (1997).
Waga, S. & Stillman, B. Anatomy of a DNA replication fork revealed by reconstitution of SV40 DNA replication in vitro. Nature 369, 207–212 (1994).
Frank, P., Braunshofer-Reiter, C. & Wintersberger, U. Yeast RNase H(35) is the counterpart of the mammalian RNase HI, and is evolutionarily related to prokaryotic RNase HII. FEBS Lett. 421, 23–26 (1998).
Reagan, M. S., Pittinger, C., Siede, W. & Friedberg, E. C. Characterization of a mutant strain of Saccharomyces cerevisiae with a deletion of the RAD 27 gene, a structural homolog of the RAD2 nucleotide excision repair gene. J. Bacteriol. 177, 364–371 (1995).
Sommers, C. H. et al. Conditional lethality of null mutations in RTH1 that encodes the yeast counterpart of a mammalian 5′- to 3′-exonuclease required for lagging strand DNA synthesis in reconstituted systems. J. Biol. Chem. 270, 4193–4196 (1995).
Budd, M. E. & Campbell, J. A yeast replicative helicase, Dna2 helicase, interacts with yeast FEN1 nucleases in carrying out its essential function. Mol. Cell. Biol. 17, 2136–2142 (1997).
Budd, M. E. & Campbell, J. A yeast gene required for DNA replication encodes a protein with homology to DNA helicases. Proc. Natl Acad. Sci. USA 92, 7642–7646 (1995).
Bae, S. H. et al. Dna2 of Saccharomyces cerevisiae possesses a single-stranded DNA-specific endonuclease activity that is able to act on double-stranded DNA in the presence of ATP. J. Biol. Chem. 273, 26880–26890 (1998).
Lee, K. W. et al. The endonuclease activity of the yeast Dna2 enzyme is essential in vivo. Nucleic Acids Res. 28, 2873–2881 (2000).
Budd, M. E., Choe, W.-C. & Campbell, J. The nuclease activity of the yeast Dna2 protein, which is related to the RecB-like nucleases, is essential in vivo. J. Biol. Chem. 275, 16518–16529 (2000).
Bae, S. H. & Seo, Y. S. Characterization of the enzymatic properties of the yeast Dna2 helicase/endonuclease suggests a new model for Okazaki fragment processing. J. Biol. Chem. 275, 38022–38031 (2000).
Kang, H. Y. et al. Genetic analyses of Schizosaccharomyces pombe dna2+ reveal that Dna2 plays an essential role in Okazaki fragment metabolism. Genetics 155, 1055–1067 (2000).
Kenny, M., Lee, S.-H. & Hurwitz, J. Multiple functions of human single-stranded-DNA binding protein in simian virus 40 DNA replication: single-strand stabilization and stimulation of DNA polymerases α and δ. Proc. Natl Acad. Sci. USA 86, 9757–9761 (1989).
Wold, M. S. Replication protein A: a heterotrimeric, single-stranded DNA binding protein required for eukaryotic DNA replication. Annu. Rev. Biochem. 66, 61–92 (1997).
Iftode, C. & Borowiec, J. A. Denaturation of the simian virus 40 origin of replication mediated by human replication protein A. Mol. Cell. Biol. 17, 3876–3883 (1997).
Budd, M. E., Choe, W.-C. & Campbell, J. DNA2 encodes a DNA helicase essential for replication of eukaryotic chromosomes. J. Biol. Chem. 270, 26766–26769 (1995).
Gomes, X. V. & Burgers, P. M. J. Two modes of FEN1 binding to PCNA regulated by DNA. EMBO J. 19, 3811–3821 (2000).
Mass, G., Nethanel, T. & Kaufmann, G. The middle subunit of replication protein A contacts growing RNA–DNA primers in replicating simian virus 40 chromosomes. Mol. Cell. Biol. 18, 6399–6407 (1998).
Tishkoff, D. X. et al. Identification and characterization of Saccharomyces cerevisiae EXO1, a gene encoding an exonuclease that interacts with MSH2. Proc. Natl Acad. Sci. USA 94, 7487–7492 (1997).
Holmes, A. M. & Haber, J. Double-strand break repair in yeast requires both leading and lagging strand DNA polymerases. Cell 96, 415–424 (1999).
Gary, R. et al. A novel role in DNA metabolism for the binding of Fen1/Rad27 to PCNA and implications for genetic risk. Mol. Cell. Biol. 19, 5373–5382 (1999).
Bae, S. H. & Seo, Y. S. In vitro evidence that purified yeast Rad27 and Dna2 are not stably associated with each other suggests that an additional protein(s) is required for a complex formation. J. Biochem. Mol. Biol. 33, 155–161 (2000).
Brill, S. J. & Stillman, B. Yeast replication factor-A functions in the unwinding of the SV40 origin of DNA replication. Nature 342, 92–95 (1989).
Tomkinson, A. E., Tappe, N. J. & Friedberg, E. C. DNA ligase I from Saccharomyces cerevisiae: physical and biochemical characterization of the CDC9 gene product. Biochemistry 31, 11762–11771 (1992).
Zuo, S. et al. Structure and activity associated with multiple forms of Schizosaccharomyces pombe DNA polymerase δ. J. Biol. Chem. 275, 5153–5162 (2000).
Brosh, R. M. et al. Replication protein A physically interacts with the Bloom's syndrome protein and stimulates its helicase activity. J. Biol. Chem. 275, 23500–23508 (2000).
Acknowledgements
We thank J. Hurwitz, S. MacNeill and A.-K. Bielinsky for critical reading of the manuscript. This work was supported by a grant from the Creative Research Initiatives of the Korean Ministry of Science and Technology given to Y.-S.S.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bae, SH., Bae, KH., Kim, JA. et al. RPA governs endonuclease switching during processing of Okazaki fragments in eukaryotes. Nature 412, 456–461 (2001). https://doi.org/10.1038/35086609
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35086609
This article is cited by
-
PARP inhibition impedes the maturation of nascent DNA strands during DNA replication
Nature Structural & Molecular Biology (2022)
-
New insights into the mechanism of RPA in preserving genome stability
Genome Instability & Disease (2022)
-
The replisome guides nucleosome assembly during DNA replication
Cell & Bioscience (2020)
-
Inhibition of DNA2 nuclease as a therapeutic strategy targeting replication stress in cancer cells
Oncogenesis (2017)
-
Control of structure-specific endonucleases to maintain genome stability
Nature Reviews Molecular Cell Biology (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.