Abstract
Background
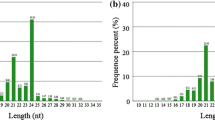
MicroRNAs (miRNAs) are about 21 snucleotide (nt) long, non-coding RNAs that play an important role in plant abiotic stress responses. Chinese jujube is a native fruit tree in China, which is also an admittedly drought-resistant plant. But the drought-related miRNAs have little been reported in jujube.
Objective
To identify possibly drought-responsive microRNAs and their target genes in Chinese Jujube.
Methods
Twelve small RNA libraries were constructed from two jujube genotypes both drought treated and control samples with three replicates to identify known and novel miRNAs in Chinese Jujube, DESeq2 was used to identify expression pattern of miRNAs between drought treatment and control samples, TargetFinder program was used to predict potential target genes of conserved and novel miRNAs, RT-qPCR were used to analysis the expression levels of drought-related miRNAs and their potential targets. The RNA ligase-mediated RLM-5′ RACE experiments were performed to validate predicted target genes of drought-related miRNAs.
Results
43 known miRNAs and 431 novel miRNAs were identified in Chinese jujube. Expression analysis showed that 28 miRNAs were differential expressed under drought stress in jujube variety “Dongzao”, including 21 up-regulated miRNAs and 7 down-regulated miRNAs, 61 miRNAs were differential expressed under drought stress in Chinese jujube variety “Zanhuangdazao”, including 23 up-regulated miRNAs and 37 down-regulated miRNAs. Depend on miRNAs target prediction, functional annotation and expression analysis, we identified 9 drought-related miRNAs, and 7 target genes of 6 miRNAs were confirmed using the modified 5′-RACE method. Also, RT-qPCR analyses revealed that relative expression of those miRNAs and their targets have negative tendency.
Conclusion
We identified 6 drought-related miRNAs by high-throughout sequencing and target gene annotation from Chinese jujube, and targets of those miRNAs were confirmed by the modified 5′-RACE method. These findings provide molecular evidence for enhancing drought tolerance in Chinese jujube and other plants.





Similar content being viewed by others
References
Akdogan G, Tufekci ED, Uranbey S, Unver T (2016) miRNA-based drought regulation in wheat. Funct Integr Genomics 16(3):221–233
Allen E, Xie Z, Gustafson AM, Carrington JC (2005) microRNA-directed phasing during trans-acting siRNA biogenesis in plants. Cell 121:207–221
Apweiler R, Bairoch A, Wu CH (2004) UniProt: the universal protein knowledgebase. Nucleic Acids Res 32:D115–D119
Ashraf M (2010) Inducing drought tolerance in plants: recent advances. Biotechnol Adv 28:169–183
Balusamy SR, Rahimi S, Yang DC (2019) Characterization of squalene-induced PgCYP736B involved in salt tolerance by modulating key genes of abscisic acid biosynthesis. Int J Biol Macromol 121:796–805
Barrera-Figueroa BE, Gao L, Diop NN, Wu Z, Ehlers JD, Roberts PA, Close TJ, Zhu JK, Liu R (2011) Identification and comparative analysis of drought-associated microRNAs in two cowpea genotypes. BMC Plant Biol 11:127
Bartel DP (2009) MicroRNAs: target recognition and regulatory functions. Cell 136:215–233
Bechtold U, Albihlal WS, Lawson T, Michael JF, Sparrow PA, Richard F, Persad R et al (2013) Arabidopsis HEAT SHOCK TRANSCRIPTION FACTORA1b overexpression enhances water productivity, resistance to drought, and infection. J Exp Bot 64:3467–3481
Bu J, Zhao J, Liu ML (2016) Expression stabilities of candidate reference genes for RT-qPCR in Chinese jujube (Ziziphus jujuba Mill.) under a variety of conditions. PLoS ONE 11(4):e0154212
Cao ZH, Zhang SZ, Wang RK, Zhang RF, Hao YJ (2013) Genome wide analysis of the apple MYB transcription factor family allows the identification of MdoMYB121 gene conferring abiotic stress tolerance in plants. PLoS ONE 8:e69955
Chaves MM, Oliveira MM (2004) Mechanisms underlying plant resilience to water deficits: prospects for water-saving agriculture. J Exp Bot 55(407):2365–2384
Cheng XW, Ma YP, Xu CX (2013) Absorption, translocation and distribution characteristics of salt ions of Chinese jujube and sour jujube under iso-osmotic potential drought, salt, and alkaline stresses. J Cent South Univ Fore Technol 33:20–25
Ci D, Song YP, Tian M, Zhang DQ (2015) Methylation of miRNA genes in the response to temperature stress in Populus simonii. Front Plant Sci 6:921
Cuming AC, Cho SH, Kamisugi Y, Graham H, Quatrano RS (2007) Microarray analysis of transcriptional responses to abscisic acid and osmotic, salt, and drought stress in the moss, Physcomitrella patens. New Phytol 176:275–287
Deng YY, Li JQ, Wu SF (2006) Integrated nr database in protein annotation system and its localization. Comput Eng 32(5):71–74
Dubos C, Stracke R, Grotewold E, Weisshaar B, Martin C, Lepiniec L (2010) MYB transcription factors in Arabidopsis. Trends Plant Sci 15:573–581
Eddy SR (1998) Profile hidden Markov models. Bioinformatics 14(9):755–763
Eldem V (2012) Genome-wide identifification of miRNAs responsive to drought in peach (Prunus persica) by high-throughput deep sequencing. PLoS ONE 7:e50298
Fedoroff NV, Battisti DS, Beachy RN, Cooper PJ, Fischhoff DA, Hodges CN, Knauf VC, Lobell D, Mazur BJ, Molden D (2010) Radically rethinking agriculture for the 21st century. Science 327:833–834
Friedlander MR, Mackowiak SD, Li N, Chen W, Rajewsky N (2012) miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res 40:37–52
Guo HS, Xie Q, Fei JF, Chua NH (2005) MicroRNA directs mRNA cleavage of the transcription factor NAC1 to down regulate auxin signals for Arabidopsis lateral root development. Plant Cell 17:1376–1386
Guo M, Liu JH, Ma X, Luo DX, Gong ZH, Lu MH (2016) The plant heat stress transcription factors (HSFs): structure, regulation, and function in response to abiotic stresses. Front Plant Sci 7:114
Hackenberg M, Gustafson P, Langridge P, Shi BJ (2015) Differential expression of microRNAs and other small RNAs in barley between water and drought conditions. Plant Biotechnol J 13:2–13
Hartl FU (1996) Molecular chaperones in cellular protein folding. Nature 381(6583):571–579
Kasuga M, Liu Q, Miura S, Yamaguchi-Shinozaki K, Shinozaki K (2007) Improving plant drought, salt, and freezing tolerance by gene transfer of a single stress-inducible transcription factor. Nat Biotechnol 17:287–291
Koonin EV, Fedorova ND, Jackson JD, Jacobs AR, Krylov DM, Makarova KS, Mazumder R et al (2004) A comprehensive evolutionary classification of proteins encoded in complete eukaryotic genomes. Genome Biol 5(2):R7
Kropat J (2005) A regulator of nutritional copper signaling in Chlamydomonas is an SBP domain protein that recognizes the GTAC core of copper response element. Proc Natl Acad Sci USA 102:18730–18735
Kurihara Y, Watanabe Y (2004) Arabidopsis micro-RNA biogenesis through Dicer-like 1 protein functions. Proc Natl Acad Sci USA 101(34):12753–12758
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10:R25
Lauter N, Kampani A, Carlson S, Goebel M, Moose SP (2005) MicroRNA172 down-regulates glossy 15 to promote vegetative phase change in maize. Proc Natl Acad Sci 102:9412–9417
Li G, Niu JF (2006) Study on building jujube plantation using wild jujube seeds in the middle barren areas of Ningxia province. China Fruits 2:58
Li JW, Fan LP, Ding SD, Ding XL (2007) Nutritional composition of five cultivars of Chinese jujube. Food Chem 103:454–460
Li BS, Qin YR, Duan H, Yin WL, Xia XL (2011) Genome-wide characterization of new and drought stress responsive microRNAs in Populus euphratica. J Exp Bot 62:3765–3779
Liu J, Liu H, Ma L, Wang S, Gao J, Li Y (2014) A Chinese jujube (Ziziphus jujuba Mill.) fruit-expressed sequence tag (EST) library: annotation and EST-SSR characterization. Sci Hortic 165:99–105
Liu H, Able AJ, Able JA (2016) SMARTER de-stressed cereal breeding. Trends Plant Sci 21:909–925
Long SP, Ort DR (2010) More than taking the heat: crops and global change. Curr Opin Plant Biol 13(3):241–248
Long RC, Li MN, Kang JM, Zhang TJ, Sun Y, Yang QC (2015) Small RNA deep sequencing identififies novel and salt-stress-regulated microRNAs from roots of Medicago sativa and Medicago truncatula. Physiol Plant 154:13–27
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15(12):550
Ma F, Huang J, Yang J, Zhou J, Sun Q, Sun J (2020) Identification, expression and miRNA targeting of auxin response factor genes related to phyllody in the witches’ broom disease of jujube. Gene 746:144656
Mallory AC, Dugas DV, Bartel DP, Bartel B (2004) MicroRNA regulation of NAC domaintargets is required for proper formation and separation of adjacent embryonic, vegetative, and floral organs. Curr Biol 14:1035–1046
Manassero NU, Viola IL, Welchen E, Gonzalez DH (2013) TCP transcription factors: architectures of plant form. Biomol Concepts 4:111–127
Mao G, Seebeck T, Schrenker D, Yu O (2013) CYP709B3, a cytochrome P450 monooxygenase gene involved in salt tolerance in Arabidopsis thaliana. BMC Plant Biol 13:169–182
Martin-Trillo M, Cubas P (2009) TCP genes: a family snapshot ten years later. Trends Plant Sci 15:31–39
Michael A, Catherine AB, Judith AB (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25:25-C29
Minoru K, Susumu G, Shuichi K (2004) The KEGG resource for deciphering the genome. Nucleic Acids Res 32(277):D280
Moldovan D, Spriggs A, Yang J, Pogson BJ, Dennis ES, Wilson IW (2010) Hypoxia-responsive microRNAs and trans-acting small interfering RNAs in Arabidopsis. J Exp Bot 61:165–177
Moxon S, Jing R, Szittya G, Schwach F, Rusholme-Pilcher RL (2008) Deep sequencing of tomato short RNAs identifies microRNAs targeting genes involved in fruit ripening. Genome Res 18:1602–1609
Mukhopadhyay P, Tyagi AK (2015) OsTCP19 influences developmental and abiotic stress signaling by modulating ABI4-mediated pathways. Sci Rep 5:9998
Niu CD, Li HY, Jiang LJ, Yan MJ, Li CY, Geng D, Xie YP, Yan Y, Shen XX, Chen PX, Dong J, Ma FW, Guan QM (2019) Genome-wide identification of drought responsive microRNAs in two sets of Malus from interspecific hybrid progenies. Hortic Res-Englang 6:75
Palatnik JF, Allen E, Wu X, Schommer C, Schwab R (2003) Control of leaf morphogenesis by microRNAs. Nature 425:257–263
Paul S, Datta SK, Datta K (2015) miRNA regulation of nutrient homeostasis in plants. Front Plant Sci 6:232
Rabbani MA, Maruyama K, Abe H, Khan MA, Katsura K, Ito Y, Yoshiwara K, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) Monitoring expression profiles of rice genes under cold, drought, and high-salinity stresses and abscisic acid application using cDNA microarray and RNA gel-blot analyses. Plant Physiol 133(4):1755–1767
Rhoades MW, Reinhart BJ, Lim LP, Burge CB, Bartel B (2002) Prediction of plant microRNA targets. Cell 110:513–520
Roman LT, Michael YG, Darren AN (2000) The COG database: a tool for genome scale analysis of protein functions and evolution. Nucleic Acids Res 128(1):33–36
Sahi C, Singh A, Kumar K, Blumwald E, Grover A (2006) Salt stress response in rice: genetics, molecular biology, and comparative genomics. Funct Integr Genomics 6:263–284
Seki M, Narusaka M, Ishida J, Nanjo T, Fujita M, Oono Y, Kamiya A, Nakajima M, Enju A, Sakurai T (2002) Monitoring the expression profiles of 7000 Arabidopsis genes under drought, cold and high-salinity stresses using a full-length cDNA microarray. Plant J 31(3):279–292
Shinozaki K, Yamaguchi-Shinozaki K (2007) Gene networks involved in drought stress response and tolerance. J Exp Bot 58:221–227
Shinozaki K, Yamaguchi-Shinozakiy K, Sekiz M (2003) Regulatory network of gene expression in the drought and cold stress responses. Curr Opin Plant Biol 6:410–417
Shiu SH, Bleecker AB (2003) Expansion of the receptor-like kinase/Pelle gene family and receptor-like proteins in Arabidopsis. Plant Physiol 132(2):530–543
Stief A (2014) Arabidopsis miR156 regulates tolerance to recurring environmental stress through SPL transcription factors. Plant Cell 26:1792–1807
Sunkar R, Zhu JK (2004) Novel and stress-regulated microRNAs and other small RNAs from Arabidopsis. Plant Cell 16:2001–2019
Umezawa T, Fujita M, Fujita Y, Yamaguchi-Shinozaki K, Shinozaki K (2006) Engineering drought tolerance in plants: discovering and tailoring genes to unlock the future. Curr Opin Biotechnol 17(2):113–122
Wang TZ, Chen L, Zhao MG, Tian QY, Zhang WH (2011) Identification of drought-responsive microRNAs in Medicago truncatula by genome-wide high-throughput sequencing. BMC Genomics 12:367
Wang C, Yang Y, Wang H, Ran X, Li B, Zhang J, Zhang H (2016) Ectopic expression of a cytochrome P450 monooxygenase gene PtCYP714A3 from Populus trichocarpa reduces shoot growth and improves tolerance to salt stress in transgenic rice. Plant Biotechnol J 14:1838–1851
Wu B, Wang M, Ma Y, Yuan L, Lu S (2012) High-throughput sequencing and characterization of the small RNA transcriptome reveal features of novel and conserved microRNAs in Panax ginseng. PLoS ONE 7:e44385
Yao Y (2010) Non-coding small RNAs responsive to abiotic stress in wheat (Triticum aestivum L.). Funct Integr Genom 10:187–190
Zhang L (2009) A genome-wide characterization of microRNA genes in maize. PLoS Genet 5:e1000716
Zhang R, Marshall D, Bryan GJ, Hornyik C (2013) Identification and characterization of miRNA transcriptome in potato by high-throughput sequencing. PLoS ONE 8:e57233
Zhao B, Liang R, Ge L, Li W, Xiao H (2007) Identification of drought-induced microRNAs in rice. Biochem Biophys Res Commun 354:585–590
Zhou LG, Liu YH, Liu ZC, Kong DY, Duan M, Luo LJ (2010) Genome-wide identification and analysis of drought-responsive microRNAs in Oryza sativa. J Exp Bot 61:4157–4168
Zhou ZS, Zeng HQ, Yang ZM (2012) Genome-wide identification of Medicago truncatula microRNAs and their targets reveals their differential regulation by heavy metal. Plant Cell Environ 35:86–99
Zhou M, Li D, Li Z, Hu Q, Yang C, Zhu L, Luo H (2013) Constitutive expression of a miR319 gene alters plant development and enhances salt and drought tolerance in transgenic creeping bentgrass. Plant Physiol 161:1375–1391
Acknowledgements
This work was sponsored by the Forestry and Grassland Science and Technology Innovation Project of Gansu Province (kjcx202006), National Natural Science Foundation of China (No. 31860400), and the 2021 Subsidy of Gansu Province Woody Oil Engineering Research Center.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: LZ YL JY. Performed the experiments: LZ YL JY. Analyzed the data: LZ YL JY. Contributed reagents/materials/analysis tools: HH QL JZ FW DW. Wrote the paper: LZ YL JY.
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
13258_2022_1274_MOESM1_ESM.xls
Supplementary file1 Supplementary Material 1: sRNAs annotation and distribution of twelve Chinese jujube samples (XLS 27 KB)
13258_2022_1274_MOESM3_ESM.xls
Supplementary file3 Supplementary Material 3: Expression analysis of known and novel miRNAs between control and drought treatment in two Chinese jujube varieties (XLS 56 KB)
13258_2022_1274_MOESM4_ESM.docx
Supplementary file4 Supplementary Material 4: The differentially expressed miRNAs between two Chinese jujube varieties (DOCX 39 KB)
Rights and permissions
About this article
Cite this article
Zhang, L., Li, Y., Yang, J. et al. Genome-wide identification of drought-responsive microRNAs and their target genes in Chinese jujube by deep sequencing. Genes Genom 45, 231–245 (2023). https://doi.org/10.1007/s13258-022-01274-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-022-01274-5