Abstract
Dermatophytes are a group of eukaryotic microorganisms characterized by high capacity to colonize keratinized structures such as the skin, hair, and nails. Over the past years, the incidence of infections caused by zoophilic species, e.g., Trichophyton verrucosum, has been increasing in some parts of the world, especially in Europe. Moreover, the emergence of recalcitrant dermatophytoses and in vitro resistant dermatophytes has become a cause of concern worldwide. Here, we analyzed the mechanisms underlying resistance to fluconazole among clinical isolates of T. verrucosum. Quantitative RT-PCR was carried out to determine the relative expression levels of mRNA transcripts of ERG3, ERG6, and ERG11 genes in the fungal samples using the housekeeping gene GAPDH as a reference. Our results showed that the upregulation of the ERG gene expression is a possible mechanism of resistance to fluconazole in this species. Furthermore, ERG11 is the most statistically significantly overexpressed gene in the pool of fluconazole-resistant T. verrucosum isolates. Additionally, we have demonstrated that exposure to fluconazole increases the levels of expression of ERG genes in fluconazole-resistant isolates of T. verrucosum. In conclusion, this study has shown one of the possible mechanisms of resistance to fluconazole among zoophilic dermatophytes, which involves the maintenance of high levels of expression of ERG genes after drug exposure.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Dermatophytes are a group of primarily pathogenic filamentous fungi classified in the Arthrodermataceae family, within the order Onygenales [1]. These eukaryotic microorganisms are characterized by a high capacity to attack keratinized structures such as the skin, hair, and nails in humans and animals causing mainly superficial cutaneous infections [2]. Indeed, a biological characteristic of dermatophytes is their ability to invade keratin-rich tissues by producing enzymes [3]. Trichophyton rubrum is the commonest causative agent with a diagnosis frequency comparable to the state of a global epidemic [4]. However, the incidence of infections caused by zoophilic dermatophytes T. mentagrophytes [2, 5, 6] and less common T. verrucosum is increasing in some parts of the world [7,8,9,10,11]. Although most of the literature data indicate the occurrence of infections caused by these species of dermatophytes in animals, zoonoses of these etiologies have been reported in humans professionally involved in animal husbandry [9, 12, 13]. Noteworthy, the reverse zoonosis caused by T. verrucosum has also been reported in Europe [8].
The emergence of recalcitrant dermatophytoses and in vitro resistant dermatophytes over the past few years has become a cause of concern worldwide [2, 14, 15]. Consequently, this has led to renewed research interest in dermatophytoses, one of the commonest fungal infections of humans and animals [16, 17]. However, molecular mechanisms underlying the reduced susceptibility to antifungals are not fully described or clearly elucidated.
Antifungal drugs commonly used in the treatment of dermatophytoses largely interfere with the different steps of the ergosterol synthesis pathway [14]. Ergosterol is an essential component of fungal plasma membranes [18]. Azoles, including fluconazole which is the main object of the present study, inhibit lanosterol 14α-demethylase (CYP51A1), i.e., a cytochrome P450-dependent enzyme, causing ergosterol depletion and disturbing the fluidity, permeability, and activity of membrane-associated proteins [19]. Azole-resistant phenotypes of dermatophytes are usually related to point mutations in the ERG11 (encoding the enzyme lanosterol 14-α demethylase) or ERG3 (encoding the enzyme sterol C-5 desaturase) and ERG6 (encoding the enzyme sterol C-24 methyltransferase) genes, their duplication conferring overexpression of target enzymes, or mutations in the cytochrome P450-encoding gene affecting drug binding [20]. Another mechanism of azole resistance is connected with overexpression of genes related with ATP binding cassette (ABC) transporters responsible for drug efflux [21].
In the last few years, the number of azole-resistant clinical isolates from the Trichophyton genus and the pool of infected patients who have failed to respond to the initial therapy for such infections have increased [2, 15, 22]. However, patient insensitivity to azole treatment has only been reported in cases caused by the anthropophilic dermatophyte T. rubrum [21]. In turn, azole-resistant phenotypes have never been found among zoophilic dermatophytes, particularly T. verrucosum.
In the present study, we evaluated the mechanisms underlying the sensitivity and resistance to fluconazole among clinical isolates of T. verrucosum obtained from human and animal dermatophytoses in Poland. The quantitative real-time PCR (qRT-PCR) technique was employed to examine the transcriptional modulation of ERG genes.
Materials and methods
Clinical dermatophyte isolates
In total, 25 clinical isolates of fluconazole-resistant Trichophyton verrucosum were obtained from humans (n = 10) and animals (n = 15) with typical symptoms of dermatomycosis from different regions of Poland (Table 1). The dermatophyte strains were isolated from outbreaks of zoophilic mycoses noted from 2011 to 2019. All human cases were documented as zoonoses. All of these isolates showed susceptibility to azoles, excluding fluconazole, and toward allylamine-type drugs. Additionally, five clinical isolates of T. verrucosum with sensitivity to fluconazole (MIC < 1 μg/ml) obtained from cattle with ringworm were used as controls.
Species identification analysis was performed as described previously by Gnat et al. [23]. All clinical isolates were identified to the species level by a combination of conventional and molecular techniques, comprising examination of macro- and micro-morphology (Fig. 1) and the internal-transcribed spacer (ITS) rDNA region sequencing with ITS1 (5′-TCCGTAGGTGAACCTGCGG-3′) and ITS4 (5′-TCCTCCGCTTATTGATATGC-3′) primer pairs [24]. All nucleotide sequences were deposited in the GenBank database (Table 1). Samples were maintained at − 80 °C in Sabouraud dextrose broth (SDB; Oxoid, Hampshire, UK) supplemented with 15% glycerol and stored in the Department of Veterinary Microbiology, University of Life Sciences in Lublin, Poland.
Micro- and macroscopic morphology of Trichophyton verrucosum after 20 days of incubation. A—Beige colonies with a friable texture and a yellow or yellow-orange reverse. The size of the colony in the range from 5 to 8 mm. The edges of the colony strongly corrugated and the image of the obverse and the reverse reminiscent of a cauliflower or a rotor of a turbine. B—Colony in a magnification × 10 (Olympus SZ61, Tokyo, Japan). C—The micromorphological image on the microscope slides exhibit circular chlamydospores. The macroconidia have not been observed (Olympus BX51, Tokyo, Japan, magnification × 400). D—The chlamydospores in a magnification × 1000
In vitro tests of susceptibility to the antifungal compounds were performed for all the tested isolates according to Clinical and Laboratory Standards Institute (CLSI) document M38, 3rd edition [25]. All chemicals used in this method were purchased from Sigma-Aldrich (Saint Louis, Missouri, USA), if not stated otherwise, and were of analytical grade. Stock solutions of the tested drugs were prepared in DMSO. Assay microtiter plates with 96 U wells were incubated for 72 h at 30 °C and then read spectrophotometrically using a Varioskan LUX multimode microplate reader (ThermoFisher) at the 530 nm wavelength (λ). The endpoint for the minimal inhibitory concentration (MIC80%) was the antifungal concentration at which prominent inhibition of growth, i.e., ≤ 80% of that of the control was observed. All isolates were tested in triplicate on different days. Prior to ERG gene expression analysis the inoculum, i.e., clinical isolates of T. verrucosum were grown for 14 days in Sabouraud glucose broth (BioMaxima, Lublin, Poland) at 25 °C. Next, they were homogenized and diluted to a final density of 1 × 103 to 3 × 103 CFU/ml. The suspension was exposed to fluconazole for 15 h (Sigma-Aldrich, Missouri, USA). The fluconazole was diluted in RPMI1640 medium (Sigma Aldrich, Missouri, USA) at the same concentration as the MIC determined for each sample. Drug-free controls were maintained for 15 h in the same medium without previous exposure to fluconazole.
RNA extraction and synthesis of cDNA
RNA extraction was performed using TRIzol reagent. Prior to the RNA extraction, 0.5 ml of TRIzol containing the fungal cell suspension was vortexed for 6 min. Afterwards, 0.2 ml of chloroform (Sigma-Aldrich, Missouri, USA) was added, mixed for 15 s, and centrifuged at 12,000 × g for 15 min at 4 °C. Next, the transparent phase was transferred to a clean tube containing 0.5 ml of isopropyl alcohol, and RNA was precipitated out of the mixture at − 20 °C overnight. The next day, the samples were centrifuged at 12,000 × g for 30 min at 4 °C. The precipitated RNA was washed twice with 1 ml of cold 70% ethyl alcohol and centrifuged at 12,000 g for 5 min at 4 °C. The samples were dried at room temperature in vacuum conditions and the RNA was dissolved in ultrapure RNase-free water. The total RNA was treated with DNase I (Sigma-Aldrich, Missouri, USA). The quantity and purity of the RNA were assessed both spectrophotometrically based on the A260/A280 and A260/A230 ratios measured by NanoDrop 2000 (Thermo Fisher scientific, USA) and visually during electrophoresis performed in 1% denaturing agarose gel. Two micrograms of the RNA obtained were reverse-transcribed into cDNA to a final volume of 40 μl using RevertAid™ Reverse Transcriptase (Thermo Fisher Scientific).
ERG gene expression
Quantitative reverse transcription PCR (qRT-PCR) was carried out with a StepOnePlus (Applied Biosystems, Foster City, USA) system using the SYBR Green GoTaq qPCR Master Mix (Promega, Madison, USA) to determine the relative expression levels of mRNA transcripts of ERG3, ERG6, and ERG11 genes in the fungal samples using the housekeeping gene GAPDH as a reference. All PCRs were performed as follows: denaturation at 95 °C for 10 min, followed by 40 cycles of 15 s at 95 °C, 30 s at 58 °C, and 30 s at 72 °C. The sequences of primers of the ERG3, ERG6, ERG11, and GAPDH genes used in the qRT-PCRs are given in Table 2. The primers were designed in the Primer 3 (http://simgene.com/Primer3) Web-based software using sequences available in NCBI (National Center for Biotechnology Information) database. The analyses of the gene expression were performed using the 2(−ΔΔCT) method of relative quantification [26].
Statistical analysis
The results are presented as the mean ± SEM. All values were compared using either Student’s t test or one-way ANOVA followed by Bonferroni’s post hoc test. Student’s t test was used to analyze ERG gene expression between fluconazole-sensitive and -resistant isolates. While ANOVA was applied to compare the relative expression of ERG3, ERG6 and ERG11 in resistant strains exposed to fluconazole. Statistical significance was considered when the p value was < 0.05. Statistical analyses were made using the R software (R Core Team, Auckland, New Zealand).
Results
The fluconazole-resistant T. verrucosum isolates used in this study showed MIC values ranging from 8 to 128 μg/ml (Table 1). The geometric mean MICs for the human and animal isolates were 44.8 μg/ml and 58.5 μg/ml, respectively. Tinea capitis (60%) and typical kerion celsi (30%) symptoms caused by fluconazole-resistant T. verrucosum were the most frequently reported disease in humans (Table 1). In turn, isolates with the highest MIC values for fluconazole (128 μg/ml) were detected in cattle and in one case of human kerion celsi. Cattle isolates with MIC values of fluconazole in the range of 0.25–0.5 μg/ml were used as control fluconazole-susceptible strains of T. verrucosum.
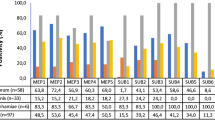
Additionally, based on the quantitative reverse transcription PCR (qRT-PCR), different levels of expression of the ERG3, ERG6, and ERG11 genes were shown in the pool of the clinical isolates of T. verrucosum. The exposure to fluconazole induced a higher level of ERG3, ERG6, and ERG11 gene expression in the cells of the resistant isolates compared to the sensitive ones (Fig. 2). However, despite these results, there was no statistical difference in the levels of ERG3 and ERG6 gene expression between the susceptible and resistant isolates after and without the exposure to fluconazole. The increase in the level of expression of these genes after the exposure to fluconazole was only 0.2 to 0.4 times higher in the phenotypically resistant isolates than in the susceptible ones. Interestingly, a statistically significant increase in the level of ERG11 gene expression was observed after and without the exposure to fluconazole in both the resistant and susceptible isolates. Moreover, the level of expression of the ERG11 gene in the fluconazole-resistant clinical isolates was statistically higher than for the ERG3 or ERG6 genes in these isolates.
ERG gene expression based on the antifungal susceptibility profile. Fungal samples of fluconazole-resistant and -sensitive Trichophyton verrucosum were grown for 14 days in Sabouraud glucose broth and exposed- or non-exposed to the drug for 15 h. The RNA was extracted and real-time PCR analysis was performed to evaluate mRNA expression of the ERG3 (a), ERG6 (b), and ERG11 (c) genes. GAPDH was used as a housekeeping gene. Ns no statistical significance and * statistical significance tested by Student’s t test between bars. ANOVA* statistical significance tested by ANOVA followed by Bonferroni’s post hoc test between relative expression of ERG3, ERG6, and ERG11 in resistant strains exposed to fluconazole
Discussion
The knowledge of the mechanisms that allow dermatophyte cells to develop drug resistance is essential for designing novel drugs and alternative therapeutic approaches [27]. This is even more important considering the infections by zoophilic dermatophytes in humans that are commonly noted in European countries [8, 28, 29], including recalcitrant dermatophytosis [27]. The frequency of infections caused by T. verrucosum varies from 3.1% in Spain, through 1.8% in Greece, and 1% in Poland to 0.01% in the Czech Republic [9, 30].
Ergosterol is a fundamental component of fungal cell membranes; therefore, many drugs, like polyenes, were designed to directly bind with this final product or, e.g., azoles, to disturb and inhibit the function of enzymes involved in its synthesis [14]. It is commonly known that the efficacy of azoles is the result of both the depletion of ergosterol in the plasma membrane and the subsequent accumulation of toxic sterol 14-α-methylated intermediates, ultimately leading to cell growth arrest [16, 19]. In consequence of perturbation of ergosterol biosynthesis during azole-treatment, the expression of some genes belonging to various sterol pathways are up or downregulated as a compensation mechanism [31, 32]. Lanosterol 14-α demethylase encoded by the ERG11 gene is the crucial enzyme [14]. An important role in this compensatory phenomenon in dermatophyte cells is also attributed to sterol C24-methyltransferase transferase and sterol C5-desaturase, i.e., products of ERG6 and ERG3 gene expression, respectively [31]. Here, we used the qRT-PCR method to determine the expression levels of three specified ERG genes to shed more light on the susceptibility and resistance to fluconazole of T. verrucosum strains. Since the mechanisms of azole resistance in zoophilic dermatophytes are not fully understood, compared to anthropophilic species, studies showing the resistance mechanism in T. verrucosum strains related to the expression of genes involved in ergosterol biosynthesis are particularly important and necessary.
Our study has demonstrated that exposure to fluconazole of T. verrucosum cells increased the expression levels of the ERG3, ERG6, and ERG11 genes both in the susceptible and resistant isolates. However, only the mRNA fold change in the ERG11 gene achieved a statistically significantly higher level in isolates with phenotypic resistance to fluconazole than in the case of the susceptible control strains. In literature, there are only fragmentary results regarding dermatophytes in this context. Diao et al. [31] found that the ERG11 gene was upregulated 13.55-fold in T. rubrum after exposure to itraconazole. Similar data obtained in investigations of isolates of Candida glabrata demonstrated that treatment with an azole drug, i.e., voriconazole, also increased the level of ERG3, ERG6, and ERG11 gene expression [33]. The ability of fluconazole to upregulate ERG11 gene expression in C. tropicalis, C. glabrata, and C. krusei cells was examined by many authors [33,34,35]. However, Alizadeh et al. [36] demonstrated differential expression of the ERG11 gene in resistant Candida albicans only after exposure to fluconazole. Nonetheless, ERG gene upregulation may play a role in azole resistance in fungi, regardless of the phylogenetic group.
It is worth noting that ERG11 is the most statistically significantly overexpressed ERG gene in the pool of fluconazole-resistant T. verrucosum isolates. Our studies show that the ERG3 and ERG6 genes seem to be less important than ERG11 for the phenomenon of azole resistance in zoophilic dermatophytes. These conclusions also raise some doubts in relation to research conducted on yeast-like fungi. Geber et al. [37] revealed that the regulatory mechanisms involved in the expression of these genes are interrelated, and deletion of ERG3 in Candida glabrata was associated with simultaneous upregulation of the ERG11 gene. Similarly, deletion of ERG11 was associated with ERG3 gene upregulation. Furthermore, in studies on Saccharomyces cerevisiae, Bammert and Fostel [38] documented global ERG gene upregulation in response to azole, terbinafine, and amorolfine treatment and mutations in the ERG1, ERG11, ERG6, ERG2, and ERG5 genes. However, Henry et al. [35] demonstrated that the increased ERG11 expression levels in drug-treated C. albicans quickly decreased upon removal of the antimycotic. In turn, Lopez-Ribot et al. [39] and Fernandes et al. [40] showed that, in the case of some species of azole-resistant Candida spp. treated with fluconazole, ERG genes are overexpressed but not genes encoding specific efflux pumps. In the study presented here, the participation of commonly known efflux pumps in the generation of the observed resistance phenotypes among T. verrucosum strains cannot be excluded. This seems very likely and will be the subject of further research, considering the fact that, unlike other azoles, fluconazole can be a good substrate for specific efflux pumps, as described elsewhere [41]. All the data presented here indicate a possible role of ERG genes in resistance to azole drugs, also in dermatophytes. However, this research should be continued.
Finally, our results have shown one of the possible mechanisms of resistance to fluconazole among T. verrucosum isolates, which involves the maintenance of high levels of expression of ERG genes after drug exposure. Further studies are necessary to determine whether other mechanisms also contribute to the maintenance and transmission of azole-resistant isolates. The gene expression profiling seems to be a useful tool for understanding the mechanisms of action and acquiring resistance to antifungal agents by zoophilic dermatophytes.
Data availability
All data generated or analyzed during this study are included in this published article.
References
de Hoog GS, Dukik K, Monod M et al (2017) Toward a novel multilocus phylogenetic taxonomy for the dermatophytes. Mycopathologia 182:5–31. https://doi.org/10.1007/s11046-016-0073-9
Gnat S, Łagowski D, Nowakiewicz A et al (2020) Population differentiation, antifungal susceptibility, and host range of Trichophyton mentagrophytes isolates causing recalcitrant infections in humans and animals. Eur J Clin Microbiol Infect Dis 39:2099–2113. https://doi.org/10.1007/s10096-020-03952-2
Begum J, Mir NA, Lingaraju MC et al (2020) Recent advances in the diagnosis of dermatophytosis. J Basic Microbiol 60:293–303. https://doi.org/10.1002/jobm.201900675
Bontems O, Fratti M, Salamin K et al (2020) Epidemiology of dermatophytoses in Switzerland according to a survey of dermatophytes isolated in Lausanne between 2001 and 2018. J Fungi 6:1–8. https://doi.org/10.3390/jof6020095
Taghipour S, Shamsizadeh F, Pchelin IM et al (2020) Emergence of Terbinafine Resistant Trichophyton mentagrophytes in Iran, Harboring mutations in the Squalene Epoxidase (SQLE) gene. Infect Drug Resist 13:845–850. https://doi.org/10.2147/IDR.S246025
Das S, De A, Saha R et al (2020) The current Indian epidemic of dermatophytosis: a study on causative agents and sensitivity patterns. Indian J Dermatol 65:118–122. https://doi.org/10.4103/ijd.IJD_203_19
Łagowski D, Gnat S, Nowakiewicz A, Osińska M (2020) Comparison of in vitro activities of 11 antifungal agents against Trichophyton verrucosum isolates associated with a variety hosts and geographical origin. Mycoses 63:294–301. https://doi.org/10.1111/myc.13042
Łagowski D, Gnat S, Nowakiewicz A et al (2019) In search of the source of dermatophytosis: epidemiological analysis of Trichophyton verrucosum infection in llamas and the breeder (case report). Zoonoses Public Health 66:982–989. https://doi.org/10.1111/zph.12648
Gnat S, Łagowski D, Nowakiewicz A et al (2018) Infection of Trichophyton verrucosum in cattle breeders, Poland: a 40-year retrospective study on the genomic variability of strains. Mycoses 61:681–690. https://doi.org/10.1111/myc.12791
Courtellemont L, Chevrier S, Degeilh B et al (2017) Epidemiology of Trichophyton verrucosum infection in Rennes University Hospital, France: a 12-year retrospective study. Med Mycol 55:720–724. https://doi.org/10.1093/mmy/myw142
Sahu P, Dayal S, Mawlong P et al (2020) Tinea corporis bullosa secondary to Trichophyton verrucosum: a newer etiological agent with literature review. Indian J Dermatol 65:76–78. https://doi.org/10.4103/ijd.IJD_483_19
Gnat S, Łagowski D, Nowakiewicz A et al (2021) Complementary effect of mechanism of multidrug resistance in Trichophyton mentagrophytes isolated from human dermatophytoses of animal origin. Mycoses 64:537–549. https://doi.org/10.1111/myc.13242
Overgaauw PAM, Avermaete KHA, va., Mertens CARM, et al (2017) Prevalence and zoonotic risks of Trichophyton mentagrophytes and Cheyletiella spp. in guinea pigs and rabbits in Dutch pet shops. Vet Microbiol 205:106–109. https://doi.org/10.1016/j.vetmic.2017.05.008
Khurana A, Sardana K, Chowdhary A (2019) Antifungal resistance in dermatophytes: recent trends and therapeutic implications. Fungal Genet Biol 132:103255. https://doi.org/10.1016/j.fgb.2019.103255
Rudramurthy SM, Shankarnarayan SA, Dogra S, et al (2018) Mutation in the squalene epoxidase gene of Trichophyton interdigitale and Trichophyton rubrum associated with allylamine resistance. Antimicrob Agents Chemother 62. https://doi.org/10.1128/AAC.02522-17
Łagowski D, Gnat S, Nowakiewicz A (2020) Mechanisms of dermatophyte resistance to antifungal substances. Postępy Mikrobiol - Adv Microbiol 59:153–165. https://doi.org/10.21307/pm-2020.59.2.012
Verma SB, Bishnoi A, Vinay K, Dogra S (2018) Emergence of recalcitrant dermatophytosis in India. Lancet Infect Dis 18:250–251. https://doi.org/10.1016/S1473-3099(18)30079-3
White TC, Marr KA, Bowden RA (1998) Clinical, cellular, and molecular factors that contribute to antifungal drug resistance. Clin Microbiol Rev 11:382–402. https://doi.org/10.1128/cmr.11.2.382
Ghannoum MA, Rice LB (1999) Antifungal agents: mode of action, mechanisms of resistance, and correlation of these mechanisms with bacterial resistance. Clin Microbiol Rev 12:501–517. https://doi.org/10.1128/cmr.12.4.501
Sanglard D (2016) Emerging threats in antifungal-resistant fungal pathogens. Front Med 3:11. https://doi.org/10.3389/fmed.2016.00011
Monod M, Feuermann M, Salamin K et al (2019) Trichophyton rubrum azole resistance mediated by a new ABC transporter, TruMDR3. Antimicrob Agents Chemother 63:e00863-e919. https://doi.org/10.1128/AAC.00863-19
Khurana A, Sardana K (2018) Reinterpreting minimum inhibitory concentration (MIC) data of itraconazole versus terbinafine for dermatophytosis - time to look beyond the MIC data? Indian J Dermatol Venereol Leprol 84:61–62. https://doi.org/10.4103/ijdvl.IJDVL_715_17
Gnat S, Łagowski D, Nowakiewicz A, Zięba P (2018) Tinea corporis by Microsporum canis in mycological laboratory staff: unexpected results of epidemiological investigation. Mycoses 61:945–953. https://doi.org/10.1111/myc.12832
White TJ, Bruns TD, Lee SB and Taylor JW (1990) Amplification and Direct Sequencing of Fungal Ribosomal RNA Genes for Phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ and White TJ, Eds, PCR Protocols: A Guide to Methods and Applications, Academic Press, New York, pp 315–322. https://doi.org/10.1016/B978-0-12-372180-8.50042-1
CLSI (2017) Reference method for broth dilution antifungal susceptibility testing of filamentous fungi. CLSI standard M38., 3rd ed. Clinical and Laboratory Standards Institute, Wayne ,PA
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Gnat S, Łagowski D, Nowakiewicz A (2020) Major challenges and perspectives in the diagnostics and treatment of dermatophyte infections. J Appl Microbiol 129:212–232. https://doi.org/10.1111/jam.14611
Gnat S, Łagowski D, Nowakiewicz A, Dyląg M (2020) Tinea corporis caused by Trichophyton equinum transmitted from asymptomatic dogs to two siblings. Brazilian J Microbiol 51:1433–1438. https://doi.org/10.1007/s42770-019-00204-0
Dworecka-Kaszak B, Bieganska MJ, Dabrowska I (2020) Occurrence of various pathogenic and opportunistic fungi in skin diseases of domestic animals: a retrospective study. BMC Vet Res 16:248. https://doi.org/10.1186/s12917-020-02460-x
Pereiro Miguens M, Pereiro M, Pereiro M (1991) Review of dermatophytoses in Galicia from 1951 to 1987, and comparison with other areas of Spain. Mycopathologia 113:65–78. https://doi.org/10.1007/BF00442412
Diao Y, Zhao R, Deng X et al (2009) Transcriptional profiles of Trichophyton rubrum in response to itraconazole. Med Mycol 47:237–247. https://doi.org/10.1080/13693780802227308
Petrucelli MF, Matsuda JB, Peroni K et al (2019) The transcriptional profile of Trichophyton rubrum co-cultured with human keratinocytes shows new insights about gene modulation by terbinafine. Pathogens 8:274. https://doi.org/10.3390/pathogens8040274
Rodrigues CF, Gonçalves B, Rodrigues ME et al (2017) The effectiveness of voriconazole in therapy of Candida glabrata’s biofilms oral infections and its influence on the matrix composition and gene expression. Mycopathologia 182:653–664. https://doi.org/10.1007/s11046-017-0135-7
Rex JH, Nelson PW, Paetznick VL et al (1998) Optimizing the correlation between results of testing in vitro and therapeutic outcome in vivo for fluconazole by testing critical isolates in a murine model of invasive candidiasis. Antimicrob Agents Chemother 42:129–134. https://doi.org/10.1128/aac.42.1.129
Henry KW, Nickels JT, Edlind TD (2000) Upregulation of ERG genes in Candida species by azoles and other sterol biosynthesis inhibitors. Antimicrob Agents Chemother 44:2693–2700. https://doi.org/10.1128/AAC.44.10.2693-2700.2000
Alizadeh F, Khodavandi A, Zalakian S (2017) Quantitation of ergosterol content and gene expression profile of ERG11 gene in fluconazole-resistant Candida albicans. Curr Med Mycol 3:13–19. https://doi.org/10.29252/cmm.3.1.13
Geber A, Hitchcock CA, Swartz JE et al (1995) Deletion of the Candida glabrata ERG3 and ERG11 genes: effect on cell viability, cell growth, sterol composition, and antifungal susceptibility. Antimicrob Agents Chemother 39:2708–2717. https://doi.org/10.1128/AAC.39.12.2708
Bammert GF, Fostel JM (2000) Genome-wide expression patterns in Saccharomyces cerevisiae: comparison of drug treatments and genetic alterations affecting biosynthesis of ergosterol. Antimicrob Agents Chemother 44:1255–1265. https://doi.org/10.1128/AAC.44.5.1255-1265.2000
Lopez-Ribot JL, Mcatee RK, Lee LN et al (1998) Distinct patterns of gene expression associated with development of fluconazole resistance in serial Candida albicans isolates from human immunodeficiency virus-infected patients with oropharyngeal candidiasis. Antimicrob Agents Chemother 42:2932–2937. https://doi.org/10.1128/aac.42.11.2932
Fernandes T, Silva S, Henriques M (2016) Effect of voriconazole on Candida tropicalis biofilms: relation with ERG genes expression. Mycopathologia 181:643–651. https://doi.org/10.1007/s11046-016-0023-6
Wang EJ, Lew K, Casciano CN et al (2002) Interaction of common azole antifungals with P glycoprotein. Antimicrob Agents Chemother 46:160–165. https://doi.org/10.1128/AAC.46.1.160-165.2002
Author information
Authors and Affiliations
Contributions
Conceptualiation: Sebastian Gnat; methodology: Sebastian Gnat and Dominik Łagowski; investigation: Dominik Łagowski, Aneta Nowakiewicz, and Mariusz Dyląg; writing—original draft preparation: Sebastian Gnat; supervision: Sebastian Gnat and Aneta Ptaszyńska.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Responsible Editor: Carlos Pelleschi Taborda
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gnat, S., Łagowski, D., Dyląg, M. et al. Modulation of ERG gene expression in fluconazole-resistant human and animal isolates of Trichophyton verrucosum. Braz J Microbiol 52, 2439–2446 (2021). https://doi.org/10.1007/s42770-021-00585-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-021-00585-1