Abstract
SPG4 mutations are the most frequent cause of autosomal-dominant hereditary spastic paraplegia (HSP). SPG4 HSP is characterized by large inter- and intrafamilial variability in age at onset (AAO) and disease severity. The broad spectrum of SPG4 mutations has recently been further extended by the finding of large genomic deletions in SPG4-linked pedigrees negative for ‘small’ mutations. We had previously reported a very large pedigree, linked to the SPG4 locus with many affected members, which showed gender difference in clinical manifestation. Screening for copy number aberrations revealed the first case of a multi-exonic duplication (exon10_12dup) in the SPG4 gene. The mutation leads to a premature stop codon, suggesting that the protein product is not functional. The analysis of 30 individuals who carry the mutation showed that males have on average an earlier AAO and are more severely affected. The present family suggests that this HSP pathogenesis may be modulated by factors related to individual background and gender as observed for other autosomal dominant conditions, such as facio-scapulohumeral muscular dystrophy or amyloidosis. Understanding why some individuals, particularly women, are ‘partially protected’ from the effects of this and other pathogenic mutations is of utmost importance.
Similar content being viewed by others
Introduction
The major feature of the heterogeneous condition hereditary spastic paraplegia (HSP) is progressive weakness and spasticity of the lower limbs. The most frequent cause is mutation of the SPG4 gene, associated with autosomal-dominant inheritance.1 SPG4 HSP shows large inter- and intrafamilial variability in age at onset (AAO) and disease severity, suggesting the existence of modifying factors.2 The broad spectrum of SPG4 mutations has recently been further extended by the finding of large genomic deletions in SPG4-linked pedigrees, which had screened negative for ‘small’ mutations.3 We previously reported on another such SPG4-linked but apparently mutation-negative pedigree, which, in addition, appeared unique in almost exclusively men expressing the phenotype.4
Subjects and methods
DNA of three definitely affected individuals was analyzed using SPG4-specific multiplex ligation-dependent probe amplification (MLPA – MRC Holland) as described previously.5 Long-range PCR on genomic DNA followed by sequencing was used to define the limits of the aberration identified. Mutation-specific PCR was used to establish the carrier status of 52 individuals.
The polymorphic variants c.131C>T or c.134C>A were screened as described previously.6 The samples were sequenced in a Mega Bace 1000 DNA Sequencer – Amersham Bioscience (GE-Healthcare). cDNA to be used in RT-PCR was prepared from peripheral blood of affected family members.
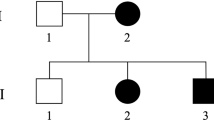
Fifty-two available family members (30 women and 22 men) from three generations were clinically re-investigated and classified by a team of neurologists, who were blind about the genotypes; these included most of the formerly reported4 as well as 14 additional individuals (Figure 1 and Table 1).
Pedigree for the 52 individuals included in this analysis. Symbol below individuals corresponds to molecular and neurological condition. N: no mutation detected; 0: asymptomatic at neurological examination; 1: asymptomatic with mild neurological abnormalities; 2: moderate neurological abnormalities, able to walk without support; 3: severe neurological abnormalities, only able to walk with cane or walker.
Results
Using MLPA, we identified a partial SPG4 duplication (exon10_12dup) in DNA of three definitely affected individuals. Long-range PCR followed by sequencing of the junction revealed that this duplication was in tandem (c.1246-2896_1493+523dup) with non-homology of the sequences contributing to the novel fusion (Supplementary Figure 1). None of the family members carried the polymorphic variants c.131C>T or c.134C>A. RT PCR-based cDNA analysis showed the duplication to also be present at mRNA level. From the total of 52 individuals investigated, 30 (14 women) carried the duplication, 12 of whom (9 women) had no clinical complaints. AAO data compiled during the re-investigation showed a strong difference between men and women (P=0.017, Figure 2).
Discussion
The molecular defect we identified corroborates the recent suggestion of copy number errors to underlie all cases of SPG4 HSP negative for small mutations.3 However, only deletions have been reported so far, whereas we now present the first instance of a partial SPG4 duplication. The lack of sequence identity at the novel junction in our case (Supplementary Figure 1) argues against non-allelic homologous recombination underlying the duplication event. The duplication is in tandem and results in an mRNA containing two adjacent exon 10–12 blocks. This arrangement causes a frameshift and creates a pre-terminal stop codon (PTC) in the second exon 10, thereby disrupting the enzymatic domain. Lack of enzymatic activity has been proposed as the disease-causing mechanism for numerous missense and truncating mutations showing the same primary effect.2, 7
The ascertainment of carrier status and AAO data by the present study allowed statistical confirmation of highly sex-dependent penetrance in our pedigree (Figures 1 and 2 and Table 1). Since the median AAO of SPG4 HSP is approximately 30 years,8 we observed more women than men showing an unusual course of the disease (Figure 2). The presence of known intragenic disease modifiers,6 potentially contributing to the distribution of AAO, was excluded. Further efforts are needed to resolve why a seemingly special mutation leads to the ‘protection’ of a defined group of carriers in this pedigree. Interestingly, a similar gender difference in clinical course has been observed for other autosomal-dominant neuromuscular disorders, such as facioscapulohumeral muscular dystrophy9 and amyloidosis.10 Understanding why some individuals, particularly women, are ‘partially protected’ from the effects of this and other pathogenic mutations remains a great challenge that might open new avenues for treating these disorders.
References
Fink JK : Hereditary spastic paraplegia. Curr Neurol Neurosci Rep 2006; 6: 65–76.
Fonknechten N, Mavel D, Byrne P et al: Spectrum of SPG4 mutations in autosomal dominant spastic paraplegia. Hum Mol Genet 2000; 9: 637–644.
Beetz C, Zuchner S, Ashley-Koch A et al: Linkage to a known gene but no mutation identified: comprehensive re-analysis of SPG4 HSP pedigrees reveals large deletions as the sole cause. Hum Mut 2007; 28: 739–740.
Starling A, Rocco P, Passos-Bueno MR, Hazan J, Marie SK, Zatz M : Autosomal dominant (AD) pure spastic paraplegia (HSP) linked to locus SPG4 affects almost exclusively males in a large pedigree. J Med Genet 2002; 39: e77.
Beetz C, Nygren AO, Schickel J et al: High frequency of partial SPAST deletions in autosomal dominant hereditary spastic paraplegia. Neurology 2006; 67: 1926–1930.
Svenson IK, Kloos MT, Gaskell PC et al: Intragenic modifiers of hereditary spastic paraplegia due to spastin gene mutations. Neurogenetics 2004; 5: 157–164.
Evans KJ, Gomes ER, Reisenweber SM, Gundersen GG : Lauring: Linking axonal degeneration to microtubule remodeling by Spastin-mediated microtubule severing. J Cell Biol 2005; 168: 599–606.
Yip AG, Durr A, Marchuk DA et al: Meta-analysis of age at onset in spastin-associated hereditary spastic paraplegia provides no evidence for a correlation with mutational class. J Med Genet 2003; 40: e106.
Zatz M, Marie SK, Cerqueira A, Vainzof M, Pavanello RC, Passos-Bueno MR : The facioscapulohumeral muscular dystrophy (FSHD1) gene affects males more severely and more frequently than females. Am J Med Genet 1998; 77: 155–161.
Bornebroek M, Westendorp R, Haan J et al: Mortality from hereditary cerebral haemorrhage with amyloidosis – Dutch type. The impact of sex, parental transmission and year of birth. Brain 1997; 120: 2243–2249.
Acknowledgements
We thank the family members for participating in the study. This research was supported by FAPESP-CEPID, CNPq and the Tom-Wahlig-Stiftung.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Supplementary Information accompanies the paper on European Journal of Human Genetics website (http://www.nature.com/ejhg)
Supplementary information
Rights and permissions
About this article
Cite this article
Mitne-Neto, M., Kok, F., Beetz, C. et al. A multi-exonic SPG4 duplication underlies sex-dependent penetrance of hereditary spastic paraplegia in a large Brazilian pedigree. Eur J Hum Genet 15, 1276–1279 (2007). https://doi.org/10.1038/sj.ejhg.5201924
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201924
Keywords
This article is cited by
-
A clinical and genetic study of SPG31 in Japan
Journal of Human Genetics (2022)
-
Genetic, clinical and biochemical characterization of a large cohort of patients with hyaline fibromatosis syndrome
Orphanet Journal of Rare Diseases (2019)
-
Mutational Spectrum of Spast (Spg4) and Atl1 (Spg3a) Genes In Russian Patients With Hereditary Spastic Paraplegia
Scientific Reports (2019)
-
Delving into the complexity of hereditary spastic paraplegias: how unexpected phenotypes and inheritance modes are revolutionizing their nosology
Human Genetics (2015)
-
Where genotype is not predictive of phenotype: towards an understanding of the molecular basis of reduced penetrance in human inherited disease
Human Genetics (2013)