Abstract
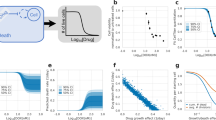
Large-scale analysis of cellular response to anticancer drugs typically focuses on variation in potency (half-maximum inhibitory concentration, (IC50)), assuming that it is the most important difference between effective and ineffective drugs or sensitive and resistant cells. We took a multiparametric approach involving analysis of the slope of the dose-response curve, the area under the curve and the maximum effect (Emax). We found that some of these parameters vary systematically with cell line and others with drug class. For cell-cycle inhibitors, Emax often but not always correlated with cell proliferation rate. For drugs targeting the Akt/PI3K/mTOR pathway, dose-response curves were unusually shallow. Classical pharmacology has no ready explanation for this phenomenon, but single-cell analysis showed that it correlated with significant and heritable cell-to-cell variability in the extent of target inhibition. We conclude that parameters other than potency should be considered in the comparative analysis of drug response, particularly at clinically relevant concentrations near and above the IC50.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Borden, E.C. & Dowlati, A. Phase I trials of targeted anticancer drugs: a need to refocus. Nat. Rev. Drug Discov. 11, 889–890 (2012).
Sos, M.L. et al. Predicting drug susceptibility of non–small cell lung cancers based on genetic lesions. J. Clin. Invest. 119, 1727–1740 (2009).
Heiser, L.M. et al. Subtype and pathway specific responses to anticancer compounds in breast cancer. Proc. Natl. Acad. Sci. USA 109, 2724–2729 (2012).
Garnett, M.J. et al. Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature 483, 570–575 (2012).
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012).
Greshock, J. et al. Molecular target class is predictive of in vitro response profile. Cancer Res. 70, 3677–3686 (2010).
Solit, D.B. et al. BRAF mutation predicts sensitivity to MEK inhibition. Nature 439, 358–362 (2006).
Staunton, J.E. et al. Chemosensitivity prediction by transcriptional profiling. Proc. Natl. Acad. Sci. USA 98, 10787–10792 (2001).
Tyson, D.R., Garbett, S.P., Frick, P.L. & Quaranta, V. Fractional proliferation: a method to deconvolve cell population dynamics from single-cell data. Nat. Methods 9, 923–928 (2012).
Hill, A. The possible effects of the aggregation of the molecules of haemoglobin on its dissociation curves. J. Physiol. (Lond.) 40, iv–vii (1910).
Chou, T.C. Derivation and properties of Michaelis-Menten type and Hill type equations for reference ligands. J. Theor. Biol. 59, 253–276 (1976).
Holford, N.H. & Sheiner, L.B. Understanding the dose-effect relationship: clinical application of pharmacokinetic-pharmacodynamic models. Clin. Pharmacokinet. 6, 429–453 (1981).
Shoemaker, R.H. The NCI60 human tumour cell line anticancer drug screen. Nat. Rev. Cancer 6, 813–823 (2006).
Hannah, R., Beck, M. & Moravec, R. CellTiter-Glo™ Luminescent cell viability assay: a sensitive and rapid method for determining cell viability. Promega Cell Notes 2, 11–13 (2001).
Cover, T.M. & Thomas, J.A. Elements of Information Theory (ed. Schilling, D.L.) (John Wiley and Sons, New York, 1991).
Berenbaum, M.C. In vivo determination of the fractional kill of human tumor cells by chemotherapeutic agents. Cancer Chemother. Rep. 56, 563–571 (1972).
Mitchison, T.J. The proliferation rate paradox in antimitotic chemotherapy. Mol. Biol. Cell 23, 1–6 (2012).
Glotzer, M., Murray, A.W. & Kirschner, M.W. Cyclin is degraded by the ubiquitin pathway. Nature 349, 132–138 (1991).
Aligue, R., Akhavan-Niak, H. & Russell, P. A role for Hsp90 in cell cycle control: Wee1 tyrosine kinase activity requires interaction with Hsp90. EMBO J. 13, 6099–6106 (1994).
Bazzaro, M. et al. Ubiquitin-proteasome system stress sensitizes ovarian cancer to proteasome inhibitor-induced apoptosis. Cancer Res. 66, 3754–3763 (2006).
Shen, L. et al. Dose-response curve slope sets class-specific limits on inhibitory potential of anti-HIV drugs. Nat. Med. 14, 762–766 (2008).
Weiss, J.N. The Hill equation revisited: uses and misuses. FASEB J. 11, 835–841 (1997).
Dancey, J. mTOR signaling and drug development in cancer. Nat. Rev. Clin. Oncol. 7, 209–219 (2010).
Hsieh, A.C. et al. Genetic dissection of the oncogenic mTOR pathway reveals druggable addiction to translational control via 4EBP-eIF4E. Cancer Cell 17, 249–261 (2010).
Feldman, M.E. et al. Active-site inhibitors of mTOR target rapamycin-resistant outputs of mTORC1 and mTORC2. PLoS Biol. 7, e38 (2009).
Sherr, C.J. Cancer cell cycles. Science 274, 1672–1677 (1996).
Keiser, M.J. et al. Predicting new molecular targets for known drugs. Nature 462, 175–181 (2009).
Lounkine, E. et al. Large-scale prediction and testing of drug activity on side-effect targets. Nature 486, 361–367 (2012).
Gaudet, S., Spencer, S.L., Chen, W.W. & Sorger, P.K. Exploring the contextual sensitivity of factors that determine cell-to-cell variability in receptor-mediated apoptosis. PLOS Comput. Biol. 8, e1002482 (2012).
Spencer, S.L., Gaudet, S., Albeck, J.G., Burke, J.M. & Sorger, P.K. Non-genetic origins of cell-to-cell variability in TRAIL-induced apoptosis. Nature 459, 428–432 (2009).
Feinerman, O., Veiga, J., Dorfman, J.R., Germain, R.N. & Altan-Bonnet, G. Variability and robustness in T cell activation from regulated heterogeneity in protein levels. Science 321, 1081–1084 (2008).
Cohen, A.A. et al. Dynamic proteomics of individual cancer cells in response to a drug. Science 322, 1511–1516 (2008).
Sigal, A. et al. Variability and memory of protein levels in human cells. Nature 444, 643–646 (2006).
Sampah, M.E., Shen, L., Jilek, B.L. & Siliciano, R.F. Dose-response curve slope is a missing dimension in the analysis of HIV-1 drug resistance. Proc. Natl. Acad. Sci. USA 108, 7613–7618 (2011).
Tourassi, G.D., Frederick, E.D., Markey, M.K. & Floyd, C.E. Jr. Application of the mutual information criterion for feature selection in computer-aided diagnosis. Med. Phys. 28, 2394–2402 (2001).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Series B Stat. Methodol. 57, 289–300 (1995).
Holm, S. A simple sequentially rejective multiple test procedure. Scand. J. Stat. 6, 65–70 (1979).
Jolliffe, I.T. Principal Component Analysis (Springer-Verlag, Berlin, 2002).
Millard, B.L., Niepel, M., Menden, M.P., Muhlich, J.L. & Sorger, P.K. Adaptive informatics for multifactorial and high-content biological data. Nat. Methods 8, 487–493 (2011).
Acknowledgements
We thank W. Chen, G. Berriz, M. Niepel, M. Hafner, D. Flusberg, T. Mitchison, D. Marks and C. Shamu for help. This work was supported by the US National Institutes of Health–Library of Integrated Network-Based Cellular Signatures Program grant HG006097 to P.K.S. and by Stand Up to Cancer grant AACR-SU2C-DT0409 to P.K.S. and J.W.G. M.F.-S. is supported by a Merck Fellowship of the Life Sciences Research Foundation.
Author information
Authors and Affiliations
Contributions
M.F.-S. designed and performed the experiments, analyzed the experimental data, performed statistical analyses and wrote the manuscript. S.H. designed and performed the experiments and wrote the manuscript. L.M.H. designed and performed the experiments and wrote the manuscript. J.W.G. designed the experiments and wrote the manuscript. P.K.S. designed the experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results,Supplementary Tables 1–3 and Supplementary Figures 1–10 (PDF 15121 kb)
Supplementary_DataSet1
Supplementary Data Set 1 (XLSX 440 kb)
Supplementary_DataSet2
Supplementary Data Set 2 (XLS 277 kb)
Supplementary_DataSet3
Supplementary Data Set 3 (XLS 44 kb)
Supplementary_DataSet4
Supplementary Data Set 4 (XLS 37 kb)
Rights and permissions
About this article
Cite this article
Fallahi-Sichani, M., Honarnejad, S., Heiser, L. et al. Metrics other than potency reveal systematic variation in responses to cancer drugs. Nat Chem Biol 9, 708–714 (2013). https://doi.org/10.1038/nchembio.1337
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1337