Abstract
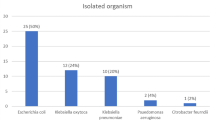
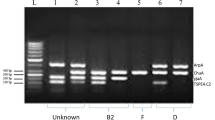
Humans frequently contract urinary tract infections (UTIs), which can be brought on by uropathogens (UPs) that are multi-drug resistant. Treatment for UTIs brought on by pathogenic UPs that produce extended-spectrum lactamases (ESBLs) is more costly and potentially fatal. As a result, the objective of this study was to use culture, biochemical, and 16S rRNA sequencing to identify and characterize UPs isolated from outpatients in Noakhali, Bangladesh, who had symptoms of UTIs. ESBL gene identification and quinolone resistance gene typing were then performed on the isolates using polymerase chain reaction (PCR). Throughout the trial’s 8-month duration, 152 (76%) of 200 urine samples were positive for the presence of UPs. The overall number of UPs recovered was 210, with 39 individuals having multiple UPs present in their samples. Among all of the isolates, Escherichia coli (45.24%, 95/210; 95% confidence interval (CI): 35.15–57.60%), Enterobacter spp. (24.76%, 52/210; CI: 19.15–35.77%), Klebsiella spp. (20.95%; 44/210; CI: 15.15–30.20%), and Providencia spp. (9.05%; 19/210; CI: 4.95–19.25%) were the four most prevalent bacteria found in the isolates. The UPs displayed a very high level of resistance to piperacillin 96.92% (126/130), ampicillin 90% (117/130), nalidixic acid 77.69% (101/130), cefazolin 70% (91/130), amoxicillin 50% (55/130), cefazolin 42.31% (55/130), nitrofurantoin 43.08% (56/130), and ciprofloxacin 33.08% (43/130), whereas resistance to netilmicin (3.85%), amikacin (4.62%), and imipenem (9.23%) was low. Individually, every species of E. coli and Providencia spp. showed greater ampicillin, amikacin, cefazolin, cefazolin, and nalidixic acid resistance than the others. The bivariate results indicate several antibiotic pairings, and isolates had meaningful associations. All MDR isolates were subjected to PCR, which revealed that blaCTX-M-15 genes predominated among the isolates, followed by the blaTEM class (37%). Isolates also had the qnrS, aac-6´-Ib-cr, and gyrA genes. The findings provide worrying indications of a major expansion of MDR isolates in the study locations, particularly the epidemiological balCTX-M 15, with the potential for the transmission of multi-drug-resistant UP strains in the population.






Similar content being viewed by others
Data availability
The data relating to this manuscript are available upon request.
References
Masajtis-Zagajewska A, Nowicki M (2017) New markers of urinary tract infection. Clin Chim Acta 471:286–291
Mishra MP, Sarangi R, Padhy RN (2016) Prevalence of multidrug resistant uropathogenic bacteria in pediatric patients of a tertiary care hospital in eastern India. J Infect Public Health 9(3):308–314
Ahmed N, Khalid H, Mushtaq M, Basha S, Rabaan AA, Garout M, Yean CY (2022) The molecular characterization of virulence determinants and antibiotic resistance patterns in human bacterial uropathogens. Antibiotics 11(4):516
Akram M, Shahid M, Khan AU (2007) Etiology and antibiotic resistance patterns of community-acquired urinary tract infections in JNMC Hospital Aligarh, India. Ann Clin Microbiol Antimicrob 6(1):1–7
Balachandran L, Jacob L, Al Awadhi R, Yahya LO, Catroon KM, Soundararajan LP,... Hussein YA (2022) Urinary tract infection in pregnancy and its effects on maternal and perinatal outcome: a retrospective study. Cureus 14(1)
Call ZD, Jang I, Geiss BJ, Dandy DS, Henry CS (2022) Progress toward a simplified UTI diagnostic: pump-free magnetophoresis for E. coli detection. Anal Chem 94(21):7545–7550
Gililland JM, Anderson LA, Erickson J, Pelt CE & Peters CL (2013) Mean 5-year clinical and radiographic outcomes of cementless total hip arthroplasty in patients under the age of 30. BioMed research international 2013
Khoshnood S, Heidary M, Mirnejad R, Bahramian A, Sedighi M, Mirzaei H (2017) Drug-resistant gram-negative uropathogens: a review. Biomed Pharmacother 94:982–994
Mollick S, Dasgupta T, Hasnain MJ, Ahmed M (2016) Isolation and characterization of pathogens responsible for urinary tract infection in Bangladesh and determination of their antibiotic susceptibility pattern. J Appl Pharmaceut Sci 6(4):072–076
Ahmed OB, Omar AO, Asghar AH, Elhassan MM, Al-Munawwarah AM, Arabia S (2013) Prevalence of TEM, SHV and CTX-M genes in Escherichia coli and Klebsiella spp urinary isolates from Sudan with confirmed ESBL phenotype. Life Sci J 10(2):191–195
Al-Naqshbandi AA, Chawsheen MA, Abdulqader HH (2019) Prevalence and antimicrobial susceptibility of bacterial pathogens isolated from urine specimens received in rizgary hospital—Erbil. J Infect Public Health 12(3):330–336
Jassim A-M (2010) In-home drug storage and self-medication with antimicrobial drugs in Basrah. Iraq Oman Med J 25(2):79
Paul R (2018) State of the globe: rising antimicrobial resistance of pathogens in urinary tract infection. J Global Infect Dis 10(3):117
Alizade H (2018) Escherichia coli in Iran: an overview of antibiotic resistance: a review article. Iran J Public Health 47(1):1
Novelli A, Rosi E (2017) Pharmacological properties of oral antibiotics for the treatment of uncomplicated urinary tract infections. J Chemother 29(1):10–18
Hasan MS, Sultana M, Hossain MA (2019) Complete genome arrangement revealed the emergence of a poultry origin superbug Citrobacter portucalensis strain NR-12. J Global Antimicrobial Resistance 18:126–129
Hossain A, Hossain SA, Fatema AN, Wahab A, Alam MM, Islam MN, Ahsan GU (2020) Age and gender-specific antibiotic resistance patterns among Bangladeshi patients with urinary tract infection caused by Escherichia coli. Heliyon 6(6):e04161
Sultana M, Parvez AK, Sultana KF, Mukharje SK, Hossain MA (2019) Characterization of extended spectrum β-Lactamase producing bactieria isolated from urinary tract infections. Bangladesh Med Res Counc Bull 45(1):23–33
Rajan S, Prabavathy JV (2012) Antibiotic sensitivity and phenotypic detection of ESBL producing E. coli strains causing urinary tract infection in a community hospital, Chennai, Tamil Nadu India. WebmedCentral Pharmaceut Sci 3(11):003840
Gundran RS, Cardenio PA, Villanueva MA, Sison FB, Benigno CC, Kreausukon K, Punyapornwithaya V (2019) Prevalence and distribution of blaCTX-M, blaSHV, blaTEM genes in extended-spectrum β-lactamase-producing E. coli isolates from broiler farms in the Philippines. BMC Vet Res 15(1):1–8
Shakya P, Shrestha D, Maharjan E, Sharma VK, Paudyal R (2017) ESBL production among E coli and Klebsiella spp causing urinary tract infection: a hospital based study. Open Microbiol J 11:23
Zowawi HM, Harris PN, Roberts MJ, Tambyah PA, Schembri MA, Pezzani MD, Paterson DI (2015) The emerging threat of multidrug-resistant Gram-negative bacteria in urology. Nat Rev Urol 12(10):570–584
Kubone PZ, Mlisana KP, Govinden U, Abia ALK, Essack SY (2020) Antibiotic susceptibility and molecular characterization of uropathogenic Escherichia coli associated with community-acquired urinary tract infections in urban and rural settings in South Africa. Tropical Med Infectious Dis 5(4):176
Ramachandran G, Rajivgandhi GN, Chackaravarthi G, Kanisha CC, Siddiqi MZ, Alharbi NS, Manoharan N (2021) Isolation and molecular identification of extended spectrum beta-lactamase producing bacteria from urinary tract infection. J Infection Public Health 14(12):1911–1916
Hoque MN, Talukder AK, Saha O, Hasan MM, Sultana M, Rahman AA, Das ZC (2022) Antibiogram and virulence profiling reveals multidrug resistant Staphylococcus aureus as the predominant aetiology of subclinical mastitis in riverine buffaloes. Vet Med Sci 8:2631–45
Clinical and Laboratory Standards Institut (2022) Performance standards for antimicrobial susceptibility testing, M100, 32nd ed. Clinical and Laboratory Standards Institute, Wayne, PA
Saha O, Rakhi NN, Hoque MN, Sultana M, Hossain MA (2021) Genome-wide genetic marker analysis and genotyping of Escherichia fergusonii strain OTSVEF-60. Braz J Microbiol 52(2):989–1004
Saha O, Hoque MN, Islam OK, Rahaman MM, Sultana M, Hossain MA (2020) Multidrug-resistant avian pathogenic Escherichia coli strains and association of their virulence genes in Bangladesh. Microorganisms 8(8):1135
Momtaz S, Saha O, Usha MK, Sultana M, Hossain MA (2018) Occurrence of pathogenic and multidrug resistant Salmonella spp in poultry slaughter-house in Bangladesh. Biores Commu (BRC) 4(2):506–515
Hoque MN, Istiaq A, Clement RA, Gibson KM, Saha O, Islam OK, Hossain MA (2020) Insights into the resistome of bovine clinical mastitis microbiome, a key factor in disease complication. Front Microbiol 11:860
Swindell SR & Plasterer TN (1997) Seqman. In Sequence data analysis guidebook (75–89). Springer, Totowa, NJ
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mole Biol Evol 33(7):1870–1874
Doi Y, Adams-Haduch JM, Endimiani A, Sidjabat HE, Gaddad SM, Paterson DL (2008) High prevalence of CTX-M-15-producing Klebsiella pneumoniae among inpatients and outpatients with urinary tract infection in Southern India. J Antimicrob Chemother 61(6):1393–1394
Hoşoğlu S, Gündeş S, Kolaylõ F, Karadenizli A, Demirdağ K, Günaydõn M, Ucmak H (2007) Extended-spectrum beta-lactamases in ceftazidime-resistant Escherichia coli and Klebsiella pneumoniae isolates in Turkish hospitals. Indian J Med Microbiol 25(4):346–350
Cattoir V, Poirel L, Rotimi V, Soussy CJ, Nordmann P (2007) Multiplex PCR for detection of plasmid-mediated quinolone resistance qnr genes in ESBL-producing enterobacterial isolates. J Antimicrob Chemother 60(2):394–397
Platell JL, Cobbold RN, Johnson JR, Heisig A, Heisig P, Clabots C, Trott DJ (2011) Commonality among fluoroquinolone-resistant sequence type ST131 extraintestinal Escherichia coli isolates from humans and companion animals in Australia. Antimicrobial Agents Chemother 55(8):3782–3787
Zhao L, Zhang J, Zheng B, Wei Z, Shen P, Li S, Xiao Y (2015) Molecular epidemiology and genetic diversity of fluoroquinolone-resistant Escherichia coli isolates from patients with community-onset infections in 30 Chinese county hospitals. J Clin Microbiol 53(3):766–770
Asare KK, Amoah S, Coomson CA Jr, Banson C, Yaro D, Mbata J, Opoku YK (2022) Antibiotic-resistant pathogenic bacterial isolates from patients attending the outpatient department of university of Cape Coast hospital, Ghana a retrospective study between 2013–2015. PLoS Global Public Health 2(5):e0000417
Haque R, Akter M, Salam M (2015) Prevalence and susceptibility of uropathogens: a recent report from a teaching hospital in Bangladesh. BMC Res Notes 8(1):1–5
Parveen R, Saha SK, Shamshuzzaman SM, Rashid AL, Chowdhury A, Muazzam N (2011) Detection of uropathogens by using chromogenic media (Hicrome UTI agar), CLED agar and other conventional media. Faridpur Med Coll J 6(1):46–50
Rahman SR, Ahmed MF, Begum A (2014) Occurrence of urinary tract infection in adolescent and adult women of shanty town in Dhaka City. Bangladesh Ethiopian J Health Sci 24(2):145–152
Niranjan V, Malini A (2014) Antimicrobial resistance pattern in Escherichia coli causing urinary tract infection among inpatients. Indian J Med Res 139(6):945
Ahmed M, Moremi N, Mirambo MM, Hokororo A, Mushi MF, Seni J, Mshana SE (2015) Multi-resistant gram negative enteric bacteria causing urinary tract infection among malnourished underfives admitted at a tertiary hospital, northwestern Tanzania. Italian J Pediat 41(1):1–5
Parveen R, Yusuf MA, Sharmin I, Islam MS, Rahim I (2015) Antibiotic sensitivity of bacteria causing urinary tract infection. Bangladesh J Infect Dis 2(1):13–18
Yousefi A, Torkan S (2017) Uropathogenic Escherichia coli in the urine samples of Iranian dogs: antimicrobial resistance pattern and distribution of antibiotic resistance genes. Biomed Res Int 2017:10
Singhal A, Sharma R, Jain M, Vyas L (2014) Hospital and community isolates of uropathogens and their antibiotic sensitivity pattern from a tertiary care hospital in North West India. Ann Med Health Sci Res 4(1):51–56
Beyene G, Tsegaye W (2011) Bacterial uropathogens in urinary tract infection and antibiotic susceptibility pattern in Jimma University Specialized Hospital, Southwest Ethiopia. Ethiop J Health Sci 21(2):141–146
Rahman MM, Haq JA, Hossain MA, Sultana R, Islam F, Islam AS (2004) Prevalence of extended-spectrum β-lactamase-producing Escherichia coli and Klebsiella pneumoniae in an urban hospital in Dhaka. Bangladesh Int J Antimicrobial Agents 24(5):508–510
Sharmin S, Alamgir F, Fahmida M, Saleh AA (2009) Antimicrobial sensitivity pattern of uropathogens in children. Bangladesh J Med Microbiol 3(2):18–22
Al Wutayd O, Al Nafeesah A, Adam I, Babikir I (2018) The antibiotic susceptibility patterns of uropathogens isolated in Qassim, Saudi Arabia. J Infect Dev Countries 12(11):946–952
Rani L, Pinnelli VBK, Hemavathi BS, Rajendran R (2012) Utility Urinary of HiChrome Tract Infection (UTI) agar medium for identification of uropathogens: a comparative study with other conventional media. J Chem Pharm Res 1(4):95–105
Soleimani N, Aganj M, Ali L, Shokoohizadeh L, Sakinc T (2014) Frequency distribution of genes encoding aminoglycoside modifying enzymes in uropathogenic E coli isolated from Iranian hospital. BMC Res Notes 7(1):1–5
Khaleque M, Akter S, Mondal DK, Akhter H, Khan SI, Begum A (2017) Molecular characterization of extended spectrum β-lactamase producing bacteria isolated from urinary tract infected patients. Bangladesh Tropical Biomed 34(3):512–523
Chervet D, Lortholary O, Zahar JR, Dufougeray A, Pilmis B, Partouche H (2018) Antimicrobial resistance in community-acquired urinary tract infections in Paris in 2015. Medecine et maladies infectieuses 48(3):188–192
Paiva MC, Nascimento AMA, Camargo ILBC, Lima-Bittencourt CI, Nardi RMD (2012) The first report of the qnrB19, qnrS1 and aac (6)-Ib-cr genes in urinary isolates of ciprofloxacin-resistant Escherichia coli in Brazil. Mem Inst Oswaldo Cruz 107:687–689
Acknowledgements
The authors acknowledge the Department of Microbiology, Noakhali Science and Technology University and Research Cell for providing the research facilities with funding.
Author information
Authors and Affiliations
Contributions
KFS, AA, SA, TJ, and OS carried out the studies (sampling, sequencing, molecular, and data analysis) and participated in drafting the manuscript. SRS and AA critically reviewed and drafted the manuscript. OS visualized figures, interpreted data and results, critically reviewed, and edited the manuscript. KFS and FA supervised the sampling and supervised the whole work. KFS and OS developed the hypothesis, supervised the whole work, and helped to prepare and critically revise the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This study obtained ethical clearance from the appropriate authorities. Moreover, verbal and written consent forms were also obtained from all of the participants.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Responsible Editor: Fernando R. Pavan
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sultana, K.F., Akter, A., Saha, S.R. et al. Bacterial profile, antimicrobial resistance, and molecular detection of ESBL and quinolone resistance gene of uropathogens causing urinary tract infection in the southeastern part of Bangladesh. Braz J Microbiol 54, 803–815 (2023). https://doi.org/10.1007/s42770-023-00942-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-023-00942-2