Abstract
Small cell lung cancer (SCLC) constitutes approximately 15% of all diagnosed lung cancers. SCLC is a particularly lethal malignancy, as the 2-year survival rate after appropriate treatment is less than 5%. The patients with SCLC have not been received a benefit of the recently developed molecular targeted treatment. Therefore, a new treatment strategy is necessary for the patients. The molecular mechanisms underlying the aggressiveness of SCLC cells and their development of treatment-resistance are still ambiguous. In this study, we newly constructed a microRNA (miRNA) expression signature of SCLC by analysis of autopsy specimens. Based on the resultant signature, four miRNAs (miR-27a-5p, miR-485-3p, miR-34-5p and miR-574-3p) were found to be candidate anti-tumor miRNAs. To investigate their functional importance, we first validated the downregulation of miR-27a-5p and miR-34b-3p in SCLC clinical specimens. Next, we demonstrated that ectopic expression of both miR-27a-5p and miR-34b-3p significantly inhibited cancer cell aggressiveness. Our in silico analyses showed that four genes (topoisomerase 2 alpha (TOP2A), maternal embryonic leucine zipper kinase (MELK), centromere protein F (CENPF) and SRY-box 1 (SOX1) were identified as miR-27a-5p- and miR-34b-3p-regulated genes. Based on immunohistochemical analysis, TOP2A, MELK and CENPF were involved in SCLC pathogenesis. These genes might contribute to high proliferation and early metastatic spread of SCLC cells. Elucidation of differentially expressed miRNA-mediated cancer pathways based on SCLC signature may provide new insights into the mechanisms of SCLC pathogenesis.
Similar content being viewed by others
Introduction
Small cell lung cancer (SCLC) constitutes approximately 15–20% of all diagnosed lung cancers.1, 2 Owing to its aggressive nature, SCLC is a particularly lethal malignancy. SCLC cells characteristically acquire rapid cell proliferation and the ability to metastasize to distant sites. The majority of patients with SCLC present with metastatic disease, and the median survival time with combination chemotherapy is under 1 year for patients with extensive disease.1, 2 The conventional first-line treatment for SCLC with extensive disease is platinum-based chemotherapy.3, 4, 5 Although the response rate of the treatment is good (70–80%), cancer cells acquire early resistance to conventional treatments. Median progression-free survival is 5–6 months,3, 4, 5 and the disease escalates its aggressiveness. The patients with SCLC have not been received a benefit of the recently developed molecular targeted treatment. Therefore, understanding the molecular mechanisms of SCLC aggressiveness through current genomic approaches is needed.
The discovery of non-coding RNA in the human genome provided new directions for the study of human cancer pathogenesis.6 MicroRNAs (miRNAs) belong to a member of non-coding RNAs that act as sequence-specific fine tuners of the expression levels of proteins and RNAs.7, 8 A single miRNA can regulate a large number of RNA transcripts in human cells.9 Thus, aberrantly expressed miRNAs cause disruption of tightly regulated RNA networks, leading to pathologic behavior of cancer cells.10, 11 Currently, numerous studies have indicated that aberrantly expressed miRNAs are deeply involved in cancer pathogenesis.10, 11
We have been revealed the anti-tumor miRNAs and their controlled cancer pathways by using miRNA expression signatures of several cancers, including lung cancer.12, 13, 14 The next challenge in our miRNA studies is to identify key molecules and novel pathways involved in the resistance of cancer cells to current treatments. Based on this, we have constructed miRNA expression signatures by analyzing autopsy specimens from patients with prostate cancer and renal cell carcinoma.15, 16 Based on these signatures, we previously identified tumor-suppressive miR-221/222-mediated castration-resistant prostate cancer pathways and miR-101-mediated sunitinib-resistant pathways.15, 16
The molecular mechanisms underlying the aggressiveness of SCLC cells and the development of therapy-resistance are still ambiguous. Thus, we have investigated the molecular pathways contributing to treatment-resistant cancer cells in an effort to develop the new therapeutic strategies in SCLC. In the present study, we newly constructed miRNA expression signatures through analysis of primary and metastatic lesions (liver and brain). Based on the signatures, we identified two tumor-suppressive miRNAs (miR-27a-5p and miR-34b-3p) that are deeply involved in SCLC pathogenesis. Elucidation of the miRNA signature of SCLC may be useful for identification of novel molecular mechanisms of SCLC recurrence, metastasis and drug resistance.
Materials and methods
Patients and clinical lung cancer specimens
Clinical lung specimens were obtained from patients admitted to the Kagoshima University Hospital from 2011 to 2015. The patients’ backgrounds and clinical characteristics are summarized in Table 1 and Supplementary Table S1. Normal tissues are summarized in Supplementary Table S2. Archival formalin-fixed, paraffin-embedded samples were used for expression analysis and immunohistochemistry. Clinical specimens were staged according to the International Association for the Study of Lung Cancer TNM classification.17 This protocol was approved by the Institutional Review Board for Clinical Research of Kagoshima University School of Medicine.
Construction of the miRNA expression signature of SCLC based on autopsy specimens
A patient (64-year-old Japanese man) who died of SCLC underwent an autopsy. The patient had an excessive tobacco-smoking custom (90 pack years). His father had also died of lung cancer. Immunohistochemical examination demonstrated consistent expression of the neuron-specific antigen (synaptophysin and CD56). Specimens were obtained from primary lung lesions and metastatic liver and brain lesions. The clinical course of the patient is summarized in Figure 1.
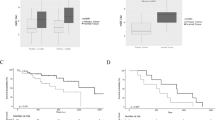
Clinical course of the SCLC patient who provided autopsy tissues. In May 2013, a 64-year-old Japanese man was admitted with increased shortness of breath. Consolidation was found in the left upper lung, and wall thickness of the pulmonary bulla was found in the right lower field in chest radiographs. He was diagnosed with lung synchronous carcinoma with multiple liver metastases. Bronchoscopic biopsy revealed small lung cell cancer (SCLC) from left B1+2 and squamous cell carcinoma from right B6. These synchronous carcinomas responded to first-line chemotherapy CBDCA+VP-16 (a total of six courses). But consolidation of the left upper lung (SCLC) worsened only 3 weeks after the last course of chemotherapy. Chemotherapy was changed to amrubicin, but disturbance of consciousness gradually appeared and worsened. Magnetic resonance imaging (MRI) findings and cytology of the cerebrospinal fluid (SCLC) were consistent with brain metastasis and meningeal carcinomatosis. In November 2013, he died, and the patient underwent autopsy. We obtained SCLC tissues of the left upper lung, liver metastasis and brain metastasis.
Expression of miRNA patterns were analyzed by the TaqMan LDA Human microRNA Panel v2.0 (Applied Biosystems, Foster City, CA, USA). The assay procedure was described previously.14 A cutoff P-value of <0.05 was used to narrow down the candidates after global normalization of the raw data. After global normalization, additional normalization was carried out with RNU48.
Cell lines, RNA isolation
Human SCLC cell lines (SBC-3 and NCI-H466 cells) were obtained from the Japanese Cancer Research Resources Bank (Osaka, Japan) and the American Type Culture Collection (Manassas, VA, USA), respectively.
Total RNA was isolated from cultured cells using Isogen (Nippon Gene, Tokyo, Japan). Total RNA was obtained from formalin-fixed, paraffin-embedded human clinical specimens using Recover All Total Nucleic Acid Isolation kit (Ambion, Austin, TX, USA) as described previously.18, 19, 20
Quantitative real-time reverse transcription-PCR
The procedure for PCR quantification was described previously.18, 19, 20 Stem–loop reverse transcription-PCRs for miR-27a-5p (P/N: 002445; Applied Biosystems), miR-485-3p (P/N: 001277), miR-34b-3p (P/N: 002102) and miR-574-3p (P/N: 002349) were used in this study. Human GUSB (P/N: Hs99999908_m1; Applied Biosystems) or RNU48 (P/N: 001006; Applied Biosystems) were used to normalize the data for quantification of mRNA and miRNAs, respectively.
Transfection with miRNA mimic and cell proliferation, migration and invasion assays
The following mature miRNA species were used in the present study: Pre-miR miRNA precursors (hsa-miR-27a-5p, P/N: AM 13096; hsa-miR-485-3p, P/N: AM 10799; hsa-miR-34b-3p, P/N: AM 12727; hsa-miR-574-3p, P/N: AM 12848; Applied Biosystems) and negative control miRNA, P/N: AM 17111; Applied Biosystems. RNAs were incubated with OPTI-MEM (Invitrogen, Carlsbad, CA, USA) and Lipofectamine RNAiMAX reagent (Invitrogen) as described previously.18, 19, 20
Cells were transfected with 10 nm miRNAs by reverse transfection as described previously.18, 19, 20 Cell migration assays and cell invasion assays were performed using modified Boyden chambers with 8 μm pores in 24-well tissue culture plates. Chambers for cell invasion assays consisted of Transwell-precoated Matrigel membrane filter inserts (BD Biosciences, Bedford, MA, USA). After 48 h of transfection, cells were plated in 24-well plates at 4 × 105 cells (SBC-3 cells) and 2 × 105 cells (NCI-H446 cells) per well, respectively. All experiments were performed in three independent trials.
Identification of oncogenic genes targeted by tumor-suppressive miRNAs
To identify genes putatively targeted by miR-27a-5p and miR-34b-3p, we performed in silico analysis using the TargetScan database and GEO expression data. First, we screened miR-27a-5p- and miR-34b-3p-targeted genes using the TargetScan database (release 7.1: http://www.targetscan.org/vert_71/). Next, we paired down the lists of genes based on a publicly available gene expression data set in a GEO database (accession number: GSE43346). Finally, we identified common genes targeted by miR-27a-5p and miR-34b-3p.
Immunohistochemistry
Three tissue autopsy specimens were used: SCLC lung tissue in the primary lesion and two metastatic tissues from the liver and brain. Specimens were immunostained following the manufacturer's protocol with the Ultra-Vision Detection System (Thermo Scientific, Fremont, CA, USA). For immunohistochemistry, we used primary rabbit polyclonal antibodies against the following: TOP2A (1:200, HPA006458; Sigma-Aldrich, St Louis, MO, USA), MELK (1:200, HPA017214; Sigma-Aldrich), CENPF (1:400, ab5; Abcam, Cambridge, UK) and SOX1 (1:500, ab87775; Abcam). The procedure was carried out as described previously.18, 19
Statistics
The relationships between two groups and the numerical values obtained by real-time reverse transcription-PCR were analyzed using Mann–Whitney U-tests. The relationships among more than three variables and numerical values were analyzed using the Bonferroni-adjusted Mann–Whitney U-test.
Results
Construction of the miRNA expression signature of SCLC specimens
Using a PCR-based array system, we analyzed differentially expressed miRNAs in the primary SCLC lesion and metastatic lesions (liver and brain) and compared them with non-cancerous lesions. Based on this analysis, we listed the top 35 downregulated miRNAs in SCLC tissues (Table 2). Among them, we focused on four miRNAs (miR-27a-5p, miR-485-3p, miR-34b-3p and miR-574-3p) that were significantly downregulated in SCLC specimens.
Expression levels of miR-27a-5p, miR-485-3p, miR-34b-3p and miR-574-3p in lung cancer clinical specimens and SCLC cell lines
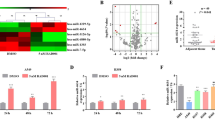
We evaluated the expression levels of four downregulated miRNAs (miR-27a-5p, miR-485-3p, miR-34b-3p and miR-574-3p) in non-cancerous (n=27), SCLC (n=11) and NSCLC (n=52) clinical specimens and in cell lines (SBC-3 and NCI-H446). The expression levels of miR-27a-5p and miR-34b-3p were significantly reduced in SCLC specimens compared with non-cancerous specimens (Figure 2). These miRNAs expression levels were also low in both SCLC cell lines (SBC-3 and NCI-H446).
Expression levels of miR-27a-5p, miR-485-3p, miR-34b-3p and miR-574-3p in lung cancer clinical specimens and SCLC cell lines. RT-PCR showed that the expression levels of miR-27a-5p and miR-34b-3p were significantly lower in SCLC clinical specimens and cell lines than in non-cancerous lung tissues. RNU48 was used as an internal control. Expression levels of (a) miR-27a-5p, (b) miR-485-3p, (c) miR-34b-3p and (d) miR-574-3p.
Effects of restoring miR-27a-5p, miR-485-3p, miR-34b-3p and miR-574-3p on cell proliferation, migration and invasion in SCLC cell lines
To investigate the functional roles of miRNAs in SCLC, we performed gain-of-function studies in SBC-3 and NCI-H446 cells by transfecting the cells with miRNA mimics. XTT assays showed that cell proliferation was significantly inhibited in miR-27a-3p transfectants of SBC-3 in comparison with the mock, although inhibition was not seen in NCI-H446. Additionally miR-34b-3p transfection significantly inhibited the proliferation of both cell lines in comparison with mock. Furthermore, cell proliferation was not inhibited in miR-485-3p and miR-574-3p transfectants of SBC-3 compared with mock, whereas it was significantly inhibited in NCI-H446 transfectants (Figure 3a).
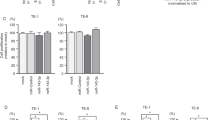
Functional significance of miR-27a-5p, miR-485-3p, miR-34b-3p and miR-574-3p in SCLC cell lines. (a) Cell proliferation was determined by XTT assays 96 h after transfection with four miRNAs at 10 nmol l−1 (miR-27a-5p, miR-485-3p, miR-34b-3p and miR-574-3p), miR-control or mock transfection. (b) Migration assays conducted 48 h after transfection. (c) Cell invasion assays conducted 48 h after transfection. *P<0.0083 and **P<0.0001.
In vitro assays showed that cell migration was inhibited in miR-485-3p transfectants of SBC-3 in comparison with the mock, whereas it was not inhibited in NCI-H446 transfectants. miR-34b-3p transfection significantly inhibited cell migration in both cell lines compared with mock. Cell migration activity was not inhibited in miR-574-3p transfectants of SBC-3 compared with mock, whereas it was significantly inhibited in NCI-H446 transfectants (Figure 3b).
Finally, Matrigel invasion assays demonstrated that cell invasive activity was significantly inhibited in miR-27a-3p, miR-485-3p and miR-574-3p transfectants of SBC-3 in comparison with the mock. Inhibition was not seen in NCI-H446 transfectants. Finally, miR-34b-3p transfection significantly inhibited cell invasion in both cell lines (Figure 3c).
Identification of putative target genes regulated by miR-27a-5p and miR-34b-3p in SCLC
To identify putative target genes subjected to miR-27a-5p and miR-34b-3p regulation, we performed in silico analysis using the TargetScan database and GEO expression data. First, we screened miR-27a-5p- and miR-34b-3p-targeted genes using the TargetScan database (release 7.1: http://www.targetscan.org/vert_71/). We found 3238 and 4165 genes that had putative target sites in their 3′-UTRs for miR-27a-5p and for miR-34b-3p, respectively. Second, we paired down the lists of genes based on a publicly available gene expression data set in the GEO database (accession number: GSE43346) and identified 7 and 17 genes, respectively. Finally, we selected the following four common genes from those lists: TOP2A, MELK, CENPF and SOX1. The flow chart outlining our strategy for identification of putative target genes of miR-27a-5p and miR-34b-3p is shown in Figure 4.
Strategy for identification of putative candidate genes controlled by miR-27a-5p and miR-34b-3p in SCLC. Outline of the identification of miR-27a-5p and miR-34b-3p target genes by in silico analysis of the TargetScan database and genome-wide gene expression analysis using a publicly available gene expression data set in the GEO database.
Immunohistochemical staining of miR-targeted proteins (TOP2A, MELK, CENPF and SOX1) in SCLC clinical specimens
To validate expression of TOP2A, MELK, CENPF and SOX1 proteins, immunohistochemistry was used to assess SCLC autopsy specimens. Immunohistochemical staining demonstrated overexpression of TOP2A in the nuclei of the primary lesion and liver metastasis (Figure 5a). The expression of MELK was high in the cytoplasm of all sites (Figure 5b). Also, the expression of CENPF was relatively high in the nuclei of the primary lesion (Figure 5c). However the expression of SOX1 was not observed in any site. Therefore, it appeared that TOP2A, MELK and CENPF play key roles as oncogenes regulated by miR-27a-5p and miR-34b-3p. TOP2A and MELK appeared to be particularly important in the development of SCLC.
Immunohistochemical staining of TOP2A, MELK and CENPF in SCLC autopsy specimens. (a) Overexpression of TOP2A in the nucleus was observed in the primary lesion and liver metastasis. (b) The expression of MELK was high in the cytoplasm. (c) The expression of CENPF was relatively high in the nuclei of the primary lesion.
Discussion
SCLC is a highly aggressive cancer with a poor prognosis because most patients are diagnosed with extensive disease.1, 2 Treatment strategies for late stage SCLC are commonly platinum-based chemotherapy.3, 4 Although SCLC cells are initially very sensitive to cytotoxic chemotherapy, in most cases, SCLC cells acquire resistance to these treatments.5 No effective treatments are approved for recurrence and the distant metastases of the disease. To improve the dismal prognosis of SCLC, effective treatment strategies are urgently needed. Uncovering the genomic characteristics of the disease might add new therapeutic targets.
A growing body of evidence has shown that dysregulated miRNAs are deeply involved in human oncogenesis, metastasis and drug resistance.11 Aberrantly expressed miRNAs can destroy tightly controlled RNA networks and promote oncogenic development. In this study, we first identified dysregulated miRNAs based on miRNA expression signatures of autopsy specimens from a patient with SCLC. The patient had undergone several therapeutic treatments, thus we considered that these specimens were likely treatment-resistant SCLC cells.
Our present data showed that several miRNAs, such as miR-27a-5p, miR-485-3p, miR-34b-3p and miR-574-3p, were markedly downregulated in cancerous tissues based on profiles of SCLC autopsy specimens. Among these miRNAs, clinical specimens confirmed that the expression levels of miR-27a-5p and miR-34b-3p were significantly reduced in SCLC tissues compared with non-cancerous tissues. Additionally, ectopic expression of these two miRNAs significantly inhibited cancer cell aggressiveness, suggesting that miR-27a-5p and miR-34b-3p functioned as tumor suppressors in SCLC cells. Based on these results, we focused on these two miRNAs and explored the molecular networks that they regulated.
By miRNA database searching (miRBase; http://www.mirbase.org/), pre-miR-27a produces two types of mature miRNAs, miR-27a-5p and miR-27a-3p. The miR-27a-5p is passenger strand of pre-miR-27a, whereas miR-27a-3p is guide strand of it. Most of past articles focused on the functional significance of the miR-27a-3p in several cancers. The functional significance of miR-27a-3p has confusion in cancer cells, including lung cancer. Previous studies have shown that miR-27a-3p is frequently upregulated and plays functional roles in multiple tumor types, including pancreatic cancer, breast cancer, ovarian cancer, esophageal cancer, renal cell carcinoma, hepatocellular carcinoma, glioma and gastric cancer.21 The guide strand of miR-27a-3p promotes tumorigenesis in several types of cancer.21, 22 On the other hand, miR-27a-3p was directly regulated tyrosine kinase receptors, EGFR and MET in lung cancer.23 Cancer stem cells is a promising target for cancer therapy in cases of cancer cell aggressiveness and drug resistance. Downregulation of miR-27a-3p was observed in sphere-forming cells in SCLC.24 Inhibition of miR-27a-3p in parental cells enhanced stem-like properties of SCLC cells in vitro.24
The passenger strand of pre-miR-27a (miR-27a-5p) is downregulated in head and neck squamous cell carcinoma, and it acts as a tumor suppressor by targeting the EGFR signaling axis. miR-27a-5p simultaneously decreases expression of EGFR, AKT1 and mTOR, leading to decreased solid tumor viability.22 In the past established theory of miRNA biogenesis, the passenger strand of miRNA is degradation and not incorporated into RNA-induced silencing complex.7 Surprisingly, our recent studies showed that miR-145-3p (passenger stand of pre-miR-145) actually functioned as anti-tumor miRNA in lung cancer and bladder cancer.25, 26 Similarly, we confirmed the anti-tumor function of miR-139-3p (passenger strand of pre-miR-139) in bladder cancer.27 These findings indicate that some passenger strand of miRNAs have biological function in cells. The involvement of passenger strand miRNAs in the regulation of cellular processes is a novel concept in RNA research.
It is an important study theme to investigate molecular mechanisms of transcriptional control of miRNAs in cancer cells. Silencing mechanisms of miR-27a-5p remain largely undefined. A recent study showed that Twist-1, a transcription factor of epithelial–mesenchymal transition regulation, was negatively controlled by miR-27a-3p expression in hepatocellular carcinoma.28 Other study showed that hepatocyte growth factor-mediated MET signal induced miR-27a-3p expression in lung cancer cells.23 The further study is necessary to elucidate expression control of miR-27a-5p in SCLC.
The expression level of miR-34b-3p is decreased in several cancers, such as neuroblastoma and cervical cancer.29, 30 In neuroblastoma, miR-34b-3p functions as a tumor suppressor by targeting CCNE2 and E2F3.29 A representative tumor suppressor p53 regulated the expression control of several miRNAs. The miR-34-family (miR-34a/b/c) was direct targets of p53 regulation.31 Expression of the miR-34-family caused antitumor effects, such as inducing apoptosis and cell cycle arrest. Therefore, cancer cells enhance inactivation of miR-34-family expression through CpG methylation.31
However, the functions of these miRNAs are still not fully understood. Based on those studies and our findings, we investigated the molecular networks regulated by the miRNAs that we identified to better understand the etiology of SCLC. We hypothesized that miR-27a-5p and miR-34b-3p might coordinately regulate target genes associated with SCLC pathogenesis. Therefore, we performed in silico analysis and identified four genes (TOP2A, MELK, CENPF and SOX1) that were potential targets of miR-27a-5p and miR-34b-3p. Immunohistochemistry indicated that MELK, TOP2A and CENPF play key roles in promoting oncogenesis. We suggest that TOP2A and MELK are particularly important in SCLC.
MELK is classified as a member of the SLK/AMPK serine–threonine kinase family and known as an embryonic and neural stem cell marker. It is associated with cell survival, proliferation and apoptosis in various cancers.32, 33 Several studies have reported a correlation between MELK gene expression and tumor malignancy grade for astrocytoma and breast cancer.34, 35 It was reported that MELK could play roles in cell cycle regulation (possibly in G0–G1 and S phases) as well as in responses against radiation and 5-FU treatment in colorectal cancer cells.32 Inoue et al.36 demonstrated that MELK was highly expressed in most SCLC cell lines and primary SCLC tumors. In this study, we demonstrated that the cancerous tissues of autopsy specimens that were presumably resistant to drug therapies stained strongly for MELK, suggesting that MELK was involved in resistance to chemotherapy in SCLC.
TOP2A is a subfamily of DNA topoisomerase type II that controls and alters the topologic states of DNA during transcription. Abnormal alterations of TOP2A include changes in gene copy number and gene expression level in cancer cells.37 Aberrant expression of TOP2A is generally associated with poor prognosis in breast cancer, ovarian cancer, oral cancer, esophageal cancer and lung cancer.38, 39, 40 In particular, multidrug resistance protein and TOP2A are involved in drug resistance in NSCLC.41 We confirmed the upregulation of TOP2A in SCLC autopsy specimens by immunohistochemistry.
CENPF encodes a protein that associates with the centromere–kinetochore complex. This protein is a member of the centromere protein family and acts in a critical chromosomal segregation process, including kinetochore assembly and spindle checkpoint signaling during mitosis.42 Overexpression of CENPF has been observed in prostate cancer and breast cancer.42, 43 Our recent study showed that downregulated tumor-suppressive miR-205 enhanced prostate cancer aggressiveness through direct regulation of CENPF.42
In conclusion, the expression of miR-27a-5p and miR-34b-3p was downregulated in SCLC autopsy specimens. They both act as tumor suppressors in SCLC cells. Oncogenic MELK, TOP2A and CENPF are regulated by these miRNAs, and high expression of those oncogenes in SCLC autopsy specimens indicates clinical importance. Elucidation of miR-27a-5p- and miR-34b-3p-mediated molecular networks may lead to a better understanding of SCLC aggressiveness and the development of new treatment strategies.
Accession codes
References
Chute, J. P., Chen, T., Feigal, E., Simon, R. & Johnson, B. E. Twenty years of phase III trials for patients with extensive-stage small-cell lung cancer: perceptible progress. J. Clin. Oncol. 17, 1794–1801 (1999).
Lassen, U., Osterlind, K., Hansen, M., Dombernowsky, P., Bergman, B. & Hansen, H. H. Long-term survival in small-cell lung cancer: posttreatment characteristics in patients surviving 5 to 18+ years—an analysis of 1714 consecutive patients. J. Clin. Oncol. 13, 1215–1220 (1995).
Amarasena, I. U., Walters, J. A., Wood-Baker, R. & Fong, K. Platinum versus non-platinum chemotherapy regimens for small cell lung cancer. Cochrane Database Syst. Rev. 8, CD006849 (2015).
Satouchi, M., Kotani, Y., Shibata, T., Ando, M., Nakagawa, K., Yamamoto, N. et al. Phase III study comparing amrubicin plus cisplatin with irinotecan plus cisplatin in the treatment of extensive-disease small-cell lung cancer: JCOG 0509. J. Clin. Oncol. 32, 1262–1268 (2014).
van Meerbeeck, J. P., Fennell, D. A. & De Ruysscher, D. K. Small-cell lung cancer. Lancet 378, 1741–1755 (2011).
Carthew, R. W. & Sontheimer, E. J. Origins and mechanisms of miRNAs and siRNAs. Cell 136, 642–655 (2009).
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Filipowicz, W., Bhattacharyya, S. N. & Sonenberg, N. Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nat. Rev. Genet. 9, 102–114 (2008).
Friedman, R. C., Farh, K. K., Burge, C. B. & Bartel, D. P. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 19, 92–105 (2009).
Nelson, K. M. & Weiss, G. J. MicroRNAs and cancer: past, present, and potential future. Mol Cancer Ther 7, 3655–3660 (2008).
Wiemer, E. A. The role of microRNAs in cancer: no small matter. Eur. J. Cancer. 43, 1529–1544 (2007).
Itesako, T., Seki, N., Yoshino, H., Chiyomaru, T., Yamasaki, T., Hidaka, H. et al. The microRNA expression signature of bladder cancer by deep sequencing: the functional significance of the miR-195/497 cluster. PLoS ONE 9, e84311 (2014).
Fukumoto, I., Kinoshita, T., Hanazawa, T., Kikkawa, N., Chiyomaru, T., Enokida, H. et al. Identification of tumour suppressive microRNA-451a in hypopharyngeal squamous cell carcinoma based on microRNA expression signature. Br. J. Cancer 111, 386–394 (2014).
Moriya, Y., Nohata, N., Kinoshita, T., Mutallip, M., Okamoto, T., Yoshida, S. et al. Tumor suppressive microRNA-133a regulates novel molecular networks in lung squamous cell carcinoma. J. Hum. Genet. 57, 38–45 (2012).
Goto, Y., Kojima, S., Nishikawa, R., Kurozumi, A., Kato, M., Enokida, H. et al. MicroRNA expression signature of castration-resistant prostate cancer: the microRNA-221/222 cluster functions as a tumour suppressor and disease progression marker. Br. J. Cancer 113, 1055–1065 (2015).
Goto, Y., Kurozumi, A., Nohata, N., Kojima, S., Matsushita, R., Yoshino, H. et al. The microRNA signature of patients with sunitinib failure: regulation of UHRF1 pathways by microRNA-101 in renal cell carcinoma. Oncotarget 7, 59070–59096 (2016).
Shepherd, F. A., Crowley, J., Van Houtte, P., Postmus, P. E., Carney, D., Chansky, K. et al. The International Association for the Study of Lung Cancer Lung Cancer Staging Project: proposals regarding the clinical staging of small cell lung cancer in the forthcoming (seventh) edition of the tumor, node, metastasis classification for lung cancer. J. Thorac. Oncol. 2, 1067–1077 (2007).
Mataki, H., Seki, N., Chiyomaru, T., Enokida, H., Goto, Y., Kumamoto, T. et al. Tumor-suppressive microRNA-206 as a dual inhibitor of MET and EGFR oncogenic signaling in lung squamous cell carcinoma. Int. J. Oncol. 46, 1039–1050 (2015).
Kamikawaji, K., Seki, N., Watanabe, M., Mataki, H., Kumamoto, T., Takagi, K. et al. Regulation of LOXL2 and SERPINH1 by antitumor microRNA-29a in lung cancer with idiopathic pulmonary fibrosis. J. Hum. Genet. 61, 985–993 (2016).
Mataki, H., Enokida, H., Chiyomaru, T., Mizuno, K., Matsushita, R., Goto, Y. et al. Downregulation of the microRNA-1/133a cluster enhances cancer cell migration and invasion in lung-squamous cell carcinoma via regulation of Coronin1C. J. Hum. Genet. 60, 53–61 (2015).
Zhou, L., Liang, X., Zhang, L., Yang, L., Nagao, N., Wu, H. et al. MiR-27a-3p functions as an oncogene in gastric cancer by targeting BTG2. Oncotarget 7, 51943–51954 (2016).
Wu, X., Bhayani, M. K., Dodge, C. T., Nicoloso, M. S., Chen, Y., Yan, X. et al. Coordinated targeting of the EGFR signaling axis by microRNA-27a*. Oncotarget 4, 1388–1398 (2013).
Acunzo, M., Romano, G., Palmieri, D., Lagana, A., Garofalo, M., Balatti, V. et al. Cross-talk between MET and EGFR in non-small cell lung cancer involves miR-27a and Sprouty2. Proc. Natl Acad. Sci. USA 110, 8573–8578 (2013).
Miao, Y., Li, J., Qiu, X., Li, Y., Wang, Z. & Luan, Y. miR-27a regulates the self renewal of the H446 small cell lung cancer cell line in vitro. Oncol. Rep. 29, 161–168 (2013).
Mataki, H., Seki, N., Mizuno, K., Nohata, N., Kamikawaji, K., Kumamoto, T. et al. Dual-strand tumor-suppressor microRNA-145 (miR-145-5p and miR-145-3p) coordinately targeted MTDH in lung squamous cell carcinoma. Oncotarget 7, 72084–72098 (2016).
Matsushita, R., Yoshino, H., Enokida, H., Goto, Y., Miyamoto, K., Yonemori, M. et al. Regulation of UHRF1 by dual-strand tumor-suppressor microRNA-145 (miR-145-5p and miR-145-3p): Inhibition of bladder cancer cell aggressiveness. Oncotarget 7, 28460–28487 (2016).
Yonemori, M., Seki, N., Yoshino, H., Matsushita, R., Miyamoto, K., Nakagawa, M. et al. Dual tumor-suppressors miR-139-5p and miR-139-3p targeting matrix metalloprotease 11 in bladder cancer. Cancer Sci. 107, 1233–1242 (2016).
Zhao, N., Sun, H., Sun, B., Zhu, D., Zhao, X., Wang, Y. et al. miR-27a-3p suppresses tumor metastasis and VM by down-regulating VE-cadherin expression and inhibiting EMT: an essential role for Twist-1 in HCC. Sci. Rep. 6, 23091 (2016).
Maugeri, M., Barbagallo, D., Barbagallo, C., Banelli, B., Di Mauro, S., Purrello, F. et al. Altered expression of miRNAs and methylation of their promoters are correlated in neuroblastoma. Oncotarget 7, 83330–83341 (2016).
Revathidevi, S., Manikandan, M., Rao, A. K., Vinothkumar, V., Arunkumar, G., Rajkumar, K. S. et al. Analysis of APOBEC3A/3B germline deletion polymorphism in breast, cervical and oral cancers from South India and its impact on miRNA regulation. Tumour Biol. 37, 11983–11990 (2016).
Hermeking, H. The miR-34 family in cancer and apoptosis. Cell Death Differ. 17, 193–199 (2010).
Choi, S. & Ku, J. L. Resistance of colorectal cancer cells to radiation and 5-FU is associated with MELK expression. Biochem. Biophys. Res. Commun. 412, 207–213 (2011).
Lin, M. L., Park, J. H., Nishidate, T., Nakamura, Y. & Katagiri, T. Involvement of maternal embryonic leucine zipper kinase (MELK) in mammary carcinogenesis through interaction with Bcl-G, a pro-apoptotic member of the Bcl-2 family. Breast Cancer Res. 9, R17 (2007).
Ganguly, R., Mohyeldin, A., Thiel, J., Kornblum, H. I., Beullens, M. & Nakano, I. MELK-a conserved kinase: functions, signaling, cancer, and controversy. Clin. Transl. Med. 4, 11 (2015).
Ganguly, R., Hong, C. S., Smith, L. G., Kornblum, H. I. & Nakano, I. Maternal embryonic leucine zipper kinase: key kinase for stem cell phenotype in glioma and other cancers. Mol. Cancer Ther. 13, 1393–1398 (2014).
Inoue, H., Kato, T., Olugbile, S., Tamura, K., Chung, S., Miyamoto, T. et al. Effective growth-suppressive activity of maternal embryonic leucine-zipper kinase (MELK) inhibitor against small cell lung cancer. Oncotarget 7, 13621–13633 (2016).
Chen, T., Sun, Y., Ji, P., Kopetz, S. & Zhang, W. Topoisomerase IIalpha in chromosome instability and personalized cancer therapy. Oncogene 34, 4019–4031 (2015).
Ceppi, P., Longo, M., Volante, M., Novello, S., Cappia, S., Bacillo, E. et al. Excision repair cross complementing-1 and topoisomerase IIalpha gene expression in small-cell lung cancer patients treated with platinum and etoposide: a retrospective study. J. Thorac. Oncol. 3, 583–589 (2008).
Meng, H., Chen, R., Li, W., Xu, L. & Xu, L. Correlations of TOP2A gene aberrations and expression of topoisomerase IIalpha protein and TOP2A mRNA expression in primary breast cancer: a retrospective study of 86 cases using fluorescence in situ hybridization and immunohistochemistry. Pathol. Int. 62, 391–399 (2012).
Dingemans, A. M., Witlox, M. A., Stallaert, R. A., van der Valk, P., Postmus, P. E. & Giaccone, G. Expression of DNA topoisomerase IIalpha and topoisomerase IIbeta genes predicts survival and response to chemotherapy in patients with small cell lung cancer. Clin. Cancer Res. 5, 2048–2058 (1999).
Huang, H., Liu, J., Meng, Q. & Niu, G. Multidrug resistance protein and topoisomerase 2 alpha expression in non-small cell lung cancer are related with brain metastasis postoperatively. Int. J. Clin. Exp. Pathol. 8, 11537–11542 (2015).
Nishikawa, R., Goto, Y., Kurozumi, A., Matsushita, R., Enokida, H., Kojima, S. et al. MicroRNA-205 inhibits cancer cell migration and invasion via modulation of centromere protein F regulating pathways in prostate cancer. Int. J. Urol. 22, 867–877 (2015).
O'Brien, S. L., Fagan, A., Fox, E. J., Millikan, R. C., Culhane, A. C., Brennan, D. J. et al. CENP-F expression is associated with poor prognosis and chromosomal instability in patients with primary breast cancer. Int. J. Cancer 120, 1434–1443 (2007).
Acknowledgements
The present study was supported by KAKENHI(C) grant 15K10801 and 16K19458.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Mizuno, K., Mataki, H., Arai, T. et al. The microRNA expression signature of small cell lung cancer: tumor suppressors of miR-27a-5p and miR-34b-3p and their targeted oncogenes. J Hum Genet 62, 671–678 (2017). https://doi.org/10.1038/jhg.2017.27
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2017.27
This article is cited by
-
Recent advances on high-efficiency of microRNAs in different types of lung cancer: a comprehensive review
Cancer Cell International (2023)
-
MicroRNA in lung cancer—a novel potential way for early diagnosis and therapy
Journal of Applied Genetics (2023)
-
Circulating miR-148b-3p and miR-27a-3p can be potential biomarkers for diagnosis of pre-diabetes and type 2 diabetes: integrating experimental and in-silico approaches
BMC Endocrine Disorders (2022)
-
KLF5-induced BBOX1-AS1 contributes to cell malignant phenotypes in non-small cell lung cancer via sponging miR-27a-5p to up-regulate MELK and activate FAK signaling pathway
Journal of Experimental & Clinical Cancer Research (2021)
-
Piperlongumine inhibits the growth of non-small cell lung cancer cells via the miR-34b-3p/TGFBR1 pathway
BMC Complementary Medicine and Therapies (2021)