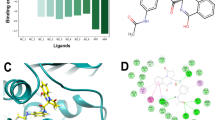
Abstract
For any antimicrobial assay, a standard drug is used to compare the bactericidal efficiency of the bioactive compound under screening. The standard drugs have different targets that may be intracellular or membrane located. The location of the target is believed to be determining the bioactivity of the drug depending on the drug's access to its target. Therefore, different drugs must have a different magnitude in exhibiting the biological effect. However, in most of the published literature about the screening of bioactive compounds on antimicrobial activity, generally, the standard drug is randomly chosen while comparing against the bioactive compound of interest. Further, the antimicrobial activity is inferred by comparing the randomly chosen standard drugs without knowing the physicochemical parameters of the standard drug and the test molecule. It is just like an unfair comparison of the impact of a bullet with the impact of an explosive in a combat scene. Computer-based strategies for structure-based drug discovery presents a valuable alternative to the costly and time-consuming process of random screening. The docking studies provide better insights into the binding mechanism of substrate and inhibitor at the molecular level. The evaluation of such a comparison of bioactive compounds against randomly selected standard drugs through a customized virtual screening pipeline showed 57% false positives, 18% true positive, 17% true negative, 8% false-negative results. This study directs for mandatory cheminformatics-based assessment of the bioactive compounds before choosing the standard drug to compare with.










Similar content being viewed by others
References
Bagchi B, Kar S, Dey SK, Bhandary S, Roy D, Mukhopadhyay TK, Das S, Nandy P (2013) In situ synthesis and antibacterial activity of copper nanoparticle loaded natural montmorillonite clay based on contact inhibition and ion release. Colloids Surf B Biointerfaces 108:358–365
Biswas SK, Chowdhury A, Raihan SZ, Muhit MA, Akbar MA, Mowla R (2012) Phytochemical investigation with assessment of cytotoxicity and antibacterial activities of chloroform extract of the leaves of Kalanchoe plnnata. Am J Plant Physiol 7(1):41–46
Campoli-Richards DM, Monk JP, Price A, Benfield P, Todd PA, Ward A (1988) Ciprofloxacin. Drugs 35(4):373–447
Chung PY, Chung LY, Navaratnam P (2014) Potential targets by pentacyclic triterpenoids from Callicarpa farinosa against methicillin-resistant and sensitive Staphylococcus aureus. Fitoterapia 94:48–54
Clark DE (2008) What has virtual screening ever done for drug discovery? Expert Opin Drug Discov 3(8):841–851
Duch W, Swaminathan K, Meller J (2007) Artificial intelligence approaches for rational drug design and discovery. Curr Pharm Des 13(14):1497–1508
Ghazal SA, Abuzarqa M, Mahasneh AM (1992) Antimicrobial activity of Polygonum equisetiforme extracts and flavonoids. Phytother Res 6(5):265–269
Hara K, Kohno S, Kadota J-I, Tomono K, Hirakata Y, Maesaki S, Nakatomi M, Asai S, Mizukane R, Okuno K, Fukushima K, Itoh N, Inoue Y, Koike T, Onishi K, Omichi M, Yamada G, Hiraga Y, Watanabe A, Utsunomiya Y (1997) Clinical evaluation of ciprofloxacin for bacterial pneumonia—phase III comparative study of intravenous ciprofloxacin versus ceftazidime. J Jap Soc Chemother 45:901–922. https://doi.org/10.11250/chemotherapy1995.45.901
Hughes JP, Rees S, Kalindjian SB, Philpott KL (2011) Principles of early drug discovery. Br J Pharmacol 162(6):1239–1249
Isitua CC, Ibeh IN, Olayinka JN (2016) Antibacterial activity of Moringa oleifera Lam leaves on enteric human pathogens. Indian J Appl Res 6(8):553–557
Jorgensen WL (2009) Efficient drug lead discovery and optimization. Acc Chem Res 42(6):724–733
Kraus CH (2008) Low hanging fruit in infectious disease drug development. Curr Opin Microbiol 11:434–438
Kuzu OF, Gowda R, Sharma A, Robertson GP (2014) Leelamine mediates cancer cell death through inhibition of intracellular cholesterol transport. Mol Cancer Ther 13(7):1690–1703
Lage OM, Ramos MC, Calisto R, Almeida E, Vasconcelos V, Vicente F (2018) Current screening methodologies in drug discovery for selected human diseases. Mar Drugs 16(8):279
Lee JA, Uhlik MT, Moxham CM, Tomandl D, Sall DJ (2012) Modern phenotypic drug discovery is a viable, neoclassic pharma strategy. J Med Chem 55:4527–4538
Mahesh B, Satish S (2008) Antimicrobial activity of some important medicinal plants against plant and human pathogens. World J Agric Sci 4:839–843
Masadeh MM, Alzoubi KH, Al-Azzam SI, Khabour OF, Al-Buhairan AM (2016) Ciprofloxacin-induced antibacterial activity is atteneuated by pretreatment with antioxidant agents. Pathogens 5(1):28
Rashid MA, Gustafson KR, Boyd MR (2001) New cytotoxic N-nethylated β-carboline alkaloids from the marine Ascidian Eudistoma g ilboverde. J Nat Prod 64(11):1454–1456
Siddiqui SA, Rahman A, Rahman MO, Akbar MA, Ali MA, Al-Hemaid FM, Elshikh MS, Farah MA (2019) A novel triterpenoid 16-hydroxy betulinic acid isolated from Mikania cordata attributes multi-faced pharmacological activities. Saudi Jo Biol Sci 26(3):554–562
Spížek J, Novotná J, Řezanka T, Demain AL (2010) Do we need new antibiotics? The search for new targets and new compounds. J Ind Microbiol Biotechnol 37(12):1241–1248
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31(2):455–461
Ulloora S, Kumar S, Shabaraya R, Adhikari AV (2013) New dihydropyridine derivatives: anti-inflammatory, analgesic and docking studies. Med Chem Res 22(4):1549–1562
Wheeler G, Field R, Tomlinson M (2012) Phenotypic screens with model organisms. In: Press CU (ed) Chemical genomics. Cambridge University Press, New York, pp 121–136
Wu G, Robertson DH, Brooks CL III, Vieth M (2003) Detailed analysis of grid-based molecular docking: a case study of CDOCKER—a CHARMm-based MD docking algorithm. J Comput Chem 24(13):1549–1562
Yang G, Yue J, Gong X, Qian B, Wang H, Deng Y, Zhao Y (2014) Blueberry leaf extracts incorporated chitosan coatings for preserving post harvest quality of fresh blueberries. Postharvest Biol Technol 92:46–53
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kumar, H.S.S., Kumar, S.R., Kumar, N.N. et al. Molecular docking studies of gyrase inhibitors: weighing earlier screening bedrock. In Silico Pharmacol. 9, 2 (2021). https://doi.org/10.1007/s40203-020-00064-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s40203-020-00064-9