Abstract
Since the outbreak of the novel coronavirus disease 2019 (COVID-19) in Wuhan, China, many health care systems have been heavily engaged in treating and preventing the disease, and the year 2020 may be called as “historic COVID-19 vaccine breakthrough”. Due to the COVID-19 pandemic, many companies have initiated investigations on developing an efficient and safe vaccine against the virus. From Moderna and Pfizer in the United States to PastocoVac in Pasteur Institute of Iran and the University of Oxford in the United Kingdom, different candidates have been introduced to the market. COVID-19 vaccine research has been facilitated based on genome and structural information, bioinformatics predictions, epitope mapping, and data obtained from the previous developments of severe acute respiratory syndrome coronavirus (SARS-CoV or SARS-CoV-1) and middle east respiratory syndrome coronavirus (MERS-CoV) vaccine candidates. SARS-CoV genome sequence is highly homologous to the one in COVID-19 and both viruses use the same receptor, angiotensin-converting enzyme 2 (ACE2). Moreover, the immune system responds to these viruses, partially in the same way. Considering the on-going COVID-19 pandemic and previous attempts to manufacture SARS-CoV vaccines, this paper is going to discuss clinical cases as well as vaccine challenges, including those related to infrastructures, transportation, possible adverse reactions, utilized delivery systems (e.g., nanotechnology and electroporation) and probable vaccine-induced mutations.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Being considered the newest addition to the Coronaviridae family, severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) is the virus responsible for the coronavirus disease 2019 (COVID-19), which was declared a pandemic on March 11, 2020, by the World Health Organization (WHO) [1]. The term ‘Corona’ represents crown-like spikes on the outer surface of the coronaviruses, which contain the largest genomes among all the RNA viruses [2, 3]. Up to July 16, 2022, the number of confirmed COVID-19 cases were 566,641,094 including 6,386,256 deaths and 537,853,204 recoveries [4].
Based on the recent WHO updates, the most common symptoms of COVID-19 are fever, cough, tiredness, and loss of taste/smell [5]. The less common ones are sore throat, headache, aches and pains, diarrhea, rashes, or discoloration of fingers or toes, and red or irritated eyes. The disease is assessed into mild, severe, and critical (i.e., respiratory failure, septic shock, multiple organ dysfunction, or failure) categories based on the clinical manifestations and severity [6, 7].
Apart from the health-related complications of the disease, its economic burden cannot be ignored. It was reported that global gross domestic product (GDP) dropped about 4.5% in 2020 [8]. Numerous people who have found themselves jobless and lost their insurance have faced a grave situation. The average cost for hospital care for COVID-19 patients ranges from 51,000 to 78,000 USD based on their age [9]. The only similar conditions we have experienced in the twenty-first century are the Middle East respiratory syndrome (MERS) and severe acute respiratory syndrome coronavirus (SARS-CoV) outbreaks in 2012 and 2003, respectively. Although SARS-CoV, MERS-CoV, and SARS-CoV-2 belong to the same coronaviruses genus, SARS-CoV-2 is associated with milder infections. This explains why SARS-CoV-2 spreads far easier [10].
Herd immunity occurs when enough of a population is protected against an infection that its transmission ceases, indirectly safeguard those who are not immune. The herd immunity threshold for COVID-19 is at least 60–70% [11]. So far, global vaccination seems to be the only weapon in providing such conditions. In taking advantage of the high genetic similarity between SARS-CoV and SARS-CoV-2 (i.e., 79.6%), various vaccine candidates from nearly all over the world have been introduced throughout the pandemic, with Pfizer’s “BNT162b1” and Modena’s “mRNA-1273” being the forerunners in acquiring USFDA’s authorization for emergency use [12]. From live attenuated or inactivated viruses, viral vectors, and virus-like particles (VLPs) to recombinant proteins and nucleic acids (RNAs and DNAs), diverse platforms have been used in the SARS-CoV-2 vaccine manufacturing process [13, 14]. Each platform has its pros and cons in fields such as storage conditions, price, side effects, efficacy, and safety. It is also affected by multiple factors, one of the most important of which is the issue of mutations.
Considering the importance of proper vaccination and the challenges mentioned above, it is necessary to thoroughly identify all available SARS-CoV-2 vaccines by reviewing their mechanism of action, delivery systems, clinical efficacy, side effects, and possible vaccine-induced mutations. These issues will be comprehensively discussed in the present article, aiming to better the global vaccination process and policymaker decisions.
A comparison between SARS-CoV and SARS-CoV-2
SARS-CoV is a virus responsible for the epidemic that started in 2003 and ended shortly after, resulting in only a few vaccines starting phase 1 clinical trial and other vaccines staying at the pre-clinical phase. Considering this issue, comparing SARS-CoV with SARS-CoV-2 is almost unfeasible; however, lessons regarding SARS-CoV vaccines can be taken to enhance the safety and efficacy of SARS-CoV-2 vaccines. Since a better understanding of SARS-CoV-2 leads to improved vaccine manufacture, this section provides information on this virus and its structure [15].
The phylogenetic tree analysis of SARS-CoV-2 shows that, like SARS-CoV, it belongs to a different clade from MERS-CoV [10]. Based on the genomic sequence comparison, SARS-CoV-2 shares about 79.6% and 50% overall genomic similarity with SARS-CoV and MERS-CoV, respectively [13]. While SARS-CoV and SARS-CoV-2 share their entry receptor, angiotensin-converting enzyme 2 (ACE2), MERS-CoV uses dipeptidyl peptidase (DPP)-4 [16]. Regarding their structural proteins (Fig. 1), spike glycoprotein (S) on the particle's surface consists of two subunits that can bind to the cellular receptors and mediate infection, following which the cell begins replicating in the cytoplasm. Membrane (M) protein increases the membrane curvature, promoting the viral assembly. Envelope (E) protein is involved mainly in numerous functions in the viral replication cycle, such as assembly, release, and pathogenesis. Nucleocapsid (N) protein inhibits interferon (IFN) and plays a significant role in virus transcription and assembly [3, 17, 18]. The results showed 76%, 90.1%, 90.6%, and 94.7% similarities between S, M, N, and E proteins of SARS-CoV and SARS-CoV-2, respectively [19]. Furthermore, some similarities and differences exist between SARS-CoV and SARS-CoV-2 regarding receptor binding domain, host cell entry, and protease activation [20]. SARS-CoV and SARS-CoV-2 bind to the ACE2 receptor of human cells in many tissues, including the lungs, kidneys, heart, and testis, by their receptor-binding domain (RBD) in the S1 subunit [21]. Then, their envelope and the host cell membrane fuse to release the viral nucleocapsid into the target cells [22]. To this end, the S protein should be activated, so the S2 subunit (cleaved from the S1 subunit by host cell proteases) assists the fusion process and transports the virion into the host cells [18].
By comparing the full-length S protein sequences, the most probable alterations were found on the S1 subunit suggesting that the neutralizing antibodies that were once effective against SARS-CoV might not offer protection against SARS-CoV-2. Although cross-reactivity between the antibodies seems to be cross-reactivity, cross-neutralization appears to be relatively rare [10, 23].
SARS-CoV-2 binding affinity is 10–20 times higher than SARS-CoV, causing a more efficient cell entry. SARS-CoV-2 RBD, despite its higher affinity, is possibly less exposed than SARS-CoV RBD. This may be due to the “lying-down” position of its RBD, which can lead to ineffective receptor binding compared to the “standing-up” state in SARS-CoV [18]. On the other hand, the immune evasion caused by this position contributes to the conformational masking strategies [18, 20, 23].
A protein sequence alignment analysis was performed to look deeper into the encoded proteins of SARS-CoV-2 and SARS-CoV. Most of them were substantially homologous (95%–100%) and two SARS-CoV-2 proteins (ORF8 and ORF10) had no counterparts in SARS-CoV, making it clinically meaningful to analyze the biological function of these two specific proteins [3].
SARS-CoV and SARS-CoV-2 interactions with the immune system
Pathophysiology of SARS-CoV-2 and SARS-CoV infections closely resemble each other, with aggressive inflammatory responses heavily involved in damage to the airways. Put differently, the viral infection rate and the host response are two contributory factors in this regard [24]. SARS-CoV-2 enters the body through the nasal-oral cavity [25]. Once inhaled, the virus primarily binds to the host cells through its target receptor. Earlier work on SARS-CoV demonstrated that the virus targets cells which express ACE2, such as airway epithelial cells, alveolar epithelial cells, vascular endothelial cells, and macrophages in the lungs. Since SARS-CoV-2 uses the same receptor, these cells are likely to be targeted [24].
SARS-CoV-2, as a cytopathic virus and as part of its replicative cycle, prompts death and injury of virus-infected cells and tissues, which in the airway epithelial cells can lead to elevated levels of virus-linked pyroptosis (a highly inflammatory form of programmed cell death that is frequently seen with cytopathic viruses). This probably triggers an inflammatory response. The released pathogen-associated molecular patterns (PAMPs) such as viral RNA, and damage-associated molecular patterns (DAMPs), including ATP and DNA, are detected by alveolar epithelial cells and alveolar macrophages, which use a variety of pattern-recognition receptors (PRRs). Subsequently, local inflammation occurs following hypersecretion of pro-inflammatory cytokines and chemokines, including interleukin (IL)-6, interferon gamma-induced protein (IP)-10, macrophage inflammatory protein 1α (MIP1α), MIP1β and monocyte chemoattractant protein 1 (MCP1); all of which parallel the observations in SARS-CoV. This results in the recruitment of immune cells, notably monocytes and T lymphocytes but not neutrophils, into the infected location, provoking further inflammation [24, 26].
In a healthy immune response, the primary inflammation caused by either SARS-CoV or SARS-CoV-2 attracts virus-specific T cells to the site of infection to eradicate the infected cells before the virus spreads and ultimately lead to minimal lung damage, clearance of the virus, and lastly, recovery. However, it has been reported that alveolar dysfunction in two COVID-19 cases led to cytokine storm and failure of multiple organs, including the lungs and heart [27].
In a dysfunctional immune response, the primary inflammation may lead to the gathering of immune cells in the lungs and the overproduction of pro-inflammatory cytokines (also known as a cytokine storm). This causes acute respiratory disease syndrome (ARDS) and eventually lung damage. Patients with severe COVID-19 required intensive care in hospitals displayed higher blood plasma levels of IL-2, IL-7, IL-10, IP-10, MCP1, MIP1α, and tumor necrosis factor (TNF). IL-6 level in these patients continued to increase over time and was higher in non-survivors than survivors. Furthermore, patients with severe disease, compared to patients with a mild infection, showed a significantly higher percentage of CD14 + and CD16 + inflammatory monocytes in peripheral blood. These cells secrete inflammatory cytokines that help the cytokine storm (IP-10, MCP1, and MIP1α) [24, 25]. Subsequently, the cytokine storm might affect other organs, leading to multi-organ damage. For instance, elevated levels of cytokines such as TNF were reported to cause septic shock, myocardial injury, and circulatory failure in some patients, especially those over 60 years of age. Furthermore, non-neutralizing antibodies produced by B cells may boost SARS-CoV-2 infection through antibody-dependent enhancement (ADE), further worsening organ damage [24].
In SARS-CoV-2 infection, production of type I and III interferons decreases, leading to an overall reduction in the transcription of antiviral genes [25]. Research on SARS-CoV found that several viral structural and non-structural proteins can antagonize interferon responses, thus, eluding the immune system. Coronaviruses are implied to possess the ability to escape from immune detection and curb human immune responses, which somewhat explains why they usually have a more extended incubation period [16]. Antagonism happens at different stages of the interferon signaling pathway, including pattern-recognition receptor (PRR) signaling through TNF receptor-associated factor 3 (TRAF3), interferon regulatory factor 3 (IRF3), and other molecules, downstream interferon signaling through STAT1, PRR recognition of viral RNA and host mRNA degradation and host protein translation. Some of these pathways are present in SARS-CoV-2 [24]. SARS-CoV-2 infection can also decrease T cells and enhance the exhaustion of effector T cells, thereby reducing the immune response against the virus. This exhaustion results from a higher expression of inhibitory receptors on the cell’s surface, which is influenced by cytokines like IL-6, IL-10, and TNF-α [25, 28,29,30,31].
Following the viral clearance of either SARS-CoV or SARS-CoV-2, a group of memory T cells is produced to encounter re-infection. Re-stimulated CD4 + memory T cells activate B cells and other immune cells by cytokine production, while cytotoxic memory T cells assist in eliminating the infected cells during a future infection. In addition to this mechanism, studies on SARS-CoV revealed that produced antibodies were present in the blood for at least six months to two years. According to a study in China, 93.88% of patients after one year and 89.58% after two years tested positive for IgG against SARS-CoV [31]. Whether these results can be extrapolated to all COVID-19 patients worldwide remains unknown and requires further investigations [25].
How antibodies decay is found to vary by individuals, target antigen, antibody isotype, and assay used in studies; anti-N antibodies are the fastest to decrease, followed by anti-RBD and anti-S antibodies [32,33,34]. The half-life of anti-S antibodies has been estimated to be 126–238 days in different studies [35,36,37]. Data are showing that SARS-CoV-2-specific IgG was detectable in 90% of seroconversions up to one year post-infection [34, 38].
Factors affecting the severity of the infection
During the current pandemic, numerous studies have addressed the challenges with vaccine efficacy. Some challenges are related to the virus, while others are linked to epidemiological conditions, vaccine platforms, and gender. Most efforts have been focused on stimulating neutralizing antibody production, route of administration, dosage, and injection intervals [39]. The physiological heterogeneity among different individuals plays a critical role in vaccine efficacy [40]. Moreover, factors such as eligibility for all age groups, genetics, gender, ethnicity, history of former infection, and the emergence of new variants of concern (VOC) are considered obstacles to vaccine production [41]. Although results are conflicting regarding the impact of the different factors on the severity of the disease and vaccine responses, a brief explanation of how such issues can affect the overall mortality and morbidity from SARS-CoV and SARS-CoV-2 is provided below.
Genetics
Results from previous studies regarding the effect of host genetic factors on the severity of SARS-CoV are crucial to be used in the SARS-CoV-2 pandemic [42,43,44], which are discussed as follows.
SARS-CoV
According to an in vivo study, there is a locus on chromosome 3 (including 23 genes and 13 non-coding RNAs), which contributes to SARS-CoV vascular cuffing and inflammation of the lungs [45]. Genetic polymorphism in those genes is associated with different clinical manifestations in patients. Moreover, human cyclophilin A, a peptidyl-prolyl isomerase, contributes to the viral core sequestration. This protein binds to the N protein of the virus and interacts with the coronavirus proteins and genome. Single nucleotide polymorphism in the cyclophilin A gene contributes to heterogeneity of COVID-19 severity in different individuals [45]. It is also suggested that ACE2 receptor polymorphism brings about varieties within individuals [45, 46]. The severity of the infection and susceptibility to SARS are affected by a particular polymorphism of the CLEC4M gene (in the variable tandem repeats in exon) since L-SIGN, the coded protein of this gene is a receptor for the virus [45, 47]. Diverse quantities in mannose-binding lectin (MBL) may affect protection against SARS. For example, a lower amount of MLB worsened the severity of SARS [46, 47]. It was also reported that Fc gamma RIIA-R/R131 genotype polymorphism, which is in the human Fc gamma receptor IIA gene, was related to the severity of SARS infection [45]. Polymorphism of IFNγ + 874A and RANTES-28G alleles, MIF gene (on influenza), and human leukocyte antigen (HLA) also contributed to SARS' severity and mortality rate. Interactions of all these variables are contentious issues that need further investigation [45,46,47].
SARS-CoV2
ACE2
ACE2 polymorphisms, such as p.Arg514Gly in the African/African-American population, were linked to cardiovascular and pulmonary disorders by altering the interaction between angiotensinogen and ACE2 receptor [48]. It has been reported that higher expression of ACE2 receptors positively correlated with the severity of the SARS-CoV-2 infection [45]. Since ACE2 is the key receptor for SARS-CoV-2, individuals with diabetes, hypertension, and chronic obstructive pulmonary disease [45] would be more susceptible to SARS-CoV-2 infection. Close contact is essential for recognizing RBD. Changes in ACE2 residues at the binding interface affect affinity, one of the most important determinants of host sensitivity [49]. hACE2 K353 and K31 are the significant areas that form hydrogen bonds with the backbones of N501 and Q493, respectively, in the receptor-binding motif and play a role in the tight binding of the SARS-CoV-2 S protein [50]. The whole-genome sequencing of 1200 individuals and chip genotyping of more than 15,000 participants found two observed missense variants of ACE2, K26R, and S331F, which are responsible for reducing the receptor affinity of the viral Spike protein [51]. It has been discovered that 14 ACE2 polymorphisms with increased susceptibility (I21V, E23K, K26R, N64K, T92I, Q102P, D206G, G211R, R219C, E329G, H378R, V447F, A501T, and N720D) had greater allele frequencies in European (non-Finnish) groups than in East Asian populations. Two resistance-related ACE2 polymorphisms (E35K and F72V) show greater allele frequencies in East Asian groups but are low or absent in European (non-Finnish) populations. These findings are consistent with the pandemic scenario and could help explain why COVID-19 prevalence and fatality rates in Europe and East Asia are so different [49]. Due to these findings, genetic variations in the ACE2 gene among individuals are remarkable factors that affect disease severity [51].
Inflammatory factors
Higher amounts of ferritin, D-dimer, C-reactive protein, and increased levels of IL-2, IL-6, IL-10, and TNF-α have been associated with the severity and mortality rates of SARS-CoV-2 infection [45]. Some reports indicated that polymorphisms of rs1800896 in IL-10, rs2275913 in IL-17A, and rs763780 loci in the IL-17F gene were correlated with death rates in different countries [52]. However, in Northwestern Mexico, there was no association between the rs1800871 and rs1800872 polymorphisms and COVID-19 severity and mortality [53]. To find an association between polymorphism of TNF-α gene and COVID-19 severity, a study showed that the presence of TNF-α polymorphism affected people's sensitivity to COVID-19 infection [54].
Transmembrane protease, serine 2 (TMPRSS2)
TMPRSS2 is a serine protease in various human tissues and is involved in SARS-CoV-2 infection. Genetic variations on this protein influence virus clearance in the host and predisposition to infection. Generally, polymorphisms at TMPRSS2 RS2070788, RS7364083, and RS9974589 are crucial to consider. Also, the expression of TMPRSS2 is increased with the polymorphism at TMPRSS2 rs8134378 in men and favors the fusion of the virus membrane [45, 46, 55].
HLA loci
Numerous studies stated the association between SARS-CoV-2 and HLA gene [47]. One study found that the HLA-A*24:02 allele influences susceptibility to COVID-19. Moreover, HLA-B*46:01 affects and enervates the immune system, thus enhancing the severity of the coronavirus infection in the Asian population. Due to some observation, repetition, and conservation of HLA-B*15:03 between various viruses, one can assume that this allele somewhat protects coronaviruses [45]. New outcomes suggest that individuals with HLA-A*11:01, HLA-A*02:06, or HLA- B*54:01 are probably preserved from SARS-CoV-2 infection [56]. Other studies on SARS-CoV and MERS-CoV reported that several HLA genotypes are associated with susceptibility or resistance, which include HLA- B*07:03, HLA-B*46:01, HLA-C*08:01, HLA-C*15:02, HLA-DRB1*03:01, HLA-DRB1*11:01, and HLA- DRB1*12:02 [57, 58].
Gender
It is demonstrated that men have a higher mortality rate and worse outcomes during the COVID-19 pandemic and SARS epidemic [57]. It has been shown that vaccines are more efficient in women as they lead to higher antibody responses, though vaccine side effects are more exacerbated within this gender [58, 59]. Several factors, including the immune system, physiological factors, sex hormones, lifestyle, socio-cultural behaviors, and prevalence of underlying diseases, are involved in morbidity and mortality [57, 60].
SARS-CoV
Meaningful differences were observed in animal models regarding the level of pro-inflammatory cytokines, IL-6, and specific chemokines. Pro-inflammatory cytokines, IL-6 and C–C motif chemokine ligand 2 (CCL2) and C-X-C motif chemokine ligand 1 (CXCL1) expression were reported to increase in the lungs of male mice. As a result, inflammatory responses in male mice lasted longer, causing higher mortality and lung immunopathology. Moreover, ACE2 expression was lower in females with heart failure than in men [59]. In a group of SARS-CoV cases in Beijing, the infection rate was similar in males and females; however, the studies revealed gender-related differences regarding case fatality rates (CFR), with males experiencing higher CFRs than females [61]. A study on a mouse model of SARS-CoV infection displayed greater susceptibility to the disease in male mice compared to females of the same age [62].
SARS-CoV-2 interaction with different cell types
Estrogens may downregulate the expression of ACE2 receptor genes located on X chromosomes and are associated with interferon expression in previous research [57, 59, 63]. Although some findings reported a relation between ACE2 receptors and mortality rate in men, the exact association is still a mystery. Besides, several studies noted that males have more ACE2 receptors on the endothelium of the lungs [59, 64]. The expression of TMPRSS2 mRNA is not different among males and females in the lung tissue, but it may be regulated by androgens of prostate cells [57, 63]. Moreover, as men express more TMPRSS2 protein in their androgen-sensitive tissues like the prostate and testis, they are more sensitive to the infection [55].
A single X chromosome in men compared to two in women increases inflammation within this gender [64]. Aging significantly reduces immune factors like B cells, T cells, and natural killer cells in males [59]. Sex hormones also regulate the function of the immune system [57, 59, 63]. Higher levels of IL-6 can cause severe disease among males [57]. These differences may eventually lead to a faster clearance of pathogens in females [46].
About early antiviral responses, Toll-like receptor 7 (TLR7) plays a role in innate sensing of the virus and has higher expression in females [57, 59, 64]. IFN-α is also expressed more in adult females than adult males [47]. Altogether, females have more efficient innate immune responses and a higher level of inflammatory cytokine production than males [58, 65]. Viral detection takes longer in males [63]. Macrophage and neutrophil activity and pattern recognition receptors are also said to differ among these two genders but have not been completely clarified [58].
In adaptive immune responses, antibody levels are higher in females since estrogens lead to escalation of the somatic hypermutations, gender dissimilarities in germinal center formation, fewer stringent selection against autoreactive B cells, and the epigenetic obtainability of B cell loci [63, 64]. T cells are also affected by gender. Overall, T-cell responses are more robust in females than males [58]. It has been shown that ACE2 genes are expressed in the testicles at high levels, and that is why viral clearance takes longer for males compared to female counterparts [59].
Age
During the time of the COVID-19 pandemic, the elderly population became more susceptible to the disease, and they were at increased risk of death; therefore, it is crucial to compare mortality and morbidity variations between individuals of different ages, since these factors can lead to diverse responses to viruses as well as vaccines [66,67,68]. The relation between ACE2 expression and aging is still in investigation. ACE2 expression was lower in the heart, but its activity was raised in aged animals [69]. In the lungs, ACE2 mRNA expression was higher in adult females, whereas protein levels and activity were reduced in the aged females compared to males. In the kidneys, ACE2 activity is sex-dependent rather than age-dependent, with an elevation in males [70]. Estrogens and androgens level is also reduced as people get older, which may impact ACE2 expression.
Moreover, higher levels of ACE2 in children probably lead to a protective effect against SARS-CoV-2 [71]. This is most likely due to children’s unique ACE plasma profile that can be identified from birth. A rise in urine and plasma levels of ACE2, as well as an increase in local placental production and activity of ACE2, was recognized during mid to late pregnancy as a result of estradiol, known as the modulator of the ACE/ACE2. ACE can cross the placenta, allowing the mother to pass on her immunity and other protective soluble elements to the infant [72,73,74]. The number of lymphocytes significantly decreases at the early stages of the disease in adults, while children have normal white blood cells and lymphocytes count [67]. It is also mentioned that “trained immunity” in children can lead to a milder disease [71]. Breastmilk containing some antiviral proteins such as Lactoferrin and Casein, along with maternal antibodies, are other factors protecting infants against infection [66].
In 2021, after global vaccination, several studies were done to evaluate age as one of the factors for vaccine immune responses. Xia et al. claimed that people aged 60 and above who received the Sinopharm inactivated vaccine showed fewer neutralizing antibodies than those aged 18–59. In another study, Müller et al. concluded that 31.3% of people aged 80 and older given the BNT162b2 vaccine display no neutralizing antibodies after the second injection, compared to 2.2% of those younger than 60 [39]. Therefore the diversity in the responses among age categories was enumerated as a challenge for vaccine production.
Ethnicity and geographical differences
African-American race is considered to have a greater mortality rate [75]. A study showed that ACE2 expression was high in East Asian females [76]. TMPRSS2 allele frequency is also different among populations as it is a lower frequency in East Asians [55]. Thus, differences in gene expression, immune system, or even genetic background play a role in diverse responses to SARS-CoV-2 [76]. Furthermore, some national variations such as their access to health care, quality of nutrition, substance abuse, social distancing policies, social behavior, the incidence of obesity, diabetes, being over 65 years old, access to health care, and socioeconomic development level should be considered to explain why mortality rates vary from place to place [75, 77, 78].
Infection history
At the beginning of the global vaccination, the debatable issue was whether previously infected patients with COVID-19 were required to receive the same full range of vaccination regimens as those who were not experienced. Hence, the rate of specific anti-SARS-CoV-2 neutralizing antibodies in the serum samples was examined. Among vaccine regimens using the BNT162b2nAb, titers were considerably lower in uninfected people receiving both doses than in previously infected patients receiving only one dose. Thereby, to maximize the herd [79] of the entire regimens, it is recommended to vaccinate previously infected patients with only a single dose [39, 80,81,82].
SARS-CoV and SARS- CoV-2 vaccines
COVID-19 vaccine producers have taken the previous investigations on SARS vaccines (in 2003) into account. During the SARS outbreak, it took almost four months to develop a range of usable antigens for the animal, and cell culture trials before the genome sequence of the coronavirus became available. The first human trial on the SARS vaccine was conducted in Beijing (December 2004) after the epidemic had ended. Therefore, studies stopped at the preclinical stage, and still, there are no available vaccines against SARS-CoV and MERS-CoV [83, 84] (Table 3). The lack of SARS-CoV and MERS-CoV vaccines could be related to insufficient funding and a poor understanding of the viruses’ biology [84].
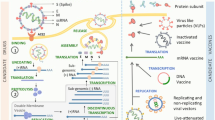
The mechanism of action of SARS-CoV and SARS-CoV-2 vaccines significantly resemble each other. Scientists attempt to find ways to expose the immune system to the spike protein on the virus membrane to stimulate virus identification during further exposure and induce antibody responses (Tables 1, 2 and 3). To this end, the same vaccine platforms were used to develop vaccines against SARS-CoV and SAR-CoV-2, which are discussed as follows. SARS-CoV, SAR-CoV-2, and MERS vaccine evolution is briefly summarized in Fig. 2.
Vaccine platforms
Virus-based vaccines
Virus-based vaccines are the platform most commonly used for other diseases, either as an inactivated (killed) or live attenuated virus that contains most of the virus antigens [14]. In killed vaccines, the virus is inactivated by radiation, heat, or formaldehyde. This platform offers a broad antigenic profile while being safe (i.e., the incapability to replicate) and less expensive than DNA/RNA vaccines. In live attenuated vaccines, serial passage of the virus leads to a reduction in its virulence. Hence, this platform produces strong and long-lasting humoral and cellular immune responses and offers a broad antigenic profile. Meanwhile, virus-based vaccines are associated with safety concerns due to their higher risk of virus replication, and thus, they are rarely used in immunocompromised patients [140,141,142].
Protein-based vaccines
Instead of using the whole virus, many researchers choose protein subunit vaccines and use antigens with strong immunogenicity, for example, the S protein or only the RBD [14]. The recombinant RBD vaccines compose of several conformational neutralizing epitopes, stimulating effective neutralizing antibodies against SARS-CoV [143]. Another protein-based vaccine strategy includes virus-like particles (VLP), offering a broad antigenic profile via imitating the SARS-CoV-2 structure on the surface of a non-replicative empty virus shell without genetic material [14, 140]. This platform is considered safer with lower cost than conventional vaccine platforms [141, 142].
Viral vector-based vaccines
Several viruses, e.g., vesicular stomatitis virus (VSV), influenza, measles, and adenovirus, could also be engineered as replicative or non-replicative recombinant vectors which express coronavirus S protein [14]. Vaccines produced by this platform are categorized as replicating and non-replicating vectors. In non-replicating kinds, virus replication is prevented by removing a gene responsible for encoding a structural protein. As the name suggests, replicating vectors can replicate in the cells and, therefore, require a smaller dosage. However, there are some safety concerns regarding their administration in immunocompromised patients [141, 142].
Nucleic acid-based vaccines
DNA-based vaccines
DNA vaccines are usually created using plasmid DNA containing eukaryotic expression elements to encode one or more antigens. DNA vaccines are stable, generally administered through intramuscular and intradermal injection, and activate both humoral and cellular immune responses. One obstacle is that they shall cross two cellular membranes before entering the nucleus. Another is that no DNA vaccine has previously been approved for human administration [14, 142].
RNA-based vaccines
The mRNA, encapsulated in lipid nanoparticles, enters the cytoplasm as a template to be translated, making multiple copies of the antigen. It may code the full-length S protein or a fraction [14, 141]. Like vector and DNA-based vaccines, mRNA vaccines can induce both humoral and cellular immunity. mRNA vaccines are considered to be safe, fast, and efficient. The only setback is the low temperature needed for transportation and storage [141, 142].
DNA/ RNA as vaccine delivery system for COVID-19
The key challenge in developing DNA/RNA vaccines is the host cells' probability of being picked up. There are two standard methods of introducing the DNA/RNA into the host immune cells [14]: first, using viral vectors; and second, using a delivery system to carry the DNA/RNA across the cell membrane to boost the synthesis of S protein.
Electroporation
This method temporarily applies a high-voltage electric pulse to the living cells. Consequently, DNA can pass through the membrane with more permeability. INOVIO pharmaceutical uses this technology (CELLECTRA®) to produce the COVID-19 vaccine [144].
Oligonucleotides (DNA/RNA) delivery
Oligonucleotides are macromolecules that present high therapeutic indexes remarkably when the formulation is tailored to reach specific tissues and sites of action. In the process of oligonucleotide delivery, nanoparticles need to encapsulate adequate amounts of nucleic acid and have specific tissue targeting properties. Thus, combining LNPs (Lipid-Based Nanoparticles) and immune-modulatory oligonucleotide adjuvants induces synergistic effects for immunologic responses [145, 146].
Adeno-associated virus (AAV) and lentivirus-based vaccines
Seven front runners (in clinical stages) of COVID-19 vaccine candidates mainly use nucleic acid vaccines which contain non-replicating viral vectors such as adeno-associated virus or lentivirus for vaccine delivery [146]. AAV encodes different antigens, stimulating a robust immune response in vaccinated individuals via various delivery methods. With these promising qualities, rAAV vectors are widely used for vaccine development (e.g., AAV-2 serotype as a vector for delivery of SARS-CoV immunogen) [147,148,149].
Lipid nanoparticle (LNP) systems
Viruses and nanoparticles are similar in size. Nano-materials are also ideal for antigen delivery as adjuvants and copies of viral structures, allowing nanotechnology to assist vaccine development [150]. Nanotechnology aids novel vaccine design, especially in the case of COVID-19. The first vaccine candidate launched into clinical trials is an mRNA vaccine delivered via lipid nanoparticles [146]. The announcement of the Pfizer and Moderna vaccines breakthrough in November 2020 has gained the attention of scientists and manufacturers. Practically, a vital part of these vaccines' success is based on their drug delivery particle, the lipid nanoparticle carrier. This system solves the delivery challenges by transporting the vaccine to the proper cellular populations and subcellular locations [92].
Moreover, LNPs can fulfill the basic conditions of an RNA/DNA delivery system. They can also preserve nucleic acids from digestion when they move to the target cell. Last, LNPs can be produced by catatonic outer membranes to allow cell entry [151,152,153].
LNPs improve the stability of mRNA-based vaccines such as mRNA-1273 Moderna. Nanocarriers present these payloads (DNA-RNA) to antigen-presenting cells (APCs). These carriers can provide innate adjuvant behavior and synchronize delivery of both antigen and adjuvant to target immune cells [154, 155].
Silica nanoparticle for DNA and RNA delivery
This method has been proven to be safe. It is an alternative to the LNP method to cover LNP deficiencies (i.e., insufficient delivery of nucleic acids into the cells). Another crucial advantage is that the mesoporous silica-nanoparticles (MSNs) have excellent biocompatibility and chemical stability attaching to oligonucleotides, including DNA, RNA, and siRNA [146, 150].
COVID-19 vaccine challenges
Probably, the best way to limit infections is vaccination. It has two outstanding usages; first, decreasing the number of infected people and hence lessening virus spreading; second, preventing multi-infection, and if not, reducing post-recovery syndromes [15]. The manufacturers have been facing some challenges during the development of the COVID-19 vaccine, as outlined below.
Animal models
The virus's ability to infect a particular cell or tissue (viral tropism) may lead to the suspension of the vaccine development at the preclinical phase and is the rationale behind animal testing [156]. This big challenge must be addressed before the clinical phases [15]. Another challenge is that the vaccine candidate may trigger an immune response in animal models, which may not be replicated in humans. This is because animals may not produce biological characteristics akin to the human model, which makes the mortality and efficacy rates unreliable [157]. Nevertheless, pigs make good models since they are susceptible to SARS-CoV-2 and have a human-like nature [158]. In vitro experiments on animal models do not necessarily assure a similar efficacy for humans.
Viral vector-based vaccines
The viral vector’s challenge lies in the production of the vaccine. This incomplete process results in the defective vector turning into some plasmids. Viral vector-based vaccine production has an impact on the cost and recovery yield. Enhancing the downstream process reduces the price and increases the recovery yield [150].
S protein
Vaccines block the interaction between ACE2 receptors on the host cells and S proteins [159]. While not much biological data exists on the whole mechanism [158]. If there is enough concentration of S protein in the body, its bioavailability causes the infection to spread [159]. Moreover, the ratio of the titer of antibody IgG to nucleotides and virion depends on the S protein. The formation of S protein complexes decreases the IgG efficacy [159]. Vaccine developers focus more on S protein as a functional site of the virus due to its ability to attach to the receptor. Vaccines can block this and make virus-neutralizing antibodies in the lung cells [158]. Mutation of S protein, although it may lead to stronger binding between S protein and receptor by affecting the function, might not affect viral pathogenicity [159].
Humoral immune responses
Recombinant protein vaccines cause humoral immune responses, which cannot provide decent immunogenicity for the body [150]. These proteins may have higher efficacy if adjuvants are added to their formulations; for instance, Novavax, adjuvanted with matrix M1, uses the noted strategy [83]. Recently, Sanofi adjuvanted recombinant protein SARS-CoV-2 vaccine candidate in collaboration with the US Biomedical Advanced Research and Development Authority (BARDA) and GSK is under major examination focusing on original D.614 virus as well as B.1.351 variant among 18 and older, in diverse regions [160]. This platform is suitable for middle- and low-income countries since it is inexpensive and safer than other platforms [161].
Glycosylation
Glycosylation of the vaccine upon administration in humans is a big challenge. Through glycosylation, the vaccine camouflages itself so the immunogenic antigens cannot be visible to the immune system leading to evasion from the host immune system. This phenomenon may negatively impact the effectiveness of the vaccine [159].
mRNA vaccines
mRNA is a platform that has been much used in designing the new vaccines for SARS-CoV-2. It limits the spreading of infections; however, drawbacks, including safety and immunogenicity, are among the challenges hindering vaccine development. Moreover, RNA is destroyed quickly, another con of this platform [159]. The risk of infection still exists in vaccine platforms that use weakened or killed viruses. However, they are more reliable and safer than others. These vaccines have some non-specific effects like activating memory cells, i.e., antibodies produced by mRNA vaccines target S protein (skipping the glycosylation), unlike vaccines containing killed or weakened viruses [159]. Consequently, vaccines that only contain specific antigens, such as Pfizer/BioNTech (BNT162b2) or Moderna/NIAID (mRNA-1273), are better able to launch an adaptive immune attack against the virus. By introducing these latter antigens, the immune system is distracted without being able to target the virus effectively with induced antibodies or T cells [162].
Mutations' effect on vaccine efficacy
A mutation is an expected change in genome sequencing that makes organisms different. So far, many mutations have been reported, but most do not affect transmission and pathogenicity. Usually, due to the ability of proofreading during replication, the rate of mutations is less than other RNA viruses. However, it has been suggested that a G614 mutation in the virus spike glycoprotein (compared to the previous variant of D614) could increase the transmission and infectivity of the virus but did not affect disease severity [163]. Moreover, studies have shown that certain medicines could affect the rate of coronavirus mutation. Coronavirus mutation rates increased when β-D-N4-hydroxycytidine-5'-isopropyl ester (NHC) exerted antiviral activity. This compound inhibited the replication of highly pathogenic human coronaviruses. EIDD-2801, an antiviral drug known as molnupiravir, is more effective at combating MERS-CoV infection when mutation rates are higher [164].
These are variants with various mutations in their genome sequences which may cause further challenges in vaccine efficacy:
-
Variant B.1.1.7 (also known as 20I / 501Y.V1 and VOC 202,012/01): this variant was discovered for the first time in the United Kingdom. Significant mutations in the deletion processes are found in strains N5014, P681H, H69-V70, and Y144/145 of B.1.1.7. These mutations of N501Y increased the affinity to the receptor binding, explaining the rapid spread of B.1.1.7 [165]. One of the mutations in the B.1.1.7 variant is a mutation in the N501Y receptor binding domain that has increased the virus transmission rate [158]. A study of variant B.1.1.7 of SARS-CoV-2 showed that its mortality rate was 30 percent higher than other previous variants [165].
-
Variant B.1.351 (also known as 20H / 501Y.V2): this variant was found in South Africa. The N501Y and E484K mutations have also been found within this variant, the latter is said to affect neutralization by polyclonal and monoclonal antibodies. There has been no evidence that this variant is associated with the disease severity [158]. Numerous mutations have been identified in NTD, three in RBD, and one at the furin cleavage site. Due to the emergence of new variants, the efficacy of current monoclonal antibodies (mAb) therapies and vaccines is at risk. In conclusion, many mutations occur, either in the antigenic supersite (NTD16,17) or in the ACE2-binding site (which is a major target of antiviral antibodies) [165, 166].
-
Variant B.1.1.28 (with the new name P.1 that was discovered in the four Brazilian passengers in Tokyo, Japan): The P.1 variant also has 17 mutations and three deletions, as well as the E484K and N501Y mutations. The possibility exists that certain mutations within this variant could impair the ability of antibodies produced by infection or vaccination to neutralize or detect the virus [158]. Viruses with co-mutations with the P.1 variant carry an increased risk of spreading the disease. It is a fact that the common mutation in the variant permitted contamination similar to that in the South African variant, as well as creating new risk factors [165, 166].
-
Variant B.1.617.2 (also known as Delta) was first detected in Maharashtra in October 2020. It has spread nationwide since then. Compared to the wild-type Wuhan-1 bearing D614G, B.1.617.2 is sixfold less sensitive to neutralizing serum antibodies from recovered individuals and eight-fold more sensitive to antibodies induced by vaccination. A lower level of neutralizing antibody against B.1.617.2 was found in ChAdOx1 vaccinates than in BNT162b2 [167].
-
South Africa first reported the variant B.1.1.529 (named Omicron) to WHO on November 24, 2021. A recent epidemiological report on South Africa showed three distinct peaks in reported cases. In the spike, Omicron has 26–32 mutations, which make it highly divergent. There is a possibility that some of these mutations involve immune escape potential and higher transmission; however, existing data are conflicting [166].
Problems with vaccination strategies
Cold chain
Currently, companies are dealing with conserving the vaccines in the cold chain since biological compounds have better stability in liquid form. They invest 80% of their production expenses [150]. Converting liquid from vaccine to dry powder (currently available for intranasal vaccines) makes transportation more accessible, reduces vaccine cost, and makes it more heat-resistant [150]. Lyophilized spike protein was fully functional when incubated at 60 °C during the experiment. On the other hand, the integrity of the liquid formulations was compromised on day 7 at 40 °C. However, the liquid formulation of Spike protein failed after less than one day at 60 °C [168].
Antibody-dependent enhancement (ADE)
ADE is a dangerous condition that may aggravate the inflammation in vaccinated individuals upon exposure to the SARS-CoV-19. The mechanism behind this phenomenon is yet to be discovered. However, one possibility is that the non-neutralizing anti‐S antibodies may expedite uptake by macrophages. This leads to macrophage stimulation, production of pro-inflammatory cytokines (IL‐6, IL‐8, and MCP1), and loss of tissue‐repaired cytokine (TGFβ), causing the virus to enter the host cell and spread the infection [169]. A neutralizing antibody acts as a viral receptor. It binds to the surface spike proteins of coronaviruses, causing a conformational alteration in the spike and intermediating viral access to IgG Fc receptor-expressing cells. This immunopathological event has been reported in other viral infections. Previously, it has been shown that if the target cells were reinfected by another serotype of dengue virus (i.e., secondary infection), the pre-existing antibodies could not thoroughly neutralize the virus and led to ADE. Thus, human trials should carefully evaluate vaccine safety and potentially harmful immune responses [159]. One action plan to reduce the chance of ADE is using the S1 or RBD antigen of SARS-CoV-2 instead of the complete full‐length S protein. Moreover, selecting Th1‐skewed adjuvants rather than alum adjuvants may avoid the inflammatory, immune-pathological, and ADE effects [159].
Side effects
During the process of SARS and MERS vaccine manufacturing, trials failed due to significant and potentially lethal side effects. For instance, a whole-virus (inactivated) vaccine was tested in ferrets and non-human primates, and a virus-like particle vaccine was tested in mice. The vaccines provided protection; however, the lungs of mice were infected with the virus [158]. One of the common immunopathological complications related to SARS-CoV and MERS-CoV vaccines was ADE [170]. In addition, elevated temperature (range 1–2.5 °C peaked on the second day), nasal discharge, and sneezing were observed in ferrets during the SARS-CoV whole killed and adenovirus vector-based vaccine study [171]. According to the above-mentioned adverse effects, development processes end at the preclinical phase.
Nonetheless, everything has changed when it comes to the current COVID-19 pandemic. One year has passed since the first wave of the pandemic, and still, we are facing this virus’ mutations. Despite all the expected adverse effects, vaccine manufacturers should try their best not to eradicate but at least alleviate vaccines' side effects necessarily. Most SARS-CoV-2 vaccines use only small portions of the virus or the virus’ RNA as a platform. This may sidestep the problems with SARS-CoV vaccines containing more virus parts. Therefore, many different methods for vaccines are being tested by global laboratories to reduce these side effects [172].
Despite estimated side effects, approved vaccines worth the risk of not contracting COVID-19. The first dose and second dose of vaccination have unpleasant but not serious side effects, except in pregnant women and children [172]. These adverse effects are similar in different COVID-19 vaccines that reached the third clinical trial phase. These effects are also classified based on their duration and intensity [172].
According to the studies on SARS-CoV and MERS-CoV vaccine development process, there is a concern about the use of coronavirus S-based vaccines since inflammatory and pathological immune effects such as pulmonary eosinophilic infiltration and ADE may occur. Earlier studies on SARS and MERS vaccine candidates have pointed to the risk of ADE; however, there is no clear evidence for SARS-CoV-2 [95, 173].
Since side effects of COVID-19 vaccines are mild and transient, they are usually out of concern. There may be pain, redness, or swelling at the injection site. Tiredness, headache, muscle pain, chills, fever, and nausea are common side effects in the rest of the body [174]. These symptoms may be signs of immune system response in the activation of T-cells and B-cells. However, our attention was drawn to an issue, society’s uncertainty about being vaccinated. Therefore, we cannot assure vaccines are safe unless most of society has taken a shot [24, 175].
Reinfections
When a virus variant in circulation causes a second infection, which was not known to be present at the first infection, it is probably considered reinfection. Some SARS-CoV-2 reinfections (primarily mild) have been documented in several studies [176]. Kojima and Klausner reviewed several clinical and epidemiological studies. They found that the risk of reinfection with SARS-CoV-2 decreased by 80.5–100 percent among those who had had COVID-19 [176]. Vaccination efficacy against SARS-CoV-2 may vary in cases with a history of previous COVID-19 infection. Some studies compared the risk of reinfection between previously infected individuals who had never been vaccinated and those who had received vaccination after infection. A case–control study from May–June 2021 in Kentucky found that previously infected individuals who were unvaccinated had 2.3 times greater odds of reinfection than previously infected but vaccinated individuals. Delta was not the dominant variant in the United States at the time of the study [177]. A series of matched-cohort studies involving 1,531,736 individuals vaccinated with mRNA vaccines between December 21, 2020, and September 19 found that prior SARS-CoV-2 infection was associated with a significantly lower risk for breakthrough infection among vaccine receivers [178]. Such findings provide evidence for the claim that post-infection immunity plus post-vaccination immunity may result in antibody responses bigger than post-infection immunity (hybrid immunity). However, a severe SARS-CoV-2 has been reported after recovery from breakthrough infection by an alpha variant is fully vaccinated (COVISHIELD®) health workers [177]. Whole genome sequencing confirmed that reinfection had happened with the Delta variant.
After vaccination, the ratio of reinfection has decreased. Besides, Omicron may have an increased risk for reinfection than other variants of concern (e.g., those who have previously had COVID-19 are more likely to become infected with Omicron), but the information is limited [166]. The immunological black box of COVID-19 has not been fully decoded, and more cohort and retrospective studies are required to improve our understanding of vaccine efficacy upon emergence of new variants (Table 4).
Conclusion
During the SARS-CoV outbreak in 2002–2003, although 44 vaccines were launched in preclinical stages, only six vaccines got into clinical trials. In contrast to SARS-CoV-2, the prevalence of SARS-CoV was not that high, and the epidemic ended after a while. Therefore, the development of all SARS vaccines was left incomplete. Consequently, at the beginning of the COVID-19 outbreak, scientists used previous SARS-CoV data to extend COVID-19 vaccines. While published data on COVID-19 vaccines showed considerable efficacy and immunogenicity, future studies will provide more accurate information on probable vaccination impacts on receivers considering age, gender, and ethnicity, providing an opportunity to prevent people from transmitting the virus. COVID-19 vaccine developers are currently facing many challenges, including cold supply chain, vaccine stability duration, transportation difficulties, complications concerning booster doses, short time follow-up duration, not enough data about immunization and vaccines' long-term effects, the probability of asymptomatic infections after vaccination, interaction of vaccines with other medicines and more notably, limited number of receivers.
References
Cucinotta D, Vanelli M. WHO declares COVID-19 a pandemic. Acta Biomed. 2020;91(1):157–60.
Ouassou H, Kharchoufa L, Bouhrim M, et al. The pathogenesis of Coronavirus Disease 2019 (COVID-19): Evaluation and prevention. J Immunol Res. 2020;2020:1357983.
Xu J, Zhao S, Teng T, et al. Systematic comparison of two animal-to-human transmitted human Coronaviruses: SARS-CoV-2 and SARS-CoV. Viruses. 2020;12(2):244.
Alimetov, A. Worldmeters. Coronavirus update (live). 2022. https://www.worldometers.info/coronavirus/. Accessed 2 June 2022.
WHO. World Health Organization. Coronavirus disease (COVID-19). 2022. https://www.who.int/health-topics/coronavirus. Accessed 16 Mar 2021.
da Rosa Mesquita R, Francelino Silva Junior LC, Santos Santana FM, et al. Clinical manifestations of COVID-19 in the general population: systematic review.Wien Klin Wochenschr. 2021;133(7–8):377–382.
Cascella M, Rajnik M, Aleem A, Dulebohn SC, Di Napoli R. Features, evaluation, and treatment of Coronavirus (COVID-19). 2022 May 4. In: StatPearls [Internet]. Treasure Island: StatPearls Publishing; 2022.
Mousavi T, Abdolahi M. The economic status of OIC member states during and after the COVID-19 pandemic. 2021;29:10–14.
Hackett M. Average cost of hospital care for COVID-19 ranges from $51,000 to $78,000, Based on age. 2020. Healthcare Finance. Available from: https://www.healthcarefinancenews.com/news/average-cost-hospital-care-covid-19-ranges-51000-78000-based-age.
Petrosillo N, Viceconte G, Ergonul O, Ippolito G, Petersen E. COVID-19, SARS and MERS: are they closely related? Clin Microbiol Infect. 2020;26(6):729–34.
Aschwanden C. Five reasons why COVID herd immunity is probably impossible. Nature. 2021;591(7851):520–2.
FDA. US Food and Drug Administration. FDA takes additional action in fight against COVID-19 by issuing emergency use authorization for second COVID-19 vaccine. 2020. https://www.fda.gov/news-events/press-announcements/fda-takes-additional-action-fight-against-covid-19-issuing-emergency-use-authorization-second-covid. Accessed 16 Mar 2021.
Liu J, Xie W, Wang Y, Xiong Y, Chen S, Han J, Wu Q. A comparative overview of COVID-19, MERS and SARS: Review article. Int J Surg. 2020;81:1–8.
de Queiroz NMGP, Marinho FV, Chagas MA, et al. Vaccines for COVID-19: perspectives from nucleic acid vaccines to BCG as delivery vector system. Microbes Infect. 2020;22(10):515–24.
Shih HI, Wu CJ, Tu YF, Chi CY. Fighting COVID-19: A quick review of diagnoses, therapies, and vaccines. Biomed J. 2020;43(4):341–54.
Prompetchara E, Ketloy C, Palaga T. Immune responses in COVID-19 and potential vaccines: Lessons learned from SARS and MERS epidemic. Asian Pac J Allergy Immunol. 2020;38(1):1–9.
Kaur SP, Gupta V. COVID-19 Vaccine: A comprehensive status report. Virus Res. 2020;288:198114.
Fani M, Teimoori A, Ghafari S. Comparison of the COVID-2019 (SARS-CoV-2) pathogenesis with SARS-CoV and MERS-CoV infections. Future Virol. 2020. https://doi.org/10.2217/fvl-2020-0050.
Ahmed SF, Quadeer AA, McKay MR. Preliminary identification of potential vaccine targets for the COVID-19 Coronavirus (SARS-CoV-2) based on SARS-CoV immunological studies. Viruses. 2020;12(3):254.
Rossi GA, Sacco O, Mancino E, Cristiani L, Midulla F. Differences and similarities between SARS-CoV and SARS-CoV-2: spike receptor-binding domain recognition and host cell infection with support of cellular serine proteases. Infection. 2020;48(5):665–9.
Salamanna F, Maglio M, Landini MP, Fini M. Body localization of ACE-2: On the trail of the keyhole of SARS-CoV-2. Front Med (Lausanne). 2020;7:594495.
Geravandi S, Mahmoudi-Aznaveh A, Azizi Z, Maedler K, Ardestani A. SARS-CoV-2 and pancreas: a potential pathological interaction? Trends Endocrinol Metab. 2021;32(11):842–5.
Bates TA, Weinstein JB, Farley SE, Leier HC, Messer WB, Tafesse FG. Cross-reactivity of SARS-CoV structural protein antibodies against SARS-CoV-2. Preprint. bioRxiv. 2020;2020.07.30.229377.
Tay MZ, Poh CM, Rénia L, MacAry PA, Ng LFP. The trinity of COVID-19: immunity, inflammation and intervention. Nat Rev Immunol. 2020;20(6):363–74.
Shah VK, Firmal P, Alam A, Ganguly D, Chattopadhyay S. Overview of immune response during SARS-CoV-2 infection: lessons from the past. Front Immunol. 2020;11:1949.
Ferreira AC, Soares VC, de Azevedo-Quintanilha IG, et al. SARS-CoV-2 engages inflammasome and pyroptosis in human primary monocytes [published correction appears in Cell Death Discov. 2021 May 19;7(1):116]. Cell Death Discov. 2021;7(1):43.
Wang C, Xie J, Zhao L, et al. Alveolar macrophage dysfunction and cytokine storm in the pathogenesis of two severe COVID-19 patients. EBioMedicine. 2020;57:102833.
Moss P. The T cell immune response against SARS-CoV-2. Nat Immunol. 2022;23(2):186–93.
Diao B, Wang C, Tan Y, et al. Reduction and functional exhaustion of T cells in patients with Coronavirus Disease 2019 (COVID-19). Front Immunol. 2020;11:827.
Liu L, Xu L, Lin C. T cell response in patients with COVID-19. Blood Science. 2020;2(3):76–8.
Wu LP, Wang NC, Chang YH, et al. Duration of antibody responses after severe acute respiratory syndrome. Emerg Infect Dis. 2007;13(10):1562–4.
Peluso MJ, Takahashi S, Hakim J, et al. SARS-CoV-2 antibody magnitude and detectability are driven by disease severity, timing, and assay. Sci Adv. 2021;7(31):eabh3409.
Post N, Eddy D, Huntley C, et al. Antibody response to SARS-CoV-2 infection in humans: A systematic review. PLoS ONE. 2020;15(12):e0244126.
He Z, Ren L, Yang J, et al. Seroprevalence and humoral immune durability of anti-SARS-CoV-2 antibodies in Wuhan, China: a longitudinal, population-level, cross-sectional study. Lancet. 2021;397(10279):1075–84.
Dan JM, Mateus J, Kato Y, et al. Immunological memory to SARS-CoV-2 assessed for up to eight months after infection. Preprint. bioRxiv. 2020;2020.11.15.383323.
Wheatley AK, Juno JA, Wang JJ, Selva KJ, Reynaldi A, Tan H-X, Lee WS, Wragg KM, Kelly HG, Esterbauer R, Davis SK, Kent HE, Mordant FL, Schlub TE, Gordon DL, Khoury DS, Subbarao K, Cromer D, Gordon TP, Chung AW, Davenport MP, Kent SJ. Evolution of immune responses to SARS-CoV-2 in mild-moderate COVID-19. Nat Commun. 2021;12(1):1162.
Cohen KW, Linderman SL, Moodie Z, Czartoski J, Lai L, Mantus G, Norwood C, Nyhoff LE, Edara VV, Floyd K, de Rosa SC, Ahmed H, Whaley R, Patel SN, Prigmore B, Lemos MP, Davis CW, Furth S, O'Keefe J, Gharpure MP, Gunisetty S, Stephens KA, Antia R, Zarnitsyna VI, Stephens DS, Edupuganti S, Rouphael N, Anderson EJ, Mehta AK, Wrammert J, Suthar MS, Ahmed R, McElrath MJ. Longitudinal analysis shows durable and broad immune memory after SARS-CoV-2 infection with persisting antibody responses and memory B and T cells. medRxiv [Preprint]. 2021 Jun 18:2021.04.19.21255739. https://doi.org/10.1101/2021.04.19.21255739. Update in: Cell Rep Med. 2021;2(7):100354.
Alfego D, Sullivan A, Poirier B, Williams J, Adcock D, Letovsky S. A population-based analysis of the longevity of SARS-CoV-2 antibody seropositivity in the United States. eClinicalMedicine. 2021;36:100902.
Cheng ZJ, Xue M, Zheng P, et al. Factors affecting the antibody immunogenicity of vaccines against SARS-CoV-2: A focused review. Vaccines (Basel). 2021;9(8):869.
Anderson RM, Vegvari C, Truscott J, Collyer BS. Challenges in creating herd immunity to SARS-CoV-2 infection by mass vaccination. Lancet. 2020;396(10263):1614–6.
Brodin P. Immune determinants of COVID-19 disease presentation and severity. Nat Med. 2021;27(1):28–33.
Liu Q, Xu K, Wang X, Wang W. From SARS to COVID-19: What lessons have we learned? J Infect Public Health. 2020;13(11):1611–8.
Zhu Z, Lian X, Su X, Wu W, Marraro GA, Zeng Y. From SARS and MERS to COVID-19: a brief summary and comparison of severe acute respiratory infections caused by three highly pathogenic human coronaviruses. Respir Res. 2020;21(1):224.
Buchy P, Buisson Y, Cintra O, Dwyer DE, Nissen M, Ortiz de Lejarazu R, Petersen E. COVID-19 pandemic: Lessons learned from more than a century of pandemics and current vaccine development for pandemic control. Int J Infect Dis. 2021;112:300–317.
Ovsyannikova IG, Haralambieva IH, Crooke SN, Poland GA, Kennedy RB. The role of host genetics in the immune response to SARS-CoV-2 and COVID-19 susceptibility and severity. Immunol Rev. 2020;296(1):205–19.
Ghafouri-Fard S, Noroozi R, Vafaee R, et al. Effects of host genetic variations on response to, susceptibility and severity of respiratory infections. Biomed Pharmacother. 2020;128:110296.
Di Maria E, Latini A, Borgiani P, Novelli G. Genetic variants of the human host influencing the coronavirus-associated phenotypes (SARS, MERS and COVID-19): rapid systematic review and field synopsis. Hum Genomics. 2020;14(1):30.
Hou Y, Zhao J, Martin W, et al. New insights into genetic susceptibility of COVID-19: an ACE2 and TMPRSS2 polymorphism analysis. BMC Med. 2020;18(1):216.
Chen F, Zhang Y, Li X, Li W, Liu X, Xue X. The impact of ACE2 polymorphisms on COVID-19 disease: Susceptibility, severity, and therapy. Front Cell Infect Microbiol. 2021;11:753721.
Sehailia M, Chemat S. Antimalarial-agent artemisinin and derivatives portray more potent binding to Lys353 and Lys31-binding hotspots of SARS-CoV-2 spike protein than hydroxychloroquine: potential repurposing of artenimol for COVID-19. J Biomol Struct Dyn. 2021;39(16):6184–94.
Lanjanian H, Moazzam-Jazi M, Hedayati M, et al. SARS-CoV-2 infection susceptibility influenced by ACE2 genetic polymorphisms: insights from Tehran Cardio-Metabolic Genetic Study. Sci Rep. 2021;11(1):1529.
Karcioglu Batur L, Hekim N. Correlation between interleukin gene polymorphisms and current prevalence and mortality rates due to novel coronavirus disease 2019 (COVID-2019) in 23 countries. J Med Virol. 2021;93(10):5853–63.
Avendaño-Félix M, Ochoa-Ramírez LA, Ramos-Payán R, et al. Lack of effects of the genetic polymorphisms of interleukin-10 in clinical outcomes of COVID-19. Viral Immunol. 2021;34(8):567–72.
Saleh A, Sultan A, Elashry MA, et al. Association of TNF-α G-308 a promoter polymorphism with the course and outcome of COVID-19 patients. Immunol Invest. 2022;51(3):546–57.
Singh H, Choudhari R, Nema V, Khan AA. ACE2 and TMPRSS2 polymorphisms in various diseases with special reference to its impact on COVID-19 disease. Microb Pathog. 2021;150:104621.
Flannery DD, Gouma S, Dhudasia MB, Mukhopadhyay S, Pfeifer MR, Woodford EC, Gerber JS, Arevalo CP, Bolton MJ, Weirick ME, Goodwin EC, Anderson EM, Greenplate AR, Kim J, Han N, Pattekar A, Dougherty J, Kuthuru O, Mathew D, Baxter AE, Vella LA, Weaver J, Verma A, Leite R, Morris JS, Rader DJ, Elovitz MA, Wherry EJ, Puopolo KM, Hensley SE. SARS-CoV-2 seroprevalence among parturient women in Philadelphia. Sci Immunol. 2020;5(49):eabd5709.
Maleki Dana P, Sadoughi F, Hallajzadeh J, et al. An insight into the sex differences in COVID-19 patients: What are the possible causes? Prehosp Disaster Med. 2020;35(4):438–41.
Klein SL, Marriott I, Fish EN. Sex-based differences in immune function and responses to vaccination. Trans R Soc Trop Med Hyg. 2015;109(1):9–15.
Pradhan A, Olsson PE. Sex differences in severity and mortality from COVID-19: are males more vulnerable? Biol Sex Differ. 2020;11(1):53.
Moroianu L.A, Moroianu M, Toma A, Barbu R, Ardeleanu V, Nitoi, L.C. (2021). Psychopathology in patients diagnosed with Sars Cov 2: a brief report. Mediterr J Clin Psychol. 9(1). https://doi.org/10.6092/2282-1619/mjcp-2982.
Xie X, Chen J, Wang X, Zhang F, Liu Y. Age- and gender-related difference of ACE2 expression in rat lung. Life Sci. 2006;78(19):2166–71.
Channappanavar R, Fett C, Mack M, Ten Eyck PP, Meyerholz DK, Perlman S. Sex-based differences in susceptibility to Severe Acute Respiratory Syndrome Coronavirus Infection. J Immunol. 2017;198(10):4046–53.
Scully EP, Haverfield J, Ursin RL, Tannenbaum C, Klein SL. Considering how biological sex impacts immune responses and COVID-19 outcomes. Nat Rev Immunol. 2020;20(7):442–7.
Conti P, Younes A. Coronavirus COV-19/SARS-CoV-2 affects women less than men: clinical response to viral infection. J Biol Regul Homeost Agents. 2020;34(2):339–43.
Falahi S, Kenarkoohi A. Sex and gender differences in the outcome of patients with COVID-19. J Med Virol. 2021;93(1):151–2.
Root-Bernstein R. Age and location in severity of COVID-19 pathology: Do lactoferrin and pneumococcal vaccination explain low infant mortality and regional differences? BioEssays. 2020;42(11):e2000076.
Cao Q, Chen YC, Chen CL, Chiu CH. SARS-CoV-2 infection in children: Transmission dynamics and clinical characteristics. J Formos Med Assoc. 2020;119(3):670–3.
Furman D, Hejblum BP, Simon N, Jojic V, Dekker CL, Thiébaut R, Tibshirani RJ, Davis MM. Systems analysis of sex differences reveals an immunosuppressive role for testosterone in the response to influenza vaccination. Proc Natl Acad Sci USA. 2014;111(2):869–74.
Kai H, Kai M. Interactions of coronaviruses with ACE2, angiotensin II, and RAS inhibitors—lessons from available evidence and insights into COVID-19. Hypertens Res. 2020;43(7):648–54.
Viveiros A, Gheblawi M, Aujla PK, Sosnowski DK, Seubert JM, Kassiri Z, Oudit GY. Sex- and age-specific regulation of ACE2: Insights into severe COVID-19 susceptibility. J Mol Cell Cardiol. 2022;164:13–6.
Cristiani L, Mancino E, Matera L, et al. Will children reveal their secret? The coronavirus dilemma. Eur Respir J. 2020;55(4):2000749.73.
Ciaglia E, Vecchione C, Puca AA. COVID-19 infection and circulating ACE2 levels: Protective role in women and children. Front Pediatr. 2020;8:206.
Saheb Sharif-Askari N, Saheb Sharif-Askari F, Alabed M, et al. Airways expression of SARS-CoV-2 receptor, ACE2, and TMPRSS2 is lower in children than adults and increases with smoking and COPD. Mol Ther Methods Clin Dev. 2020;18:1–6.
Li G, He X, Zhang L, Ran Q, Wang J, Xiong A, Wu D, Chen F, Sun J, Chang C. Assessing ACE2 expression patterns in lung tissues in the pathogenesis of COVID-19. J Autoimmun. 2020;112:102463.
Navar AM, Purinton SN, Hou Q, Taylor RJ, Peterson ED. The impact of race and ethnicity on outcomes in 19,584 adults hospitalized with COVID-19. PLoS ONE. 2021;16(7):e0254809.
Chen J, Jiang Q, Xia X, et al. Individual variation of the SARS-CoV-2 receptor ACE2 gene expression and regulation. Aging Cell. 2020;19(7):e13168.
Baqui P, Bica I, Marra V, Ercole A, Van der Schaar M. Ethnic and regional variations in hospital mortality from COVID-19 in Brazil: a cross-sectional observational study. Lancet Glob Health. 2020;8(8):e1018–26.
Platt L, Warwick R. COVID-19 and ethnic inequalities in England and Wales [published online ahead of print, 2020 Jun 3].Fisc Stud. 2020; https://doi.org/10.1111/1475-5890.12228.
Roper RL, Rehm KE. SARS vaccines: where are we? Expert Rev Vaccines. 2009;8(7):887–98.
Saadat S, Rikhtegaran Tehrani Z, Logue J, et al. Binding and neutralization antibody titers after a single vaccine dose in health care workers previously infected with SARS-CoV-2. JAMA. 2021;325(14):1467–9.
Hall VJ, Foulkes S, Saei A, et al. COVID-19 vaccine coverage in health-care workers in England and effectiveness of BNT162b2 mRNA vaccine against infection (SIREN): a prospective, multicentre, cohort study. Lancet. 2021;397(10286):1725–35.
Levi R, Azzolini E, Pozzi C, et al. One dose of SARS-CoV-2 vaccine exponentially increases antibodies in individuals who have recovered from symptomatic COVID-19. J Clin Invest. 2021;131(12):e149154.
Dong Y, Dai T, Wang B, et al. The way of SARS-CoV-2 vaccine development: success and challenges. Signal Transduct Target Ther. 2021;6(1):387.
Boudjelal M, Nehdi A, Islam I. Why do SARS-COV vaccines not exist? The pharma scientific intelligence and business model must be revisited. Expert Opin Drug Discov. 2020;15(11):1233–5.
Mathew S, Faheem M, Hassain NA, et al. Platforms exploited for SARS-CoV-2 vaccine development. Vaccines (Basel). 2020;9(1):11.
WHO. World Health Organization. Draft landscape and tracker of COVID-19 candidate vaccines. 2020. https://www.who.int/docs/default-source/a-future-for-children/novel-coronavirus_landscape_covid-19.pdf?sfvrsn=4d8bd201_1. Accessed 16 Mar 2021.
Baviskar T, Raut D, Bhatt LK. Deciphering vaccines for COVID-19: where do we stand today? Immunopharmacol Immunotoxicol. 2021;43(1):8–21.
Chauhan N, Soni S, Gupta A, Aslam M, Jain U. Interpretative immune targets and contemporary position for vaccine development against SARS-CoV-2: A systematic review. J Med Virol. 2021;93(4):1967–82.
Silveira MM, Moreira GMSG, Mendonça M. DNA vaccines against COVID-19: Perspectives and challenges. Life Sci. 2021;267:118919.
Lee P, Kim CU, Seo SH, Kim DJ. Current status of COVID-19 vaccine development: Focusing on antigen design and clinical trials on later stages. Immune Netw. 2021;21(1):e4.
Rego GNA, Nucci MP, Alves AH, Oliveira FA, Marti LC, Nucci LP, Mamani JB, Gamarra LF. Current clinical trials protocols and the global effort for immunization against SARS-CoV-2. Vaccines (Basel). 2020;8(3):474.
Meo SA, Bukhari IA, Akram J, Meo AS, Klonoff DC. COVID-19 vaccines: comparison of biological, pharmacological characteristics and adverse effects of Pfizer/BioNTech and Moderna Vaccines. Eur Rev Med Pharmacol Sci. 2021;25(3):1663–9.
Team CC-R, Administration FaD. Allergic reactions including anaphylaxis after receipt of the first dose of moderna COVID-19 vaccine - United States, December 21, 2020-January 10, 2021. MMWR Morb Mortal Wkly Rep. 2021;70(4):125–129.
Tanne JH. Covid-19: Moderna plans booster doses to counter variants. BMJ. 2021;372:n232.
Tregoning JS, Brown ES, Cheeseman HM, Flight KE, Higham SL, Lemm NM, Pierce BF, Stirling DC, Wang Z, Pollock KM. Vaccines for COVID-19. Clin Exp Immunol. 2020;202(2):162–92.
Jones I, Roy P. Sputnik V COVID-19 vaccine candidate appears safe and effective. Lancet. 2021;397(10275):642–3.
Baraniuk C. Covid-19: What do we know about Sputnik V and other Russian vaccines? BMJ. 2021;372:n743.
Logunov DY, Dolzhikova IV, Shcheblyakov DV, et al. Safety and efficacy of an rAd26 and rAd5 vector-based heterologous prime-boost COVID-19 vaccine: an interim analysis of a randomised controlled phase 3 trial in Russia [published correction appears in Lancet. 2021;397(10275):670]. Lancet. 2021;397(10275):671–681.
kegame S, Siddiquey MNA, Hung CT, Haas G, Brambilla L, Oguntuyo KY, Kowdle S, Vilardo AE, Edelstein A, Perandones C, Kamil JP, Lee B. Neutralizing activity of Sputnik V vaccine sera against SARS-CoV-2 variants. Res Sq [Preprint]. 2021 Apr 8:rs.3.rs-400230. https://doi.org/10.21203/rs.3.rs-400230/v1. Update in: Nat Commun. 2021;12(1):4598.
Palacios R, Patiño EG, de Oliveira PR, et al. Double-blind, randomized, placebo-controlled phase III clinical trial to evaluate the efficacy and safety of treating healthcare professionals with the adsorbed COVID-19 (Inactivated) vaccine manufactured by Sinovac - PROFISCOV: A structured summary of a study protocol for a randomised controlled trial. Trials. 2020;21(1):853.
Kanno AI, Barbosa MMF, Moraes L, Leite LCC. SARS-CoV-2 vaccine development and how Brazil is contributing. Genet Mol Biol. 2021;44(1 Suppl 1):e20200320.
Zhang Y, Zeng G, Pan H, et al. Safety, tolerability, and immunogenicity of an inactivated SARS-CoV-2 vaccine in healthy adults aged 18–59 years: a randomised, double-blind, placebo-controlled, phase 1/2 clinical trial. Lancet Infect Dis. 2021;21(2):181–92.
Mahase E. Covid-19: Where are we on vaccines and variants? BMJ. 2021;372:n597.
Zhu FC, Guan XH, Li YH, et al. Immunogenicity and safety of a recombinant adenovirus type-5-vectored COVID-19 vaccine in healthy adults aged 18 years or older: a randomised, double-blind, placebo-controlled, phase 2 trial. Lancet. 2020;396(10249):479–88.
Ryzhikov A, et al. A single blind, placebo-controlled randomized study of the safety, reactogenicity and immunogenicity of the “EpiVacCorona” Vaccine for the prevention of COVID-19, in volunteers aged 18–60 years (phase I–II). Russ J Infect Immun. 11.2(2021):283–296. WEB. https://doi.org/10.15789/2220-7619-ASB-1699.
Polack FP, Thomas SJ, Kitchin N, et al. Safety and efficacy of the BNT162b2 mRNA Covid-19 vaccine. N Engl J Med. 2020;383(27):2603–15.
Walsh EE, Frenck RW Jr, Falsey AR, et al. Safety and immunogenicity of two RNA-based Covid-19 vaccine candidates. N Engl J Med. 2020;383(25):2439–50.
Muik A, Wallisch AK, Sänger B, et al. Neutralization of SARS-CoV-2 lineage B.1.1.7 pseudovirus by BNT162b2 vaccine-elicited human sera.Science. 2021;371(6534):1152–1153.
FDA. U.S. Food and Drug Administration. Pfizer-BioNTech COVID-19 Vaccine EUA fact sheet for healthcare providers administrating Vaccine, Food and Drug Administration (FDA). 2022. https://www.fda.gov/media/153715/download. Accessed 19 Jun 2022.
WHO. World Health Organization. mRNA vaccines against COVID-19: Pfizer-BioNTech COVID-19 vaccine BNT162b2. 2020. https://apps.who.int/iris/bitstream/handle/10665/338096/WHO-2019-nCoV-vaccines-SAGE_evaluation-BNT162b2-2020.1-eng.pdf?sequence=1&isAllowed=y. Accessed 16 Mar 2021.
Xia S, Zhang Y, Wang Y, et al. Safety and immunogenicity of an inactivated SARS-CoV-2 vaccine, BBIBP-CorV: a randomised, double-blind, placebo-controlled, phase 1/2 trial. Lancet Infect Dis. 2021;21(1):39–51.
Baraniuk C. What do we know about China's covid-19 vaccines? BMJ. 2021;373:n912.
Wang H, Zhang Y, Huang B, et al. Development of an inactivated vaccine candidate, BBIBP-CorV, with potent protection against SARS-CoV-2. Cell. 2020;182(3):713-721.e9.
Karim SSA. Vaccines and SARS-CoV-2 variants: the urgent need for a correlate of protection. Lancet. 2021;397(10281):1263–4.
Mallapaty S, Callaway E. What scientists do and don't know about the Oxford-AstraZeneca COVID vaccine. Nature. 2021;592(7852):15–7.
FDA. U.S. Food and Drug Administration. Janssen COVID-19 Vaccine (Johnson & Johnson). 2022. https://www.cdc.gov/vaccines/covid-19/info-by-product/janssen/index.html. Accessed 16 Mar 2021.
Yang S, Li Y, Dai L, et al. Safety and immunogenicity of a recombinant tandem-repeat dimeric RBD-based protein subunit vaccine (ZF2001) against COVID-19 in adults: two randomised, double-blind, placebo-controlled, phase 1 and 2 trials. Lancet Infect Dis. 2021;21(8):1107–19.
Bao ZJJ. Chinese Covid-19 Vaccine efficacy better than expected interview with Mr. Liu Jingzhen, Chairman of Sinopharm. Sinopharm. 2021. http://www.sinopharm.com/en/s/1395-4173-38923.html. Accessed 16 Mar 2021.
Le Thanh T, Andreadakis Z, Kumar A, et al. The COVID-19 vaccine development landscape. Nat Rev Drug Discov. 2020;19(5):305–6.
Abdoli A, Aalizadeh R, Aminianfar H, et al. Safety and potency of BIV1-CovIran inactivated vaccine candidate for SARS-CoV-2: A preclinical study. Rev Med Virol. 2022;32(3):e2305.
McPherson C, Chubet R, Holtz K, et al. Development of a SARS Coronavirus vaccine from recombinant spike protein plus delta inulin adjuvant. Methods Mol Biol. 2016;1403:269–84.
Fontanet A, Autran B, Lina B, Kieny MP, Karim SSA, Sridhar D. SARS-CoV-2 variants and ending the COVID-19 pandemic. Lancet. 2021;397(10278):952–4.
Forni G, Mantovani A, COVID-19 Commission of Accademia Nazionale dei Lincei Rm. COVID-19 vaccines: where we stand and challenges ahead. Cell Death Differ. 2021;28(2):626–639.
Fiolet T, Kherabi Y, MacDonald CJ, Ghosn J, Peiffer-Smadja N. Comparing COVID-19 vaccines for their characteristics, efficacy and effectiveness against SARS-CoV-2 and variants of concern: a narrative review. Clin Microbiol Infect. 2022;28(2):202–21.
Hernández‐Bernal F, Ricardo‐Cobas MC, Martín‐Bauta Y, et al. Safety, tolerability, and immunogenicity of a SARS‐CoV‐2 recombinant spike RBD protein vaccine: a randomised, double‐blind, placebo‐controlled, phase 1–2 clinical trial (ABDALA Study). eClinicalMedicine. 2022;46:101383.
Limonta-Fernández M, Chinea-Santiago G, Martín-Dunn AM, Gonzalez-Roche D, Bequet-Romero M, Marquez-Perera G, González-Moya I, Canaan-Haden-Ayala C, Cabrales-Rico A, Espinosa-Rodríguez LA, Ramos-Gómez Y, Andujar-Martínez I, González-López LJ, de la Iglesia MP, Zamora-Sanchez J, Cruz-Sui O, Lemos-Pérez G, Cabrera-Herrera G, Valdes-Hernández J, Martinez-Diaz E, Pimentel-Vazquez E, Ayala-Avila M, Guillén-Nieto G. The SARS-CoV-2 receptor-binding domain expressed in Pichia pastoris as a candidate vaccine antigen. preprint. medRxiv. 2021:2021.2006.2029.21259605.
Moodie EEM, Basta NE. Center for Genetic Engineering and Biotechnology (CIGB). CdIGyB. Phase III clinical study of Abdala. https://www.cigb.edu.cu/en/product/abdala-cigb-66-2/. Accessed 16 Mar 2021.
Pasteur Institut. Coronavirus: Institut pasteur warns against false information circulating on social media. https://www.pasteur.fr/en/research-journal/news/coronavirus-institut-pasteur-warns-against-false-information-circulating-social-media. Accessed 16 March 2021.
Pasteur Institut. MV-SARS-CoV-2 vaccine candidate: a new partnership between institut pasteur, cepi, thémis and msd. 2020. https://www.pasteur.fr/en/press-area/press-documents/mv-sars-cov-2-vaccine-candidate-new-partnership-between-institut-pasteur-cepi-themis-and-msd. Accessed 16 Mar 2021.
Liu YV, Massare MJ, Barnard DL, et al. Chimeric Severe Acute Respiratory Syndrome Coronavirus (SARS-CoV) S glycoprotein and influenza matrix 1 efficiently form virus-like particles (VLPs) that protect mice against challenge with SARS-CoV. Vaccine. 2011;29(38):6606–13.
Martin JE, Louder MK, Holman LA, et al. A SARS DNA vaccine induces neutralizing antibody and cellular immune responses in healthy adults in a Phase I clinical trial. Vaccine. 2008;26(50):6338–43.
Kusters IC, Matthews J, Saluzzo JF. Manufacturing vaccines for an emerging viral infection–specific issues associated with the development of a prototype SARS vaccine. Vaccines for Biodefense and Emerging and Neglected Diseases. 2009;147–156.
Lin JT, Zhang JS, Su N, et al. Safety and immunogenicity from a phase I trial of inactivated severe acute respiratory syndrome coronavirus vaccine. Antivir Ther. 2007;12(7):1107–13.
Alharbi NK, Qasim I, Almasoud A, et al. Humoral immunogenicity and efficacy of a single dose of ChAdOx1 MERS vaccine candidate in dromedary camels. Sci Rep. 2019;9(1):16292.
Modjarrad K, Roberts CC, Mills KT, et al. Safety and immunogenicity of an anti-Middle East respiratory syndrome coronavirus DNA vaccine: a phase 1, open-label, single-arm, dose-escalation trial. Lancet Infect Dis. 2019;19(9):1013–22.
Addo M. The development of a vaccine against MERS virus gets international support. German Center for Infection Research (DZIF). 2018. https://www.dzif.de/en/development-vaccine-against-mers-virus-gets-international-support. Accessed 02 Mar 2021.
Gupta T, Gupta SK. Potential adjuvants for the development of a SARS-CoV-2 vaccine based on experimental results from similar coronaviruses. Int Immunopharmacol. 2020;86:106717.
olegatti PM, Ewer KJ, Aley PK, et al. Safety and immunogenicity of the ChAdOx1 nCoV-19 vaccine against SARS-CoV-2: a preliminary report of a phase 1/2, single-blind, randomised controlled trial [published correction appears in Lancet. 2020 Aug 15;396(10249):466] [published correction appears in Lancet. 2020 Dec 12;396(10266):1884].Lancet. 2020;396(10249):467–478.
van Doremalen N, Haddock E, Feldmann F, et al. A single dose of ChAdOx1 MERS provides protective immunity in rhesus macaques. Science Advances. 2020 Jun;6(24):eaba8399.
Nidom RV, Ansori ANM, Indrasari S, Normalina I, Kusala MKJ, Saefuddin A, Nidom CA. Recent updates on COVID-19 Vaccine platforms and its immunological aspects: A review. SRP. 2020;11(10):807–18. https://doi.org/10.31838/srp.2020.10.121.
Li Y, Tenchov R, Smoot J, Liu C, Watkins S, Zhou Q. A comprehensive review of the global efforts on COVID-19 vaccine development. ACS Cent Sci. 2021;7(4):512–33.
Sharma O, Sultan AA, Ding H, Triggle CR. A review of the progress and challenges of developing a vaccine for COVID-19. Front Immunol. 2020;11:585354.
Jiang S, He Y, Liu S. SARS vaccine development. Emerg Infect Dis. 2005;11(7):1016–20.
Matone B. Inovio receives new $5 million grant to accelerate scale up of smart delivery device for its Covid-19 vaccine. https://www.biospace.com/article/releases/inovio-receives-new-5-million-grant-to-accelerate-scale-up-of-smart-delivery-device-for-its-covid-19-vaccine/. Accessed 10 Feb 2021.
Thi TTH, Suys EJA, Lee JS, Nguyen DH, Park KD, Truong NP. Lipid-based nanoparticles in the clinic and clinical trials: From cancer nanomedicine to COVID-19 vaccines. Vaccines (Basel). 2021;9(4):359.
Theobald N. Emerging vaccine delivery systems for COVID-19: Functionalised silica nanoparticles offer a potentially safe and effective alternative delivery system for DNA/RNA vaccines and may be useful in the hunt for a COVID-19 vaccine. Drug Discov Today. 2020;25(9):1556–8.
Du L, Zhao G, Lin Y, et al. Intranasal vaccination of recombinant adeno-associated virus encoding receptor-binding domain of severe acute respiratory syndrome coronavirus (SARS-CoV) spike protein induces strong mucosal immune responses and provides long-term protection against SARS-CoV infection. J Immunol. 2008;180(2):948–56.
Bouquet C, Vignal Clermont C, Galy A, et al. Immune response and intraocular inflammation in patients with leber hereditary optic neuropathy treated with intravitreal injection of recombinant adeno-associated virus 2 carrying the ND4 gene: A Secondary analysis of a phase 1/2 clinical trial. JAMA Ophthalmol. 2019;137(4):399–406.
Zabaleta N, Dai W, Bhatt U, Chichester JA, Sanmiguel J, Estelien R, Michalson KT, Diop C, Maciorowski D, Qi W, Hudspeth E, Cucalon A, Dyer CD, Pampena MB, Knox JJ, LaRocque RC, Charles RC, Li D, Kim M, Sheridan A, Storm N, Johnson RI, Feldman J, Hauser BM, Zinn E, Ryan A, Kobayashi DT, Chauhan R, McGlynn M, Ryan ET, Schmidt AG, Price B, Honko A, Griffiths A, Yaghmour S, Hodge R, Betts MR, Freeman MW, Wilson JM, Vandenberghe LH. Immunogenicity of an AAV-based, room-temperature stable, single dose COVID-19 vaccine in mice and non-human primates. bioRxiv [Preprint]. 2021 Jan 19:2021.01.05.422952. https://doi.org/10.1101/2021.01.05.422952. Update in: Cell Host Microbe. 2021;29(9):1437–1453.e8.
Wang J, Peng Y, Xu H, Cui Z, Williams RO. The COVID-19 vaccine race: Challenges and opportunities in vaccine formulation. AAPS PharmSciTech. 2020;21(6):225.
Schoenmaker L, Witzigmann D, Kulkarni JA, et al. mRNA-lipid nanoparticle COVID-19 vaccines: Structure and stability. Int J Pharm. 2021;601:120586.
Pilkington EH, Suys EJA, Trevaskis NL, et al. From influenza to COVID-19: Lipid nanoparticle mRNA vaccines at the frontiers of infectious diseases. Acta Biomater. 2021;131:16–40.
McKay PF, Hu K, Blakney AK, et al. Self-amplifying RNA SARS-CoV-2 lipid nanoparticle vaccine candidate induces high neutralizing antibody titers in mice. Nat Commun. 2020;11(1):3523.
Scioli Montoto S, Muraca G, Ruiz ME. Solid lipid nanoparticles for drug delivery: Pharmacological and biopharmaceutical aspects. Front Mol Biosci. 2020;7:587997.
Florindo HF, Kleiner R, Vaskovich-Koubi D, et al. Immune-mediated approaches against COVID-19. Nat Nanotechnol. 2020;15(8):630–45.
McFadden G, Mohamed MR, Rahman MM, Bartee E. Cytokine determinants of viral tropism. Nat Rev Immunol. 2009;9(9):645–55.
Zumla A, Chan JF, Azhar EI, Hui DS, Yuen KY. Coronaviruses - drug discovery and therapeutic options. Nat Rev Drug Discov. 2016;15(5):327–47.
Begum J, Mir NA, Dev K, Buyamayum B, Wani MY, Raza M. Challenges and prospects of COVID-19 vaccine development based on the progress made in SARS and MERS vaccine development. Transbound Emerg Dis. 2021;68:1111–24.
Wang F, Kream RM, Stefano GB. An evidence based perspective on mRNA-SARS-CoV-2 vaccine development. Med Sci Monit. 2020;26:e924700.
Trimophe, Th. Sanofi. Our adjuvanted recombinant protein-based COVID-19 vaccine candidate. 2022. https://www.sanofi.com/en/our-covid-19-vaccine-candidates/recombinant-vaccine. Accessed 19 Mar 2022.
Hotez PJ, Bottazzi ME. Developing a low-cost and accessible COVID-19 vaccine for global health. PLoS Negl Trop Dis. 2020;14(7):e0008548.
Fernández A. Glycosylation of SARS-CoV-2 steers evolutionary outcomes in the postvaccination phase. ACS Pharmacol Transl Sci. 2021;4(1):410–2.
Cevik M, Kuppalli K, Kindrachuk J, Peiris M. Virology, transmission, and pathogenesis of SARS-CoV-2. BMJ. 2020;371:m3862.
Sheahan TP, Sims AC, Zhou S, et al. An orally bioavailable broad-spectrum antiviral inhibits SARS-CoV-2 in human airway epithelial cell cultures and multiple coronaviruses in mice.Sci Transl Med. 2020;12(541):eabb5883.
Cosar B, Karagulleoglu ZY, Unal S, Ince AT, Uncuoglu DB, Tuncer G, Kilinc BR, Ozkan YE, Ozkoc HC, Demir IN, Eker A, Karagoz F, Simsek SY, Yasar B, Pala M, Demir A, Atak IN, Mendi AH, Bengi VU, Cengiz Seval G, Gunes Altuntas E, Kilic P, Demir-Dora D. SARS-CoV-2 Mutations and their Viral Variants. Cytokine Growth Factor Rev. 2022;63:10–22. https://doi.org/10.1016/j.cytogfr.2021.06.001.
WHO. World Health Organization. Enhancing readiness for omicron (B.1.1.529): Technical brief and priority actions for member states. 2021. https://www.who.int/publications/m/item/enhancing-readiness-for-omicron-(b.1.1.529)-technical-brief-and-priority-actions-for-member-states. Accessed 26 Dec 2021.
Mlcochova P, Kemp SA, Dhar MS, et al. SARS-CoV-2 B.1.617.2 Delta variant replication and immune evasion. Nature. 2021;599(7883):114–119.
Mabrouk MT, Chiem K, Rujas E, Huang WC, Jahagirdar D, Quinn B, Surendran Nair M, Nissly RH, Cavener VS, Boyle NR, Sornberger TA, Kuchipudi SV, Ortega J, Julien JP, Martinez-Sobrido L, Lovell J. Lyophilized, thermostable Spike or RBD immunogenic liposomes induce protective immunity against SARS-CoV-2 in mice. Sci Adv. 2021;7(49):eabj1476.
Padron-Regalado E. Vaccines for SARS-CoV-2: Lessons from Other Coronavirus Strains. Infect Dis Ther. 2020:1–20.
Li YD, Chi WY, Su JH, Ferrall L, Hung CF, Wu TC. Coronavirus vaccine development: from SARS and MERS to COVID-19. J Biomed Sci. 2020;27(1):104.
See RH, Petric M, Lawrence DJ, et al. Severe acute respiratory syndrome vaccine efficacy in ferrets: whole killed virus and adenovirus-vectored vaccines. J Gen Virol. 2008;89(Pt 9):2136–46.
Pang J, Wang MX, Ang IYH, Tan SHX, Lewis RF, Chen JI, Gutierrez RA, Gwee SXW, Chua PEY, Yang Q, Ng XY, Yap RK, Tan HY, Teo YY, Tan CC, Cook AR, Yap JC, Hsu LY. Potential rapid diagnostics, vaccine and therapeutics for 2019 Novel Coronavirus (2019-nCoV): A systematic review. J Clin Med. 2020;9(3).
Wan Y, Shang J, Sun S, et al. Molecular mechanism for antibody-dependent enhancement of Coronavirus entry. J Virol. 2020;94(5):e02015-e2019.
CDC. Centers for disease control and prevention. possible side effects after getting a COVID-19 vaccine. 2022. https://www.cdc.gov/coronavirus/2019-ncov/vaccines/expect/after.html#print. Accessed 29 Mar 2022.
Smith TRF, Patel A, Ramos S, et al. Immunogenicity of a DNA vaccine candidate for COVID-19. Nat Commun. 2020;11(1):2601.
Shastri J, Parikh S, Aggarwal V, et al. Severe SARS-CoV-2 breakthrough reinfection with delta variant after recovery from breakthrough infection by alpha variant in a fully vaccinated health worker. Front Med (Lausanne). 2021;8:737007.
CDC. U.S. National Center for Immunization Respiratory Diseases, Division of Viral Diseases. Science Brief: SARS-CoV-2 Infection-induced and Vaccine-induced Immunity. https://www.cdc.gov/coronavirus/2019-ncov/science/science-briefs/vaccine-induced-immunity.html. Accessed 20 Oct 2021.
Abu-Raddad LJ, Chemaitelly H, Ayoub HH, Yassine HM, Benslimane FM, Al Khatib HA, Tang P, Hasan MR, Coyle P, Al Kanaani Z, Al Kuwari E, Jeremijenko A, Kaleeckal AH, Latif AN, Shaik RM, Abdul Rahim HF, Nasrallah GK, Al Kuwari MG, Butt AA, Al Romaihi HE, Al-Thani MH, Al Khal A, Bertollini R. Association of prior SARS-CoV-2 infection with risk of breakthrough infection following mRNA vaccination in Qatar. JAMA. 2021;326(19):1930–9.
Lee JS, Kim SY, et al. Evidence of Severe Acute Respiratory Syndrome Coronavirus 2 reinfection after recovery from mild Coronavirus Disease 2019. Clin Infect Dis. 2021;73(9):E3002–8.
Tillett RL, Sevinsky JR, Hartley PD, et al. Genomic evidence for reinfection with SARS-CoV-2: a case study. Lancet Infect Dis. 2021;21(1):52–8.
Abu-Raddad LJ, Chemaitelly H, Malek JA, et al. Assessment of the Risk of Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Reinfection in an Intense Reexposure Setting. Clin Infect Dis. 2021;73(7):e1830–40.
Prado-Vivar B, Becerra-Wong M, Guadalupe JJ, et al. A case of SARS-CoV-2 reinfection in Ecuador. Lancet Infect Dis. 2021;21(6):e142.
Larson D, Brodniak SL, Voegtly LJ, et al. A case of early reinfection with Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2). Clin Infect Dis. 2021;73(9):e2827–8.
Goldman JD, Wang K, Roltgen K, Nielsen SCA, Roach JC, Naccache SN, Yang F, Wirz OF, Yost KE, Lee JY, Chun K, Wrin T, Petropoulos CJ, Lee I, Fallen S, Manner PM, Wallick JA, Algren HA, Murray KM, Su Y, Hadlock J, Jeharajah J, Berrington WR, Pappas GP, Nyatsatsang ST, Greninger AL, Satpathy AT, Pauk JS, Boyd SD, Heath JR. Reinfection with SARS-CoV-2 and Failure of Humoral Immunity: a case report. medRxiv [Preprint]. 2020 Sep 25:2020.09.22.20192443. https://doi.org/10.1101/2020.09.22.20192443.
Gupta V, Bhoyar RC, Jain A, Srivastava S, Upadhayay R, Imran M, Jolly B, Divakar MK, Sharma D, Sehgal P, Ranjan G, Gupta R, Scaria V, Sivasubbu S. Asymptomatic reinfection in 2 healthcare workers from India with genetically distinct Severe Acute Respiratory Syndrome Coronavirus 2. Clin Infect Dis. 2021;73(9):e2823–5.
To KK, Hung IF, Ip JD, Chu AW, Chan WM, Tam AR, Fong CH, Yuan S, Tsoi HW, Ng AC, Lee LL, Wan P, Tso EY, To WK, Tsang DN, Chan KH, Huang JD, Kok KH, Cheng VC, Yuen KY. Coronavirus Disease 2019 (COVID-19) re-infection by a phylogenetically distinct Severe Acute Respiratory Syndrome Coronavirus 2 strain confirmed by whole genome sequencing. Clin Infect Dis. 2021;73(9):e2946–51. https://doi.org/10.1093/cid/ciaa1275.
Acknowledgements
We would like to acknowledge Tehran University of Medical Sciences for their educational support in time of COVID-19 pandemic. This article is the outcome of an in-house, financially non-supported study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Declaration of interests
The authors have no relevant affiliations or financial involvement with any organization or entity with a financial interest in or financial conflict with the subject matter or materials discussed in the manuscript. This includes employment, consultancies, honoraria, stock ownership or options, expert testimony, grants or patents received or pending, or royalties.
Ethics approval
Not applicable.
Informed consent
Not applicable.
Conflict of interest
The authors declare that they have no conflict of interest.capture
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Abdolmaleki, G., Taheri, M.A., Paridehpour, S. et al. A comparison between SARS-CoV-1 and SARS-CoV2: an update on current COVID-19 vaccines. DARU J Pharm Sci 30, 379–406 (2022). https://doi.org/10.1007/s40199-022-00446-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40199-022-00446-8