Abstract
Lysosomes respond to environmental cues by controlling their own biogenesis, but the underlying mechanisms are poorly understood. Here we describe a protein kinase C (PKC)-dependent and mTORC1-independent mechanism for regulating lysosome biogenesis, which provides insights into previously reported effects of PKC on lysosomes. By identifying lysosome-inducing compounds we show that PKC couples activation of the TFEB transcription factor with inactivation of the ZKSCAN3 transcriptional repressor through two parallel signalling cascades. Activated PKC inactivates GSK3β, leading to reduced phosphorylation, nuclear translocation and activation of TFEB, while PKC activates JNK and p38 MAPK, which phosphorylate ZKSCAN3, leading to its inactivation by translocation out of the nucleus. PKC activation may therefore mediate lysosomal adaptation to many extracellular cues. PKC activators facilitate clearance of aggregated proteins and lipid droplets in cell models and ameliorate amyloid β plaque formation in APP/PS1 mouse brains. Thus, PKC activators are viable treatment options for lysosome-related disorders.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Luzio, J. P., Pryor, P. R. & Bright, N. A. Lysosomes: fusion and function. Nat. Rev. Mol. Cell Biol. 8, 622–632 (2007).
Xu, H. & Ren, D. Lysosomal physiology. Annu. Rev. Physiol. 77, 57–80 (2015).
Settembre, C., Fraldi, A., Medina, D. L. & Ballabio, A. Signals from the lysosome: a control centre for cellular clearance and energy metabolism. Nat. Rev. Mol. Cell Biol. 14, 283–296 (2013).
Ferguson, S. M. Beyond indigestion: emerging roles for lysosome-based signaling in human disease. Curr. Opin. Cell Biol. 35, 59–68 (2015).
Efeyan, A., Comb, W. C. & Sabatini, D. M. Nutrient-sensing mechanisms and pathways. Nature 517, 302–310 (2015).
Medina, D. L. et al. Transcriptional activation of lysosomal exocytosis promotes cellular clearance. Dev. Cell 21, 421–430 (2011).
Spampanato, C. et al. Transcription factor EB (TFEB) is a new therapeutic target for Pompe disease. EMBO Mol. Med. 5, 691–706 (2013).
Martina, J. A. et al. The nutrient-responsive transcription factor TFE3 promotes autophagy, lysosomal biogenesis, and clearance of cellular debris. Sci. Signal. 7, ra9 (2014).
Polito, V. A. et al. Selective clearance of aberrant tau proteins and rescue of neurotoxicity by transcription factor EB. EMBO Mol. Med. 6, 1142–1160 (2014).
Decressac, M. et al. TFEB-mediated autophagy rescues midbrain dopamine neurons from α-synuclein toxicity. Proc. Natl Acad. Sci. USA 110, E1817–E1826 (2013).
Tsunemi, T. et al. PGC-1α rescues Huntington’s disease proteotoxicity by preventing oxidative stress and promoting TFEB function. Sci. Transl. Med. 4, 142ra197 (2012).
Settembre, C. et al. TFEB links autophagy to lysosomal biogenesis. Science 332, 1429–1433 (2011).
Sardiello, M. et al. A gene network regulating lysosomal biogenesis and function. Science 325, 473–477 (2009).
Settembre, C. et al. A lysosome-to-nucleus signalling mechanism senses and regulates the lysosome via mTOR and TFEB. EMBO J. 31, 1095–1108 (2012).
Martina, J. A. & Puertollano, R. Rag GTPases mediate amino acid-dependent recruitment of TFEB and MITF to lysosomes. J. Cell Biol. 200, 475–491 (2013).
Roczniak-Ferguson, A. et al. The transcription factor TFEB links mTORC1 signaling to transcriptional control of lysosome homeostasis. Sci. Signal. 5, ra42 (2012).
Martina, J. A., Chen, Y., Gucek, M. & Puertollano, R. MTORC1 functions as a transcriptional regulator of autophagy by preventing nuclear transport of TFEB. Autophagy 8, 903–914 (2012).
Settembre, C. & Ballabio, A. Lysosomal adaptation: how the lysosome responds to external cues. Cold Spring Harb. Perspect. Biol. 6, a016907 (2014).
Medina, D. L. et al. Lysosomal calcium signalling regulates autophagy through calcineurin and TFEB. Nat. Cell Biol. 17, 288–299 (2015).
Wang, W. et al. Up-regulation of lysosomal TRPML1 channels is essential for lysosomal adaptation to nutrient starvation. Proc. Natl Acad. Sci. USA 112, E1373–E1381 (2015).
Chauhan, S. et al. ZKSCAN3 is a master transcriptional repressor of autophagy. Mol. Cell 50, 16–28 (2013).
Gotta, H., Adolf, W., Opferkuch, H. J. & Hecker, E. On the active principles of the Euphorbiaceae, IXa ingenane type diterpene esters from five Euphorbia species. Z. Naturforsch. B 39, 683–694 (1984).
Chen, C. S., Chen, W. N., Zhou, M., Arttamangkul, S. & Haugland, R. P. Probing the cathepsin D using a BODIPY FL-pepstatin A: applications in fluorescence polarization and microscopy. J. Biochem. Biophys. Methods 42, 137–151 (2000).
Thoreen, C. C. et al. An ATP-competitive mammalian target of rapamycin inhibitor reveals rapamycin-resistant functions of mTORC1. J. Biol. Chem. 284, 8023–8032 (2009).
Hasler, C. M., Acs, G. & Blumberg, P. M. Specific binding to protein kinase C by ingenol and its induction of biological responses. Cancer Res. 52, 202–208 (1992).
Hampson, P. et al. PEP005, a selective small-molecule activator of protein kinase C, has potent antileukemic activity mediated via the delta isoform of PKC. Blood 106, 1362–1368 (2005).
Griner, E. M. & Kazanietz, M. G. Protein kinase C and other diacylglycerol effectors in cancer. Nat. Rev. Cancer 7, 281–294 (2007).
Steinberg, S. F. Distinctive activation mechanisms and functions for protein kinase C delta. Biochem. J. 384, 449–459 (2004).
Zeidan, Y. H. & Hannun, Y. A. Activation of acid sphingomyelinase by protein kinase C delta-mediated phosphorylation. J. Biol. Chem. 282, 11549–11561 (2007).
Balakumar, P. & Jagadeesh, G. A century old renin-angiotensin system still grows with endless possibilities: AT1 receptor signaling cascades in cardiovascular physiopathology. Cell. Signal. 26, 2147–2160 (2014).
Loegering, D. J. & Lennartz, M. R. Protein kinase C and toll-like receptor signaling. Enzyme Res. 2011, 537821 (2011).
Ferron, M. et al. A RANKL-PKCβ-TFEB signaling cascade is necessary for lysosomal biogenesis in osteoclasts. Genes Dev. 27, 955–969 (2013).
Shi, Y. Serine/threonine phosphatases: mechanism through structure. Cell 139, 468–484 (2009).
Doble, B. W. & Woodgett, J. R. GSK-3: tricks of the trade for a multi-tasking kinase. J. Cell Sci. 116, 1175–1186 (2003).
Parr, C. et al. Glycogen synthase kinase 3 inhibition promotes lysosomal biogenesis and autophagic degradation of the amyloid-β precursor protein. Mol. Cell. Biol. 32, 4410–4418 (2012).
Marchand, B., Arsenault, D., Raymond-Fleury, A., Boisvert, F. M. & Boucher, M. J. Glycogen synthase kinase-3 (GSK3) inhibition induces prosurvival autophagic signals in human pancreatic cancer cells. J. Biol. Chem. 290, 5592–5605 (2015).
Stretton, C. et al. GSK3-mediated raptor phosphorylation supports amino-acid-dependent mTORC1-directed signalling. Biochem. J. 470, 207–221 (2015).
Taelman, V. F. et al. Wnt signaling requires sequestration of glycogen synthase kinase 3 inside multivesicular endosomes. Cell 143, 1136–1148 (2010).
Xue, Y. et al. GPS 2.1: enhanced prediction of kinase-specific phosphorylation sites with an algorithm of motif length selection. Protein Eng. Des. Sel. 24, 255–260 (2011).
Larocca, M. C., Ochoa, E. J., Rodriguez Garay, E. A. & Marinelli, R. A. Protein kinase C-dependent inhibition of the lysosomal degradation of endocytosed proteins in rat hepatocytes. Cell. Signal. 14, 641–647 (2002).
Jankowsky, J. L. et al. Mutant presenilins specifically elevate the levels of the 42 residue β-amyloid peptide in vivo: evidence for augmentation of a 42-specific gamma secretase. Hum. Mol. Genet. 13, 159–170 (2004).
Rozengurt, E. Mitogenic signaling pathways induced by G protein-coupled receptors. J. Cell. Physiol. 213, 589–602 (2007).
Naor, Z., Benard, O. & Seger, R. Activation of MAPK cascades by G-protein-coupled receptors: the case of gonadotropin-releasing hormone receptor. Trends Endocrinol. Metab. 11, 91–99 (2000).
Shahnazari, S. et al. A diacylglycerol-dependent signaling pathway contributes to regulation of antibacterial autophagy. Cell Host Microbe 8, 137–146 (2010).
Samie, M. & Cresswell, P. The transcription factor TFEB acts as a molecular switch that regulates exogenous antigen-presentation pathways. Nat. Immunol. 16, 729–736 (2015).
Majewski, M. et al. Protein kinase C delta stimulates antigen presentation by Class II MHC in murine dendritic cells. Int. Immunol. 19, 719–732 (2007).
Liu, D. et al. Rapid biogenesis and sensitization of secretory lysosomes in NK cells mediated by target-cell recognition. Proc. Natl Acad. Sci. USA 102, 123–127 (2005).
Hirai, S. et al. Ras-dependent signal transduction is indispensable but not sufficient for the activation of AP1/Jun by PKCδ. EMBO J. 13, 2331–2340 (1994).
Wu-Zhang, A. X. & Newton, A. C. Protein kinase C pharmacology: refining the toolbox. Biochem. J. 452, 195–209 (2013).
Guo, W. et al. RNA-binding protein FXR2 regulates adult hippocampal neurogenesis by reducing Noggin expression. Neuron 70, 924–938 (2011).
Acknowledgements
We thank X. C. Wang (Institute of Biophysics, China), S. M. Ferguson (Yale University, USA), S. C. Lin (Xiamen University, China), L. Yu (School of Life Sciences, Tsinghua University, China) and Z. H. Xu (Institute of Genetics and Developmental Biology, China) for experimental materials; A. Ballabio, X. C. Wang and J. R. Speakman for constructive suggestions and critical reading of the manuscript; and I. Hanson for proofreading services. This research was supported by grants 31230043 (to C.Yang) and 21432010 (to X.Hao and C.Yang) from the National Science Foundation of China, 2013CB910102 (to C.Yang) from the National Basic Research Program of China, and KJZD-EW-L08 (to C.Yang) from the Chinese Academy of Sciences.
Author information
Authors and Affiliations
Contributions
C.Yang, X.Hao, Y.L. and M.X. conceived the study. Y.L. and M.X. did most of the experiments. X.D. and G.T. contributed to compound screening. C.Yan, Z.S. and Y.D. purified and modified compounds. X.Huang and Y.W. performed mass spectrometry and analysis. T.L. performed flow cytometry. L.C., X.W. and L.M. performed immunoelectron microscopy. S.M., Y.J., X.L. and K.L. contributed materials. W.G., C.T. and Y.L. performed all animal experiments and analysed results. C.Yang, Y.L., M.X. and X.Hao wrote the manuscript with feedback from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 HEP14 induces TFEB nuclear translocation and lysosome biogenesis in multiple cell types.
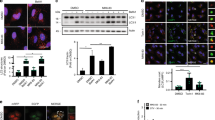
(a) Images (left) and quantifications (right) of HeLa cells treated with HEP14 (20 μM, 3 h) and co-stained with BODIPY-pepstatin A (1 μM) and LysoTracker Red (0.3 μM). n = 3 independent experiments. (b) Images (left) and quantification (right) of immunostaining of endogenous LAMP1 and cathepsin B in HeLa cells treated with HEP14 (20 μM, 3 h). n = 3 independent experiments. (c) Immunoblotting of LAMP1 (upper) and cathepsin B (lower) in HeLa cells treated with HEP14 (0, 10, 20, 40 μM, 3 h). Fold induction of proteins was normalized by LAMP1/α-tubulin or cathepsin B/α-tubulin ratios. Data are from one experiment that is representative of 3 independent experiments. (d) Immuno-electron microscopy images of lysosomes labeled with LAMP1 (arrows) in HeLa cells treated without or with HEP14 (20 μM, 3 h). Insets show magnified (2.27x) views of lysosomes indicated by red arrows. Lysosome quantification is shown at the right. 120 and 150 sections were analyzed for control and HEP14 treatment, respectively. n = 3 independent experiments. (e) Images (left) and quantifications (right) of LysoTracker Red staining in the indicated cell types treated with HEP14 (20 μM, 3 h). n = 3 independent experiments. (f) Images (left) and quantification (right) of immunostaining of endogenous LAMP1 in HeLa cells treated with HEP15 (20 μM, 3 h). n = 3 independent experiments. (g) Images (left) and quantification (right) of the subcellular locations TFEB-EGFP or mCherry-TFE3 in HeLa cells treated with HEP15 (20 μM, 3 h). n = 3 independent experiments. (h and i) Images (left) and quantifications (right) of endogenous TFEB or TFE3 in SH-SY5Y neuroblastoma (h) and HepG2 hepatocarcinoma (i) cells treated for 3 h with HEP14 (20 μM) or Torin1 (1 μM). n = 3 independent experiments. Bars represent 10 μm in all images except (d). Data (mean ± s.e.m) were compared using t-tests or ANOVA ∗∗p < 0.01, ∗∗∗p < 0.001.
Supplementary Figure 2 Requirement for PKC isoforms by HEP14-induced TFEB translocation and characterization of the interaction of HEP14 with PKCα and PKCδ.
(a) Structural comparison of ingenol and ingenol mebutate with HEP14 and HEP15. (b) siRNA knockdown effects of individual PKC isoforms. qRT-PCRs were performed on mRNA from HeLa cells 48 h after siRNA treatment of individual PKC isoforms. n = 3 independent experiments. (c) Quantification of HEP14 (20 μM, 3 h)- or Torin1 (1 μM, 3 h)-induced nuclear translocation of TFEB-EGFP in stable TFEB-EGFP-expressing cells treated with siRNA specific to individual PKC isoforms. For quantifications in (b and c), data (mean ± s.e.m.) were from n = 3 independent experiments and analyzed with ANOVA. ∗∗p < 0.01, ∗∗∗p < 0.001. (d) Purified MBP, MBP-PKCα-C1 and MBP-PKCδ-C1 used for MST assays. (e and f) HEP14 binds to MBP-PKCα-C1 (e) or MBP-PKCδ-C1 (f) in MST assays. Data (mean ± r.m.s.e, root mean square error) were from n = 3 independent experiments. (g) Purified GST, GST-PKCα and GST-PKCδ used for MST assays. (h and i) HEP14 binds to GST-PKCα (h) and GST-PKCδ (i) in MST assays. Data (mean ± r.m.s.e.) were from n = 3 independent experiments.
Supplementary Figure 3 Analysis of PKC-dependent TFEB activation and lysosome biogenesis by DAG, AngII and LPS and the requirement for protein phosphatases.
(a–c) Images (left) and quantifications (right) of nuclear translocation of endogenous TFEB induced by DAG (D-1-palmitoyl-2-oleoyl diglycerol, 100 μM, 3 h), AngII (10 μM, 6 h), or LPS (100 ng/mL, 3 h) in the absence or presence of Bis1 (2 μM) in different cell types. n = 3 independent experiments. (d) Quantification of lysosomes induced by DAG, AngII, or LPS in the absence or presence of Bis1 in different cell types. n = 3 independent experiments. (e) Images (left) and quantifications (right) of subcellular localizations of EGFP-tagged TFEB(S462A/S463A/S466A/S467A/S469A, 5S-A) in HepG2 cells treated with HEP14 (20 μM, 3 h), PMA (1 μM, 3 h), AngII (10 μM, 6 h), or LPS (100 ng/mL, 3 h). n = 3 independent experiments. (f) siRNA effects of the PPP1CA subunit of PP1 (siPPP1CA), the PPP2CA and PPP2CB subunits of PP2A (siPPP2CA, siPPP2CB), the PPP3CB and PPP3R1 subunits of PP3/calcineurin (siPPP3CB and siPPP3R1), the PPP4C subunit of PP4 (siPPP4C), the PPP5C subunit of PP5 (siPPP5C) and MCOLN1 (siMCOLN1). n = 3 independent siRNA experiments. (g) Images of HEP14 (20 μM, 3 h)-induced nuclear translocation of TFEB-EGFP in stable TFEB-EGFP-expressing HeLa cells pretreated with siRNA against PPP1CA, PPP2CA, PPP2CB, PPP3CB, PPP3R1, PPP4C, PPP5C or MCOLN1. (h) Quantification of HEP14 (20 μM, 3 h)-induced nuclear translocation of TFEB-EGFP in stable TFEB-EGFP-expressing HeLa cells pretreated with siRNA against PPP1CA, PPP2CA, PPP2CB, PPP4C and PPP5C. n = 3 independent experiments. (i) Quantification of HEP14 (20 μM, 3 h)- or Torin1 (1 μM, 3 h)-induced nuclear translocation of TFEB-EGFP in stable TFEB-EGFP-expressing HeLa cells pretreated with siRNA against PPP3CB, PPP3R1 or MCOLN1. n = 3 independent experiments. Bars represent 10 μm in all images. For all quantifications, data (mean ± s.e.m) were compared using ANOVA. ∗p < 0.05, ∗∗p < 0.01, ∗∗∗ p < 0.001. NS, not significant.
Supplementary Figure 4 Mass-spectrometry analysis of the phosphorylation of TFEB(105-300) by GSK3β and ZKSCAN3(40-200) by JNK2.
(a) Purified GST-GSK3β (indicated with ∗) used for in vitro phosphorylation assays. (b and c) Mass-spectrometry analysis of the phosphorylation at Ser134 and Ser138 of TFEB(105-300) without (b) or with (c) GST-GSK3β. Peptides containing Ser134 and Ser138 with no phosphorylation (b) and with potential phosphorylation (ph) (c) are indicated. The data are from 2 independent experiments. (d) Purified Flag-MKK7-JNK2 (activated JNK2) (indicated with ∗). (e and f) Mass-spectrometry analysis of the phosphorylation at Thr153 of ZKSCAN3(40-200) without (e) or with (f) Flag-MKK7-JNK2. Peptides containing Thr153 with no phosphorylation (e) and with potential phosphorylation (ph) (f) are shown. The data are from 2 independent experiments.
Supplementary Figure 5 HEP14 induces ZKSCAN3 cytoplasmic translocation through JNK2 and p38, but not GSK3β.
(a) The indicated cell types were transfected with EGFP-ZKSCAN3-expressing vector. 24 h later, cells were treated with HEP14 (20 μM) for 3 h and examined for cytoplasmic translocation of EGFP-ZKSCAN3 by confocal microscopy. n = 3 independent experiments. (b) Images (left) and quantification (right) of HEP14 (20 μM, 3 h)-induced EGFP-ZKSCAN3 cytoplasmic localization in HeLa cells pretreated with GSK3α or GSK3β siRNA. n = 3 independent experiments. (c) Images (left) and quantification (right) of HeLa cells transfected with constructs expressing TFEB(S134A/S138A)-EGFP and mCh-ZKSCAN3 and treated without or with HEP14 (20 μM, 3 h). n = 3 independent experiments. (d) siRNA of JNK1, JNK2, or JNK3 specifically knocks down the corresponding genes in HeLa cells. n = 3 independent experiments. (e) siRNA of p38α, p38β, p38γ or p38δ specifically knocks down the corresponding genes in HeLa cells. n = 3 independent experiments. (f) Images (left) and quantifications (right) of EGFP-ZKSCAN3 in cells treated with the indicated siRNAs and treated without or with HEP14 (20 μM, 3 h). n = 3 independent experiments. (g) Images (left) and quantification (right) of TFEB-EGFP-expressing HeLa cells transfected with an mCh-ZKSCAN3-expressing construct alone or together with a JNK2(CA)-expressing construct. n = 3 independent experiments. Bars represent 10 μm in all images. For all quantifications, data (mean ± s.e.m) were from the indicated number of independent experiments and compared using ANOVA. ∗∗p < 0.01, ∗∗∗ p < 0.001. NS, not significant.
Supplementary Figure 6 HEP14 promotes clearance of HTT polyQ aggregates and lipid droplets.
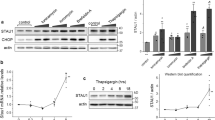
(a) HeLa cells stably expressing Tet-on Htt97Q-GFP were treated with doxycyclin (1 μg/ml, 3 h). After removal of doxycyclin, cells were treated with HEP14 (20, 40 μM) or Torin1 (5 μM). The number of polyQ foci was scored at the indicated time points. n = 3 independent experiments. (b) Tet-on Htt97Q-GFP HeLa cells were transfected with the indicated PKCδ-expressing vectors. 12 h later, cells were treated with doxycycline as in (a) in the absence or presence of BFA1 (0.4 μM, 48 h). polyQ foci were examined 48 h after removal of doxycyclin. Images (left) and quantification (middle) of polyQ foci are shown. Ectopic protein expression is shown in the right panel. n = 3 independent experiments. (c) HepG2 cells were fed with oleic acid (100 μM, 12 h) to allow lipid droplets to accumulate. Following removal of oleic acid, cells were treated with HEP14 (40 μM) for the indicated times, then lipid droplets were stained with BODIPY (left) and quantified with flow cytometry (right). The BODIPY intensity measured at each time point was normalized to the start point of HEP14 treatment. n = 3 independent experiments. (d) Lipid droplet clearance assays were performed without or with BFA1 (0.4 μM) together with HEP14 (40 μM). Left: representative images of BODIPY staining of lipid droplets and LysoTracker Red staining of lysosomes in HepG2 cells at 48 h post treatment with HEP14, BFA1, or both. Right: quantification of BODIPY staining. n = 3 independent experiments. (e) HepG2 cells were transfected with the indicated PKCδ-expressing vectors. 12 h later, lipid droplet clearance assays were performed as in (c) without or with BFA1 (0.4 μM). Images (left) and quantification (middle) of BODIPY staining are shown. Ectopic protein expression is shown at the right. n = 3 independent experiments. Bars represent 10 μm in all images. For quantifications, data (mean ± s.e.m) were compared using t-tests or ANOVA. ∗ p < 0.05, ∗∗p < 0.01, ∗∗∗ p < 0.001.
Supplementary Figure 7 Analysis of the body weights of mice with HEP14 injection and the effects of lentivirus-mediated knockdown of PKCα or PKCδ.
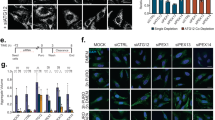
(a) Time-course measurement of the body weights of mice injected with vehicle alone (Veh) or HEP14 (5 mg/kg). n = 5 mice were measured for each group. (b) Knockdown of PKCα or PKCδ by Lenti-shPKCα or Lenti-shPKCδ vectors in mouse Neuro2a cells. Lenti-shNC represents control shRNA. The Lenti-shPKCα#1 and Lenti-shPKCδ#3 vectors were further used for preparing recombinant lentivirus and in vivo lentiviral grafting. Data are from one experiment that is representative of 3 independent experiments. (c) Graphic depiction of stereotactic injection of lentivirus expressing the indicated shRNAs into the hippocampus of mouse brain. Mice injected with virus were also i.p. injected with Veh or HEP14.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1085 kb)
Supplementary Table 1
Supplementary Information (XLSX 11 kb)
Supplementary Table 2
Supplementary Information (XLSX 11 kb)
Supplementary Table 3
Supplementary Information (XLSX 96 kb)
Supplementary Table 4
Supplementary Information (XLSX 10 kb)
Supplementary Table 5
Supplementary Information (XLSX 15 kb)
Supplementary Table 6
Supplementary Information (XLSX 11 kb)
Supplementary Table 7
Supplementary Information (XLSX 98 kb)
Rights and permissions
About this article
Cite this article
Li, Y., Xu, M., Ding, X. et al. Protein kinase C controls lysosome biogenesis independently of mTORC1. Nat Cell Biol 18, 1065–1077 (2016). https://doi.org/10.1038/ncb3407
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3407
This article is cited by
-
TFEB: a double-edged sword for tumor metastasis
Journal of Molecular Medicine (2023)
-
Past, present, and future perspectives of transcription factor EB (TFEB): mechanisms of regulation and association with disease
Cell Death & Differentiation (2022)
-
Alteration of Trop-2 expression in breast cancer cells by clinically used therapeutic agents and acquired tamoxifen resistance
Breast Cancer (2022)
-
Role of lysosomes in physiological activities, diseases, and therapy
Journal of Hematology & Oncology (2021)
-
Autophagy is affected in patients with hypokalemic periodic paralysis: an involvement in vacuolar myopathy?
Acta Neuropathologica Communications (2021)