Abstract
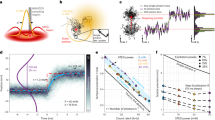
Many biological processes involve enzymes moving along DNA. Such motion might be impeded by DNA-bound proteins or DNA supercoils. Current techniques are incapable of directly measuring forces that such 'roadblocks' might impose. We constructed a setup with four independently moveable optical traps, allowing us to manipulate two DNA molecules held between beads. By tightly wrapping one DNA around the other, we created a probe that can be scanned along the contour of the second DNA. We found that friction between the two polymers remains below 1 pN. Upon encountering DNA-bound proteins substantial friction forces are measured, allowing accurate localization of protein positions. Furthermore, these proteins remained associated at low probe tensions but could be driven off using forces greater than 20 pN. Finally, the full control of the orientation of two DNA molecules opens a wide range of experiments on proteins interacting with multiple DNA regions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
West, S.C. Molecular views of recombination proteins and their control. Nat. Rev. Mol. Cell Biol. 4, 435–445 (2003).
van den Bosch, M., Lohman, P.H. & Pastink, A. DNA double-strand break repair by homologous recombination. Biol. Chem. 383, 873–892 (2002).
van Mameren, J. et al. Dissecting elastic heterogeneity along DNA molecules coated partly with Rad51 using concurrent fluorescence microscopy and optical tweezers. Biophys. J. 91, L78–L80 (2006).
van den Broek, B., Vanzi, F., Normanno, D., Pavone, F.S. & Wuite, G.J. Real-time observation of DNA looping dynamics of Type IIE restriction enzymes NaeI and NarI. Nucleic Acids Res. 34, 167–174 (2006).
Luijsterburg, M.S., Noom, M.C., Wuite, G.J. & Dame, R.T. The architectural role of nucleoid-associated proteins in the organization of bacterial chromatin: A molecular perspective. J. Struct. Biol. 156, 262–272 (2006).
Dame, R.T., Noom, M.C. & Wuite, G.J. Bacterial chromatin organization by H-NS protein unravelled using dual DNA manipulation. Nature 444, 387–390 (2006).
Berg, O.G., Winter, R.B. & von Hippel, P.H. Diffusion-driven mechanisms of protein translocation on nucleic acids. 1. Models and theory. Biochemistry 20, 6929–6948 (1981).
Gowers, D.M. & Halford, S.E. Protein motion from non-specific to specific DNA by three-dimensional routes aided by supercoiling. EMBO J. 22, 1410–1418 (2003).
von Hippel, P.H. & Berg, O.G. Facilitated target location in biological systems. J. Biol. Chem. 264, 675–678 (1989).
Epshtein, V., Toulme, F., Rahmouni, A.R., Borukhov, S. & Nudler, E. Transcription through the roadblocks: the role of RNA polymerase cooperation. EMBO J. 22, 4719–4727 (2003).
Allersma, M.W., Gittes, F., deCastro, M.J., Stewart, R.J. & Schmidt, C.F. Two-dimensional tracking of ncd motility by back focal plane interferometry. Biophys. J. 74, 1074–1085 (1998).
Gittes, F. & Schmidt, C.F. Interference model for back-focal-plane displacement detection in optical tweezers. Opt. Lett. 23, 7–9 (1998).
Brewer, L.R., Corzett, M. & Balhorn, R. Protamine-induced condensation and decondensation of the same DNA molecule. Science 286, 120–123 (1999).
Arai, Y. et al. Tying a molecular knot with optical tweezers. Nature 399, 446–448 (1999).
Vermeulen, K.C. et al. Calibrating bead displacements in optical tweezers using acousto-optic deflectors. Rev. Sci. Instrum. 77, 013704 (2006).
Gosule, L.C. & Schellman, J.A. Compact form of DNA induced by spermidine. Nature 259, 333–335 (1976).
Shimamoto, N. One-dimensional diffusion of proteins along DNA - Its biological and chemical significance revealed by single-molecule measurements. J. Biol. Chem. 274, 15293–15296 (1999).
Trabesinger, W., Schutz, G.J., Gruber, H.J., Schindler, H. & Schmidt, T. Detection of individual oligonucleotide pairing by single-molecule microscopy. Anal. Chem. 71, 279–283 (1999).
Hiller, D.A. et al. Simultaneous DNA binding and bending by EcoRV endonuclease observed by real-time fluorescence. Biochemistry 42, 14375–14385 (2003).
Charvin, G., Vologodskii, A., Bensimon, D. & Croquette, V. Braiding DNA: experiments, simulations, and models. Biophys. J. 88, 4124–4136 (2005).
Binnig, G., Quate, C.F. & Gerber, C. Atomic force microscope. Phys. Rev. Lett. 56, 930–933 (1986).
Lindsay, S.M. et al. STM and AFM images of nucleosome DNA under water. J. Biomol. Struct. Dyn. 7, 279–287 (1989).
Fritzsche, W., Takac, L. & Henderson, E. Application of atomic force microscopy to visualization of DNA, chromatin, and chromosomes. Crit. Rev. Eukaryot. Gene Expr. 7, 231–240 (1997).
Kasas, S. et al. Escherichia coli RNA polymerase activity observed using atomic force microscopy. Biochemistry 36, 461–468 (1997).
van Noort, S.J. et al. Direct visualization of dynamic protein-DNA interactions with a dedicated atomic force microscope. Biophys. J. 74, 2840–2849 (1998).
Huisstede, J.H.G., van der Werf, K.O., Bennink, M.L. & Subramaniam, V. Force detection in optical tweezers using backscattered light. Opt. Express 13, 1113–1123 (2005).
Koch, S.J., Shundrovsky, A., Jantzen, B.C. & Wang, M.D. Probing protein-DNA interactions by unzipping a single DNA double helix. Biophys. J. 83, 1098–1105 (2002).
Kowalczykowski, S.C., Dixon, D.A., Eggleston, A.K., Lauder, S.D. & Rehrauer, W.M. Biochemistry of homologous recombination in Escherichia coli. Microbiol. Rev. 58, 401–465 (1994).
Dame, R.T., Wyman, C. & Goosen, N. Structural basis for preferential binding of H-NS to curved DNA. Biochimie 83, 231–234 (2001).
Halford, S.E., Welsh, A.J. & Szczelkun, M.D. Enzyme-mediated DNA looping. Annu. Rev. Biophys. Biomol. Struct. 33, 1–24 (2004).
van den Broek, B., Noom, M.C. & Wuite, G.J.L. DNA-tension dependence of restriction enzyme activity reveals mechanochemical properties of the reaction pathway. Nucleic Acids Res. 33, 2676–2684 (2005).
Svoboda, K. & Block, S.M. Biological applications of optical forces. Annu. Rev. Biophys. Biomol. Struct. 23, 247–285 (1994).
Wuite, G.J.L., Davenport, R.J., Rappaport, A. & Bustamante, C. An integrated laser trap/flow control video microscope for the study of single biomolecules. Biophys. J. 79, 1155–1167 (2000).
Gittes, F. & Schmidt, C.F. Signals and noise in micromechanical measurements. Methods Cell Biol. 55, 129–156 (1998).
Acknowledgements
We thank E.J.G. Peterman and R.T. Dame for useful discussions. We acknowledge D.A. Hiller (Yale) and J.J. Perona (University of California Santa Barbara) for the kind gift of EcoRV. This work is part of the research program of the 'Stichting voor Fundamenteel Onderzoek der Materie (FOM)', which is financially supported by the 'Nederlandse Organisatie voor Wetenschappelijk Onderzoek (NWO)' and was supported by a NWO-Vernieuwingsimpuls grant.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3, Supplementary Note (PDF 185 kb)
Supplementary Movie 1
Trapping beads, catching two DNA molecules and wrapping of the probing DNA around the scanned DNA. The movie (sped up 3×) shows how two optically trapped DNA molecules are wound around each other. First, beads are captured in the four optical traps. Next, two DNA molecules are caught between these beads. The displacements of the beads out of their traps (indicated by crosshairs) are a signal that DNA is tethered between them. By moving one of the traps in the direction perpendicular to the field of view, the DNA molecules are then wound around each other. Finally, the DNA molecules are brought into a crossed configuration, required for the DNA scanning experiments. (MOV 5015 kb)
Rights and permissions
About this article
Cite this article
Noom, M., van den Broek, B., van Mameren, J. et al. Visualizing single DNA-bound proteins using DNA as a scanning probe. Nat Methods 4, 1031–1036 (2007). https://doi.org/10.1038/nmeth1126
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth1126
This article is cited by
-
PICH acts as a force-dependent nucleosome remodeler
Nature Communications (2022)
-
Additive manufacturing of laminar flow cells for single-molecule experiments
Scientific Reports (2019)
-
A Colorimetric and Fluorescent Probe Based on Rhodamine B for Detection of Fe3+ and Cu2+ Ions
Journal of Fluorescence (2019)
-
Directly interrogating single quantum dot labelled UvrA2 molecules on DNA tightropes using an optically trapped nanoprobe
Scientific Reports (2015)
-
STED nanoscopy combined with optical tweezers reveals protein dynamics on densely covered DNA
Nature Methods (2013)