Abstract
Objectives: To conduct a systematic survey of published techniques for automated diagnosis and prognosis of COVID-19 diseases using medical imaging, assessing the validity of reported performance and investigating the proposed clinical use-case. To conduct a scoping review into the authors publishing such work. Methods: The Scopus database was queried and studies were screened for article type, and minimum source normalized impact per paper and citations, before manual relevance assessment and a bias assessment derived from a subset of the Checklist for Artificial Intelligence in Medical Imaging (CLAIM). The number of failures of the full CLAIM was adopted as a surrogate for risk-of-bias. Methodological and performance measurements were collected from each technique. Each study was assessed by one author. Comparisons were evaluated for significance with a two-sided independent t-test. Findings: Of 1002 studies identified, 390 remained after screening and 81 after relevance and bias exclusion. The ratio of exclusion for bias was 71%, indicative of a high level of bias in the field. The mean number of CLAIM failures per study was 8.3 ± 3.9 [1,17] (mean ± standard deviation [min,max]). 58% of methods performed diagnosis versus 31% prognosis. Of the diagnostic methods, 38% differentiated COVID-19 from healthy controls. For diagnostic techniques, area under the receiver operating curve (AUC) = 0.924 ± 0.074 [0.810,0.991] and accuracy = 91.7% ± 6.4 [79.0,99.0]. For prognostic techniques, AUC = 0.836 ± 0.126 [0.605,0.980] and accuracy = 78.4% ± 9.4 [62.5,98.0]. CLAIM failures did not correlate with performance, providing confidence that the highest results were not driven by biased papers. Deep learning techniques reported higher AUC (p < 0.05) and accuracy (p < 0.05), but no difference in CLAIM failures was identified. Interpretation: A majority of papers focus on the less clinically impactful diagnosis task, contrasted with prognosis, with a significant portion performing a clinically unnecessary task of differentiating COVID-19 from healthy. Authors should consider the clinical scenario in which their work would be deployed when developing techniques. Nevertheless, studies report superb performance in a potentially impactful application. Future work is warranted in translating techniques into clinical tools.
Similar content being viewed by others
Introduction
The novel coronavirus, SARS-Cov-2 and its associated disease, COVID-19, have presented a significant and urgent threat to public health while simultaneously disrupting healthcare systems. Despite being more than 2 years since the beginning of the pandemic, outbreaks continue to threaten to overwhelm healthcare systems, and viral variants continue to introduce uncertainty [1]. Fast and accurate diagnostic and prognostic capability help quickly determine which patients need to be isolated and informs triage of patients. Reverse-transcription polymerase chain reaction (RT-PCR) is the current clinical standard for diagnosis of COVID-19, however, its low sensitivity often necessitates repeat testing [2] taking additional time. This has led to the suggestion that there is a role for radiology in diagnosing COVID-19.
Radiological professional bodies have generally recommended against the use of imaging for screening in COVID-19 but recognise the role of incidental findings and for disease staging. Early in the pandemic, the use of computed tomography (CT) for diagnosis and screening was discussed in the context of shortages of RT-PCR test kits and poor sensitivity [3]. In March of 2020, a consensus report was released [4], endorsed by the Society of Thoracic Radiology, the American College of Radiology and the Radiological Society of North America (RSNA), recommending against the use of chest CT for screening due to a low negative predictive value, but also partly due to a lack of evidence early in the pandemic. The Royal Australian and New Zealand College of Radiologists released their advice in April of 2020, which remains current, recommending against the use of chest radiograph for screening but recommending for the use of CT for staging [5]. The report, however, stops short of recommending a severity scale. By June of 2020, the World Health Organisation recommended the use of radiological imaging: (1) for diagnostic purposes in symptomatic patients when RT-PCR is not available, is available but results are delayed and when RT-PCR is negative but there is high clinical suspicion of COVID-19; (2) for triage purposes when deciding to admit to hospital and/or intensive care unit (ICU); and (3) for staging purposes when deciding appropriate therapeutic management [6]. The most recent version of the Cochrane review on the topic suggest that CT and chest X-ray (CXR) are moderately sensitive and specific to the diagnosis of COVID-19, whereas ultrasound is sensitive but not specific to the diagnosis of COVID-19 [7]. This novel application of radiology has spurred an interest in the application of machine learning techniques to automate the image interpretation tasks.
Many investigators have proposed techniques in a wide range of applications to automate image interpretation in imaging of COVID-19, including segmentation of COVID-19 related lesions, typically ground-glass opacities (GGOs), diagnosis, staging of the current disease progression and prognosis of likely future disease progression. However, the field has inspired controversy. DeGrave et al. [8] demonstrated that combining data from multiple sources, in particular where data from different classes have different acquisition and pre-processing parameters, led to a significant bias that artificially improved the measured performance in many studies. Garcia Santa Cruz et al. [9] presented a review of public CXR datasets, concluding that the most popular datasets used in the literature were at a high risk of introducing bias into reported results.
Many other reviews have been introduced on the topic, we now introduce the seminal ones. Shi et al. [10] presented a narrative review very early in the pandemic (published April of 2020) of machine learning techniques for segmentation of COVID-19-related lesions and for diagnosis, staging and prognosis of COVID-19 using CXR and CT. However, this early review did not consider potential study bias in its papers. Others have presented systematic reviews [11, 12] that, while following a more rigorous approach to inclusion also failed to asses bias when assessing results. Wynants et al. [13] present a broadly-scoped systematic review for prediction models in COVID-19, leveraging the prediction model risk of bias assessment tool (PROBAST) [14]. They reported high risk of bias across the field. Roberts et al. [15] presented a systematic review of machine learning techniques applied to CXR and CT imaging, published up to the 3rd of October, 2020, assessing bias using the Checklist for Artificial Intelligence in Medical Imaging (CLAIM) [16], Radiomics Quality Score (RQS) [17] and PROBAST [14] and reporting methodological and dataset trends. They use this to develop a set of recommendations for authors in the field.
In this review, we use similar techniques to those presented by Roberts et al. [15]. Rather than assessing papers on separate criteria, RQS and CLAIM, we assess all papers with CLAIM. We also aim to present a richer analysis of techniques and their performance, and to provide an update, including publications until 31st October, 2021. We also introduce an analysis of authors and institutions in the field, in the hope that it encourages and facilitates further collaboration.
Research questions:
-
Which techniques are most successful in differentiating COVID-19?
-
What are the clinical requirements driving the development of these tools? How would such techniques be implemented clinically?
-
Who is publishing this in this field?
Methodology
Study selection
The inclusion criteria for the review are:
-
(1)
Studies that aim to automatically (allowing for manual contouring as a preprocessing step under the assumption this could be automated) diagnose, stage or prognose COVID-19 or segment lesions associated with COVID-19; and
-
(2)
Studies that use medical imaging or signals, including CXR, CT, ultrasound, magnetic resonance imaging (MRI), or electrocardiograph (ECG) as input to their model.
rscopus version 0.6.6 [18] was used to retrieve articles according to the search criteria outlined in Panel 1. The search was performed on the 19th November, 2021. Papers meeting the inclusion criteria that were identified during the investigation but not identified in the search were also included in the study.
Panel 1: Scopus search criteria
TITLE-ABS-KEY ( ( covid OR coronavirus ) AND ( ( chest W/5 xray ) OR “computed tomography” OR ultrasound OR “magnetic resonance” OR mr OR mri OR ecg OR electrocardiograph* ) AND ( diagnos* OR staging OR identif* OR response OR prognos* OR segment* ) AND ( learn* OR convolutional OR network OR radiomic*) )
Exclusion criteria were also imposed to eliminate studies that exhibited or were likely to exhibit a high risk of bias:
-
(1)
Studies from journals with a source normalized impact per paper (SNIP), as measured in 2021, less than 1 were excluded. SNIP is a metric introduced by Scopus that measures contextual impact, normalising between fields with different citation rates. This process was manually checked by two of the authors, and journals that were likely to publish relevant studies and reputable within their fields, that would be eliminated, were included.
-
(2)
Studies that were more than 90 days old and had not attracted any citations were excluded. This criteria is included to automatically filter articles which the scientific community has deemed uninteresting, under the assumption that in such a fast moving field, 90 days should be adequate to have attracted at least one citation.
-
(3)
Studies with metadata indicating that they were Editorials, Reviews, Notes or Letters were excluded.
-
(4)
Studies where application to COVID-19 is secondary and not the primary focus of the paper were excluded.
-
(5)
Studies not meeting the minimum risk of bias assessment (see “Bias assessments” section) were excluded.
Remaining studies were assigned amongst reviewing authors, and each study was reviewed by one author, who assessed for minimum risk-of-bias, and extracted data. Studies were not de-identified before analysis.
Bias assessments
Due to reports of a high risk-of-bias in the field [9, 13, 15], we include a bias assessment. Improper study design, data collection, data partitioning and statistical methods can lead to misleading reported results [14]. This commonly manifests as a positive bias because authors (rightly) attempt to improve the performance of their proposed techniques.
The CLAIM checklist was completed for all included papers [16]. All 42 checklist items were given either a pass or fail score, or a “not applicable” score which did not count towards the failure count in cases where the checklist item was not applicable to the paper. The number of failure scores was used as a measure for bias. Similar to Roberts et al. [15], we impose a subset of CLAIM, items 7, 9, 20, 21, 22, 25, 26 and 28, as a minimum risk of bias. Any papers that did not meet all subset checklist items were excluded. CLAIM checklist reports from Roberts et al. [15] were merged and used where available to avoid duplication.
Extracted data
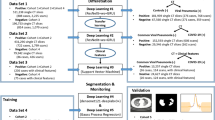
Methodological and performance results were collected per technique, where each study presents one or more technique. When multiple techniques were introduced in each study, only the highest performing technique was surveyed, unless the techniques filled different purposes (e.g., one study presenting a segmentation and diagnostic technique) or different contexts (e.g., different available clinical data to augment image input) (Table 1).
Analysis of studies
Accuracy and area under the curve (AUC) of the receiver operating characteristic (ROC), where reported, were used for performance comparison. Statistical significance was measured throughout this review using two-sided independent t-tests, with a significance threshold of p < 0.05. No adjustments were made for multiple comparisons.
Analysis of authors and publishers
Author, institution and publication metadata were extracted using rscopus 0.6.6 [18] and used to compute author h-indices. A co-author network was generated with tidygraph 1.2.0 [19] by linking authors that had published together, and the most central authors identified using the betweenness centrality.
Results
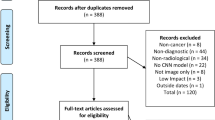
Of 1002 studies identified, 282 were assessed against the required subset of the CLAIM checklist for exclusion, after which 81 studies were included in the study (Fig. 1). A list of identified and included studies are available in Supplementary 1, Table S1, and the full set of studies identified and collected data are available in Supplementary 2. CLAIM 26 eliminated the most studies (Fig. 2, left), which pertains to the evaluation of the best-performing model. Most papers failing this subset failed to evaluate against a separate test set after presenting multiple models. CLAIM 25 eliminated the next most studies, which required an adequate description of hyperparameter selection. Only one in four of papers met the inclusion criteria, and approximately one in four of papers failed a half or more of the required CLAIM subset (Fig. 2, right). From the 81 studies included, a total of 103 separate techniques were included.
Bias
Remaining CLAIM failures in the included articles are depicted in Fig. 3 (left). The count of failures for each article became the risk-of-bias surrogate, a histogram over all papers is shown in Fig. 3 (right). The mean number of failures was 8.3 \(\pm\) 3.9 standard deviation.
Methodologies
The majority, 58%, of techniques sought to solve a diagnosis task, attempting to classify COVID-19 disease from healthy patients and/or non-COVID-19 pneumonia (Fig. 4, left), versus 31% performing prognosis (where techniques performing both are counted in both). Of the 31% of techniques attempting to solve a prognosis task, the majority used an objective prognostic outcome measure (46% progression and 16% survival) rather than matching a clinical assessment.
(Left) Machine learning tasks attempted to be solved by techniques. (Top Right) A breakdown of Diagnosis and Diagnosis & Prognosis approaches by diagnostic outcome variable classes. (Bottom Right) A breakdown of Prognosis and Diagnosis & Prognosis approaches by prognostic outcome variable. The inner ring represents the number of classes, or continuous for regression tasks, and the outer ring represents the derivation of the outcome variable. See Table 1 for definitions of derivations
Most papers used CT images, either in 3D or as 2D slices, as model input, followed by CXR and US (Fig. 5, left). Only a small minority of papers included clinical features as input. Although MRI and ECG were explicitly included within the scope of the review, no techniques using these modalities were included. No MRI papers were identified, and none of the 3 identified ECG papers that progressed beyond screening met the inclusion criteria.
(Left) The distribution of modalities used for input to techniques. (Middle) The reported AUC and (Right) accuracy of techniques by modality. Only techniques reporting AUC or accuracy are included, respectively. Results of a two-sided independent t-test are give as ‘*’ for significance or ‘ns’ for no significance
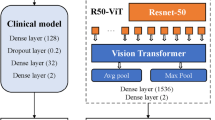
The majority of papers used a deep learning approach, the most common deep learning models used are listed in Fig. 6.
Performance
Performance is only reported here for studies where AUC or accuracy were described. The top-performing diagnostic and prognostic techniques are listed in Tables 2 and 3, respectively. Neither AUC (Fig. 7, left) nor accuracy (Fig. 7, right) significantly correlated with the number of CLAIM failures for diagnosis nor prognosis. There were no statistically significant differences between input modalities on performance (Fig. 5, middle and right), although CXR appeared to provide a higher AUC than CT, and US appeared to provide a lower accuracy than CT and CXR. Deep learning approaches had increased reported AUC (p = 0.04) and accuracy (p = 0.01), but no significant difference in bias was identified (Fig. 8).
Performance of techniques, as measured by AUC (left) and accuracy (right), plotted against CLAIM failures. Hue represents tasks, as indicated in the legend. Dashed lines indicate the mean regression for each of the tasks, and shading indicates the 95% confidence interval. All regression lines were compared with a two-sided independent t-test against a null hypothesis that gradient = 0, none of which reached significance
Comparison of (Left) AUC, (middle) accuracy and (Right) number of CLAIM fails between techniques leveraging deep learning and those leveraging classical machine learning and radiomics approaches. Results of a two-sided independent t-test are represented as ‘*’ for significance or ‘ns’ for no significance
Authors
The country of residence of authors tended to correlate with countries that were affected the most by the pandemic in early 2020 (Fig. 9).
A network analysis of connectivity between authors yielded 48 separate graphs of the 81 publications, depicted in Supplementary 1 Figure S1, and a subset in Fig. 10. The most productive research groups are summarised in Table 4.
Discussion
In this work, we present a systematic review of automated techniques for diagnosis, prognosis and segmentation of COVID-19 disease. Because the field has proven both popular and controversial, we used liberal exclusion criteria to reduce the number of lower-quality papers for manual review. In formulating the criteria, we assumed that impactful papers are likely to be published in highly cited publications and are likely to attract citations themselves. Studies published in journals with a SNIP below 1 were eliminated, which risks eliminating journals that aren’t ranked by Scopus. In order to reduce this risk, the list of eliminated journals was reviewed by all authors, and a consensus on non-indexed journals to include was reached. Further, studies that have been published for greater than 90 days yet hadn’t attracted any citations were eliminated, which risks eliminating unnoticed studies. Even after screening, 71% of papers were excluded during bias assessment (Figs. 1, 2), indicating that the majority of work in the field is at high risk of bias, including those published in reputable peer-reviewed publications.
Sources of bias
Datasets
Many studies use data from sources with minimal provenance and metadata, and often use data that was not intended for training diagnostic or prognostic tools. A number of datasets aggregate data from different sources, some of which may be aggregates themselves [9]; and many studies aggregate a number of datasets, either to increase their training size or to provide an independent test set. However, this causes a complex set or participants and leads to a high risk that the same images are present in the training and evaluation set. Other datasets present a series of CT slices without metadata indicating which images belong to which participants, leading to a high risk that adjacent axial slices from a participant may lie in the training and evaluation set. Any studies exhibiting these risks failed CLAIM 21.
Although it did not lead to exclusion in this review, some datasets also aggregate different classes from different sources. It has been established that this presents a high risk of bias, as networks are able to distinguish between classes using non-disease-related domain effects.
Data handling
Studies that did not split training and evaluation sets at the patient level also failed CLAIM 21. This mostly occurred in papers dealing with CT as 2D axial slices, some of which randomly allocated all 2D images between classes. CLAIM 26 was responsible for the most failures (45%, Figure < CLAIM subset >), which often indicated a failure to allocate an evaluation set for use after model selection.
Description of methods
The remaining CLAIM checklist items, 7, 9, 20, 22, 25 and 28, each related to adequately documenting methodology. This is important not only for reproducibility, which is important in technical publications to ensure the advancement of the field, but also could represent hidden bias. The field of machine learning requires attention to detail in implementation to prevent overfitting, data dredging or otherwise accidentally positively biasing results.
Study demographics
The majority (58%) of techniques sought to solve a diagnosis task. Although there has been limited need for diagnosis of COVID-19 using imaging, the potential for faster analysis compared with RT-PCR, especially when considering that consecutive negative RT-PCR testing is required for exclusion when the pre-test probability is high [39]. However, within this set, 38% only demonstrated an ability to differentiate COVID-19 from healthy individuals. Any clinically realistic scenario for deployment of such an algorithm would need to demonstrate an ability to aid in a differential diagnosis between similar diseases. Regardless, most professional bodies recommend the use of radiographic imaging in COVID-19 only for triage purposes [5, 6, 40] and therefore it is most likely more impactful for investigators to explore prognostic techniques.
CT scanning was the most popular modality, likely due to the image quality of tomographic imaging and the availability of public datasets. The additional context a 3D image can give may also have motivated the use of the modality, although many techniques only considered 2D axial sections. Given the clinical context and the fact that techniques are likely to be most useful during an outbreak, the use of CXR may be more convenient and practical. For example, clinical practice dictates that imaging rooms require an hour between patients for cleaning, a requirement that can be obviated with portable CXR that can move to the patient’s room [41]. Therefore, we suggest that future investigations may be more impactful in delivering a technique using CXR data, especially as no significant performance differences were seen between CXR and CT (Fig. 5).
It has been proposed that ultrasound analysis for COVID-19 could be valuable in rural and remote regions, and as a tool to facilitate social distancing in urban regions [42]. The relatively niche requirement means that systems for automated analysis of ultrasound are likely to be less impactful. This may be offset by the low cost of ultrasound, and the potential to deploy systems to developing countries. Other modalities, including MRI and even ECG, were explicitly included in the scope of this review, however no papers met the inclusion criteria for either. MRI generally yields poor contrast within the lung and provides few benefits over CT in this application. Some studies investigating ECG remain after the screening process, but either were excluded as they were not automated or did not meet the bias assessment requirements.
Study performance
Studies tended to report excellent diagnostic and prognostic performance based on imaging features. The top diagnostic techniques all reported AUC ≥ 0.98 and accuracy ≥ 96.8% (Table 2), while the prognostic techniques reported AUC ≥ 0.97 and accuracy ≥ 85.7% (Table 3). Further, these results were relatively stable across the number of CLAIM failures (Fig. 7), providing some confidence that the top results are not dominated by biased studies. Notably, though, the top performing prognostic techniques in Table 3 are binary classification tasks, which naturally yield higher metrics than those with more classes.
Observations
Data handling
Many studies used image storage formats that don’t meet medical imaging standards. Images may be stored at lower bit depth resolution, be stored using lossy compression, or be stored without requisite metadata. If these traits are consistent between classes, these issues are less likely to lead to a positive bias in reported results but may lead to lower performance. Similarly, many CT studies reported using per-image intensity normalisation for pre-processing. For quantitative modalities such as CT, this leads to a loss of information that the network is likely having to account for internally.
Input data
Studies that presented techniques under identical conditions with and without clinical data reported superior performance with the clinical data [28, 43]. This may be reporting bias, but it is likely that some combination of demographic, symptomatic and imaging data is likely to provide additional discrimination into the disease progression. Much of this information is relatively easily acquired, so there is little cost to include it.
Ethics
The majority of studies presenting novel datasets reported detail on the ethical approval. However, far fewer provided information on the consent given by participants, as required by CLAIM item 7. To be consistent with the analysis of Roberts et al., we have ignored this requirement, however we note this is an area to be improved in the medical imaging literature. Further, no studies that sourced from public datasets reported any ethical approval. The National Statement on Ethical Conduct in Human Research [44] outlines the definition of human data to include that sourced from public datasets.
Clinical translation
Few of the reviewed papers realistically considered clinical deployment. As Roberts et al. [15] highlight, no developed systems are ready to be deployed clinically, with one reason being the need to work with clinicians to ensure the developed algorithms are clinically relevant and implementable. This is highlighted by a review by Born et al. [45] who found that although 84% of clinical studies report the use of CT (with CXR only comprising 10% of studies), a much larger proportion of the AI papers were focused on X-ray. The same paper also emphasizes the need for additional stakeholder engagement, including patients, ethics committees, regulatory bodies, hospital administrators and clinicians. For clinical deployment medical imaging software generally requires validation through randomised control trials, regulatory certification (generally the software would be developed within an ISO1485 and IEC 62304 environment), and integration with existing clinical workflow (aligning with agreed standards for interoperability and upgradability, particularly the DICOM standard and required vendor tags).
Author demographics
We provide data on the authors (Fig. 9) and institutions (Table 4) publishing in the field as a landscape map for new authors. Most of the authors are located between China, The United States of America and Italy, and most of the most productive groups in China. Collaboration between groups predominantly occurred within the same country, except for a cluster of collaboration between Italy and the United States (Fig. 10).
Review limitations
Automatic filtering of studies using the SNIP of the published journal and number of citations for studies older than 90 days at the time of search was conducted. This was required in order to regulate the scope of the manually reviewed articles. This risked omitting rigorous papers that have not attracted scientific interest or are published in less circulated or newer journals. We believe this risk is low enough that the results presented are generalisable to the field.
In this work, we collect studies primarily from the Scopus database. While Scopus, alongside Web of Science, are historically the most widely used databases in bibliometric analysis, their coverage is not complete. Notwithstanding, Scopus shares 99.11% of its indexed journals with Web of Science and 96.61% with Dimensions. For this reason, we believe the methods in this review were valid and fit-for-purpose.
In this work, we use CLAIM as a surrogate measure for bias. CLAIM provides a prescriptive and objective criterion, well-suited to having a range of reviewers quickly and consistently assess a large number of papers. However, CLAIM is designed as a checklist of best practices, as opposed to an assessment of bias. The number of CLAIM failures should be interpreted by the reader as only an approximate measure of bias.
Conclusion
In this systematic review, we collected 1002 studies and have included 82 in the analysis after screening, relevance and bias assessment. A 71% exclusion ratio for bias despite extensive screening was indicative of a high level of risk-of-bias in the field. Commonly, publications sought to solve tasks with lower potential clinical impact, focusing on diagnosis rather than prognosis and differentiation of COVID-19 from controls rather than from other likely candidate diseases in a differential diagnosis. Similarly, clinical considerations and deployment were seldom discussed. Medical imaging standards were also regularly not met, with data sourced online without provenance and in compressed formats. Nevertheless, studies reported superb prognostic and diagnostic performance, and these results were robust amongst studies regardless of risk-of-bias or modality. Deep learning studies tended to report improved performance but did not report higher risk-of-bias compared with traditional machine learning approaches. We therefore conclude that the field has proven itself as a concept and that future work should focus on developing clinically useful and robust tools.
Data availability
Data is included as supplementary material in this article.
References
Karim SSA, Karim QA (2021) Omicron SARS-CoV-2 variant: a new chapter in the COVID-19 pandemic. Lancet. https://doi.org/10.1016/S0140-6736(21)02758-6
Xie X, Zhong Z, Zhao W, Zheng C, Wang F, Liu J (2020) Chest CT for typical coronavirus disease 2019 (COVID-19) pneumonia: relationship to negative RT-PCR testing. Radiology 296:E41–E45. https://doi.org/10.1148/radiol.2020200343
Kanne JP, Little BP, Chung JH, Elicker BM, Ketai LH (2020) Essentials for radiologists on COVID-19: an update—radiology scientific expert panel. Radiology 296:E113–E114. https://doi.org/10.1148/radiol.2020200527
Simpson S, Kay FU, Abbara S, Bhalla S, Chung JH, Chung M, Henry TS, Kanne JP, Kligerman S, Ko JP, Litt H (2020) Radiological Society of North America expert consensus document on reporting chest CT findings related to COVID-19: endorsed by the society of thoracic radiology, the American College of Radiology, and RSNA. Radiol Cardiothorac Imaging 2:e200152. https://doi.org/10.1148/ryct.2020200152
The Royal Australian and New Zealand College of Radiologists (2020) Guidelines for CT Chest and Chest Radiograph reporting in patients with suspected COVID-19 infection
Akl EA, Blažić I, Yaacoub S, Frija G, Chou R, Appiah JA, Fatehi M, Flor N, Hitti E, Jafri H, Jin Z-Y, Kauczor HU, Kawooya M, Kazerooni EA, Ko JP, Mahfouz R, Muglia V, Nyabanda R, Sanchez M, Shete PB, Ulla M, Zheng C, van Deventer E, del Perez MR (2021) Use of chest imaging in the diagnosis and management of COVID-19: a WHO rapid advice guide. Radiology 298:E63–E69. https://doi.org/10.1148/radiol.2020203173
Islam N, Ebrahimzadeh S, Salameh J-P, Kazi S, Fabiano N, Treanor L, Absi M, Hallgrimson Z, Leeflang MM, Hooft L, van der Pol CB, Prager R, Hare SS, Dennie C, Spijker R, Deeks JJ, Dinnes J, Jenniskens K, Korevaar DA, Cohen JF, den Bruel AV, Takwoingi Y, van de Wijgert J, Damen JA, Wang J, McInnes MD, Group CC-19 DTA (2021) Thoracic imaging tests for the diagnosis of COVID-19. Cochrane Database Syst Rev. https://doi.org/10.1002/14651858.CD013639.pub4
DeGrave AJ, Janizek JD, Lee S-I (2021) AI for radiographic COVID-19 detection selects shortcuts over signal. Nat Mach Intell 3:610–619. https://doi.org/10.1038/s42256-021-00338-7
Garcia Santa Cruz B, Bossa MN, Sölter J, Husch AD (2021) Public covid-19 X-ray datasets and their impact on model bias—a systematic review of a significant problem. Med Image Anal 74:102225. https://doi.org/10.1016/j.media.2021.102225
Shi F, Wang J, Shi J, Wu Z, Wang Q, Tang Z, He K, Shi Y, Shen D (2021) Review of artificial intelligence techniques in imaging data acquisition, segmentation, and diagnosis for COVID-19. IEEE Rev Biomed Eng 14:4–15. https://doi.org/10.1109/RBME.2020.2987975
Albahri OS, Zaidan AA, Albahri AS, Zaidan BB, Abdulkareem KH, Al-qaysi ZT, Alamoodi AH, Aleesa AM, Chyad MA, Alesa RM, Kem LC, Lakulu MM, Ibrahim AB, Rashid NA (2020) Systematic review of artificial intelligence techniques in the detection and classification of COVID-19 medical images in terms of evaluation and benchmarking: taxonomy analysis, challenges, future solutions and methodological aspects. J Infect Public Health 13:1381–1396. https://doi.org/10.1016/j.jiph.2020.06.028
Alabool H, Alarabiat D, Abualigah L, Habib M, Khasawneh AM, Alshinwan M, Shehab M (2021) Artificial intelligence techniques for containment COVID-19 pandemic: a systematic review. Syst Rev. https://doi.org/10.21203/rs.3.rs-30432/v1
Wynants L, Calster BV, Collins GS, Riley RD, Heinze G, Schuit E, Bonten MMJ, Dahly DL, Damen JA, Debray TPA, de Jong VMT, Vos MD, Dhiman P, Haller MC, Harhay MO, Henckaerts L, Heus P, Kammer M, Kreuzberger N, Lohmann A, Luijken K, Ma J, Martin GP, McLernon DJ, Navarro CLA, Reitsma JB, Sergeant JC, Shi C, Skoetz N, Smits LJM, Snell KIE, Sperrin M, Spijker R, Steyerberg EW, Takada T, Tzoulaki I, van Kuijk SMJ, van Bussel BCT, van der Horst ICC, van Royen FS, Verbakel JY, Wallisch C, Wilkinson J, Wolff R, Hooft L, Moons KGM, van Smeden M (2020) Prediction models for diagnosis and prognosis of covid-19: systematic review and critical appraisal. BMJ 369:m1328. https://doi.org/10.1136/bmj.m1328
Moons KGM, Wolff RF, Riley RD, Whiting PF, Westwood M, Collins GS, Reitsma JB, Kleijnen J, Mallett S (2019) PROBAST: a tool to assess risk of bias and applicability of prediction model studies: explanation and elaboration. Ann Intern Med 170:W1–W33. https://doi.org/10.7326/M18-1377
Roberts M, Driggs D, Thorpe M, Gilbey J, Yeung M, Ursprung S, Aviles-Rivero AI, Etmann C, McCague C, Beer L, Weir-McCall JR, Teng Z, Gkrania-Klotsas E, Rudd JHF, Sala E, Schönlieb C-B (2021) Common pitfalls and recommendations for using machine learning to detect and prognosticate for COVID-19 using chest radiographs and CT scans. Nat Mach Intell 3:199–217. https://doi.org/10.1038/s42256-021-00307-0
Mongan J, Moy L, Kahn CE (2020) Checklist for artificial intelligence in medical imaging (CLAIM): a guide for authors and reviewers. Radiol Artif Intell 2:e200029. https://doi.org/10.1148/ryai.2020200029
Lambin P, Leijenaar RTH, Deist TM, Peerlings J, de Jong EEC, van Timmeren J, Sanduleanu S, Larue RTHM, Even AJG, Jochems A, van Wijk Y, Woodruff H, van Soest J, Lustberg T, Roelofs E, van Elmpt W, Dekker A, Mottaghy FM, Wildberger JE, Walsh S (2017) Radiomics: the bridge between medical imaging and personalized medicine. Nat Rev Clin Oncol 14:749–762. https://doi.org/10.1038/nrclinonc.2017.141
Muschelli III J (2018) Gathering bibliometric information from the Scopus API using rscopus. R J
Wickham H, Averick M, Bryan J, Chang W, McGowan LD, François R, Grolemund G, Hayes A, Henry L, Hester J (2019) Welcome to the Tidyverse. J Open Source Softw 4:1686
Zheng W, Yan L, Gou C, Zhang Z-C, Zhang JJ, Hu M, Wang F-Y (2021) Learning to learn by yourself: unsupervised meta-learning with self-knowledge distillation for COVID-19 diagnosis from pneumonia cases. Int J Intell Syst 36:4033–4064. https://doi.org/10.1002/int.22449
Yang X, He X, Zhao J, Zhang Y, Zhang S, Xie P (2020) COVID-CT-dataset: a CT scan dataset about COVID-19. http://arxiv.org/200313865
Han Z, Wei B, Hong Y, Li T, Cong J, Zhu X, Wei H, Zhang W (2020) Accurate screening of COVID-19 using attention-based deep 3D multiple instance learning. IEEE Trans Med Imaging 39:2584–2594. https://doi.org/10.1109/TMI.2020.2996256
Das D, Santosh KC, Pal U (2020) Truncated inception net: COVID-19 outbreak screening using chest X-rays. Phys Eng Sci Med 43:915–925. https://doi.org/10.1007/s13246-020-00888-x
Cohen JP, Dao L, Roth K, Morrison P, Bengio Y, Abbasi AF, Shen B, Mahsa HK, Ghassemi M, Li H (2020) Predicting covid-19 pneumonia severity on chest x-ray with deep learning. Cureus 12:e9448
Jaeger S, Candemir S, Antani S, Wáng Y-XJ, Lu P-X, Thoma G (2014) Two public chest X-ray datasets for computer-aided screening of pulmonary diseases. Quant Imaging Med Surg 4:47577–47477
Kermany DS, Goldbaum M, Cai W, Valentim CCS, Liang H, Baxter SL, McKeown A, Yang G, Wu X, Yan F, Dong J, Prasadha MK, Pei J, Ting MYL, Zhu J, Li C, Hewett S, Dong J, Ziyar I, Shi A, Zhang R, Zheng L, Hou R, Shi W, Fu X, Duan Y, Huu VAN, Wen C, Zhang ED, Zhang CL, Li O, Wang X, Singer MA, Sun X, Xu J, Tafreshi A, Lewis MA, Xia H, Zhang K (2018) Identifying medical diagnoses and treatable diseases by image-based deep learning. Cell 172:1122–1131. https://doi.org/10.1016/j.cell.2018.02.010
Wang X, Jiang L, Li L, Xu M, Deng X, Dai L, Xu X, Li T, Guo Y, Wang Z, Dragotti PL (2021) Joint learning of 3D lesion segmentation and classification for explainable COVID-19 diagnosis. IEEE Trans Med Imaging 40:2463–2476. https://doi.org/10.1109/TMI.2021.3079709
Liu H, Ren H, Wu Z, Xu H, Zhang S, Li J, Hou L, Chi R, Zheng H, Chen Y, Duan S, Li H, Xie Z, Wang D (2021) CT radiomics facilitates more accurate diagnosis of COVID-19 pneumonia: compared with CO-RADS. J Transl Med 19:29. https://doi.org/10.1186/s12967-020-02692-3
Karakanis S, Leontidis G (2021) Lightweight deep learning models for detecting COVID-19 from chest X-ray images. Comput Biol Med 130:104181. https://doi.org/10.1016/j.compbiomed.2020.104181
Jin W, Dong S, Dong C, Ye X (2021) Hybrid ensemble model for differential diagnosis between COVID-19 and common viral pneumonia by chest X-ray radiograph. Comput Biol Med 131:104252. https://doi.org/10.1016/j.compbiomed.2021.104252
Wang X, Peng Y, Lu L, Lu Z, Bagheri M, Summers RM (2019) ChestX-ray: hospital-scale chest X-ray database and benchmarks on weakly supervised classification and localization of common thorax diseases. In: Lu L, Wang X, Carneiro G, Yang L (eds) Deep learning and convolutional neural networks for medical imaging and clinical informatics. Springer, Cham, pp 369–392
Tang Z, Zhao W, Xie X, Zhong Z, Shi F, Ma T, Liu J, Shen D (2021) Severity assessment of COVID-19 using CT image features and laboratory indices. Phys Med Biol 66:035015. https://doi.org/10.1088/1361-6560/abbf9e
Wu Q, Wang S, Li L, Wu Q, Qian W, Hu Y, Li L, Zhou X, Ma H, Li H, Wang M, Qiu X, Zha Y, Tian J (2020) Radiomics analysis of computed tomography helps predict poor prognostic outcome in COVID-19. Theranostics 10:7231–7244. https://doi.org/10.7150/thno.46428
Li Z, Zhong Z, Li Y, Zhang T, Gao L, Jin D, Sun Y, Ye X, Yu L, Hu Z, Xiao J, Huang L, Tang Y (2020) From community-acquired pneumonia to COVID-19: a deep learning-based method for quantitative analysis of COVID-19 on thick-section CT scans. Eur Radiol 30:6828–6837. https://doi.org/10.1007/s00330-020-07042-x
Elsharkawy M, Sharafeldeen A, Taher F, Shalaby A, Soliman A, Mahmoud A, Ghazal M, Khalil A, Alghamdi NS, Razek AAKA, Alnaghy E, El-Melegy MT, Sandhu HS, Giridharan GA, El-Baz A (2021) Early assessment of lung function in coronavirus patients using invariant markers from chest X-rays images. Sci Rep 11:12095. https://doi.org/10.1038/s41598-021-91305-0
Lu Wang L, Lo K, Chandrasekhar Y, Reas R, Yang J, Eide D, Funk K, Kinney R, Liu Z, Merrill W, Mooney P, Murdick D, Rishi D, Sheehan J, Shen Z, Stilson B, Wade AD, Wang K, Wilhelm C, Xie B, Raymond D, Weld DS, Etzioni O, Kohlmeier S (2020) CORD-19: The Covid-19 Open Research Dataset. ArXiv [Preprint]
Meng L, Dong D, Li L, Niu M, Bai Y, Wang M, Qiu X, Zha Y, Tian J (2020) A deep learning prognosis model help alert for COVID-19 patients at High-risk of death: a multi-center study. IEEE J Biomed Health Inform 24:3576–3584. https://doi.org/10.1109/JBHI.2020.3034296
Zhu X, Song B, Shi F, Chen Y, Hu R, Gan J, Zhang W, Li M, Wang L, Gao Y, Shan F, Shen D (2021) Joint prediction and time estimation of COVID-19 developing severe symptoms using chest CT scan. Med Image Anal 67:101824. https://doi.org/10.1016/j.media.2020.101824
Schalekamp S, Bleeker-Rovers CP, Beenen LFM, Quarles van Ufford HME, Gietema HA, Stöger JL, Harris V, Reijers MHE, Rahamat-Langendoen J, Korevaar DA, Smits LP, Korteweg C, van Rees Vellinga TFD, Vermaat M, Stassen PM, Scheper H, Wijnakker R, Borm FJ, Dofferhoff ASM, Prokop M (2021) Chest CT in the emergency department for diagnosis of COVID-19 pneumonia: dutch experience. Radiology 298:E98–E106. https://doi.org/10.1148/radiol.2020203465
Rubin GD, Ryerson CJ, Haramati LB, Sverzellati N, Kanne JP, Raoof S, Schluger NW, Volpi A, Yim JJ, Martin IBK, Anderson DJ, Kong C, Altes T, Bush A, Desai SR, Onathan G, Goo JM, Humbert M, Inoue Y, Kauczor H-U, Luo F, Mazzone PJ, Prokop M, Remy-Jardin M, Richeldi L, Schaefer-Prokop CM, Tomiyama N, Wells AU, Leung AN (2020) The role of chest imaging in patient management during the COVID-19 pandemic: a multinational consensus statement from the Fleischner Society. Radiology 296:172–180. https://doi.org/10.1148/radiol.2020201365
Mossa-Basha M, Azadi J, Ko J, Klein J, Meltzer C (2020) RSNA COVID-19 task force: best practices for radiology departments during COVID-19. In: Radiological Society of North America, pp 1–7
Wang J, Peng C, Zhao Y, Ye R, Hong J, Huang H, Chen L (2021) Application of a robotic tele-echography system for COVID-19 pneumonia. J Ultrasound Med 40:385–390. https://doi.org/10.1002/jum.15406
Soda P, D’Amico NC, Tessadori J, Valbusa G, Guarrasi V, Bortolotto C, Akbar MU, Sicilia R, Cordelli E, Fazzini D, Cellina M, Oliva G, Callea G, Panella S, Cariati M, Cozzi D, Miele V, Stellato E, Carrafiello G, Castorani G, Simeone A, Preda L, Iannello G, Del Bue A, Tedoldi F, Alí M, Sona D, Papa S (2021) AIforCOVID: predicting the clinical outcomes in patients with COVID-19 applying AI to chest-X-rays. An Italian multicentre study. Med Image Anal 74:102216. https://doi.org/10.1016/j.media.2021.102216
Anderson W (2011) National Statement on Ethical Conduct in Human Research. Intern Med J 41:581
Born J, Beymer D, Rajan D, Coy A, Mukherjee VV, Manica M, Prasanna P, Ballah D, Guindy M, Shaham D, Shah PL, Karteris E, Robertus JL, Gabrani M, Rosen-Zvi M (2021) On the role of artificial intelligence in medical imaging of COVID-19. Patterns N Y 2:100269. https://doi.org/10.1016/j.patter.2021.100269
Lee EH, Zheng J, Colak E, Mohammadzadeh M, Houshmand G, Bevins N, Kitamura F, Altinmakas E, Reis EP, Kim J-K, Klochko C, Han M, Moradian S, Mohammadzadeh A, Sharifian H, Hashemi H, Firouznia K, Ghanaati H, Gity M, Doğan H, Salehinejad H, Alves H, Seekins J, Abdala N, Atasoy Ç, Pouraliakbar H, Maleki M, Wong SS, Yeom KW (2021) Deep COVID DeteCT: an international experience on COVID-19 lung detection and prognosis using chest CT. Npj Digit Med 4:11. https://doi.org/10.1038/s41746-020-00369-1
Wang X, Deng X, Fu Q, Zhou Q, Feng J, Ma H, Liu W, Zheng C (2020) A Weakly-supervised framework for COVID-19 classification and lesion localization from chest CT. IEEE Trans Med Imaging 39:2615–2625. https://doi.org/10.1109/TMI.2020.2995965
Tabik S, Gomez-Rios A, Martin-Rodriguez JL, Sevillano-Garcia I, Rey-Area M, Charte D, Guirado E, Suarez JL, Luengo J, Valero-Gonzalez MA, Garcia-Villanova P, Olmedo-Sanchez E, Herrera F (2020) COVIDGR dataset and COVID-SDNet methodology for predicting COVID-19 based on chest X-ray images. IEEE J Biomed Health Inform 24:3595–3605. https://doi.org/10.1109/JBHI.2020.3037127
Rajaraman S, Sornapudi S, Alderson PO, Folio LR, Antani SK (2020) Analyzing inter-reader variability affecting deep ensemble learning for COVID-19 detection in chest radiographs. PLoS ONE 15:e0242301. https://doi.org/10.1371/journal.pone.0242301
Liang S, Liu H, Gu Y, Guo X, Li H, Li L, Wu Z, Liu M, Tao L (2021) Fast automated detection of COVID-19 from medical images using convolutional neural networks. Commun Biol 4:35. https://doi.org/10.1038/s42003-020-01535-7
Pathan S, Siddalingaswamy PC, Kumar P, Pai MMM, Ali T, Acharya UR (2021) Novel ensemble of optimized CNN and dynamic selection techniques for accurate Covid-19 screening using chest CT images. Comput Biol Med 137:104835. https://doi.org/10.1016/j.compbiomed.2021.104835
Monshi MMA, Poon J, Chung V, Monshi FM (2021) CovidXrayNet: optimizing data augmentation and CNN hyperparameters for improved COVID-19 detection from CXR. Comput Biol Med 133:104375. https://doi.org/10.1016/j.compbiomed.2021.104375
Arora V, Ng EY-K, Leekha RS, Darshan M, Singh A (2021) Transfer learning-based approach for detecting COVID-19 ailment in lung CT scan. Comput Biol Med 135:104575. https://doi.org/10.1016/j.compbiomed.2021.104575
Delli Pizzi A, Chiarelli AM, Chiacchiaretta P, Valdesi C, Croce P, Mastrodicasa D, Villani M, Trebeschi S, Serafini FL, Rosa C, Cocco G, Luberti R, Conte S, Mazzamurro L, Mereu M, Patea RL, Panara V, Marinari S, Vecchiet J, Caulo M (2021) Radiomics-based machine learning differentiates “ground-glass” opacities due to COVID-19 from acute non-COVID-19 lung disease. Sci Rep 11:17237. https://doi.org/10.1038/s41598-021-96755-0
Fang C, Bai S, Chen Q, Zhou Y, Xia L, Qin L, Gong S, Xie X, Zhou C, Tu D, Zhang C, Liu X, Chen W, Bai X, Torr PHS (2021) Deep learning for predicting COVID-19 malignant progression. Med Image Anal 72:102096. https://doi.org/10.1016/j.media.2021.102096
Pu J, Leader JK, Bandos A, Ke S, Wang J, Shi J, Du P, Guo Y, Wenzel SE, Fuhrman CR, Wilson DO, Sciurba FC, Jin C (2021) Automated quantification of COVID-19 severity and progression using chest CT images. Eur Radiol 31:436–446. https://doi.org/10.1007/s00330-020-07156-2
Quan H, Xu X, Zheng T, Li Z, Zhao M, Cui X (2021) DenseCapsNet: detection of COVID-19 from X-ray images using a capsule neural network. Comput Biol Med 133:104399. https://doi.org/10.1016/j.compbiomed.2021.104399
Mishra NK, Singh P, Joshi SD (2021) Automated detection of COVID-19 from CT scan using convolutional neural network. Biocybern Biomed Eng 41:572–588. https://doi.org/10.1016/j.bbe.2021.04.006
Paluru N, Dayal A, Jenssen HB, Sakinis T, Cenkeramaddi LR, Prakash J, Yalavarthy PK (2021) Anam-Net: anamorphic depth embedding-based lightweight cnn for segmentation of anomalies in COVID-19 chest CT images. IEEE Trans Neural Netw Learn Syst 32:932–946. https://doi.org/10.1109/TNNLS.2021.3054746
Goncharov M, Pisov M, Shevtsov A, Shirokikh B, Kurmukov A, Blokhin I, Chernina V, Solovev A, Gombolevskiy V, Morozov S, Belyaev M (2021) CT-based COVID-19 triage: Deep multitask learning improves joint identification and severity quantification. Med Image Anal 71:102054. https://doi.org/10.1016/j.media.2021.102054
Chassagnon G, Vakalopoulou M, Battistella E, Christodoulidis S, Hoang-Thi T-N, Dangeard S, Deutsch E, Andre F, Guillo E, Halm N, El Hajj S, Bompard F, Neveu S, Hani C, Saab I, Campredon A, Koulakian H, Bennani S, Freche G, Barat M, Lombard A, Fournier L, Monnier H, Grand T, Gregory J, Nguyen Y, Khalil A, Mahdjoub E, Brillet P-Y, Tran Ba S, Bousson V, Mekki A, Carlier R-Y, Revel M-P, Paragios N (2021) AI-driven quantification, staging and outcome prediction of COVID-19 pneumonia. Med Image Anal 67:101860. https://doi.org/10.1016/j.media.2020.101860
Zhang Z, Chen B, Sun J, Luo Y (2021) A bagging dynamic deep learning network for diagnosing COVID-19. Sci Rep 11:16280. https://doi.org/10.1038/s41598-021-95537-y
Fung DLX, Liu Q, Zammit J, Leung CK-S, Hu P (2021) Self-supervised deep learning model for COVID-19 lung CT image segmentation highlighting putative causal relationship among age, underlying disease and COVID-19. J Transl Med 19:318. https://doi.org/10.1186/s12967-021-02992-2
Wang D, Huang C, Bao S, Fan T, Sun Z, Wang Y, Jiang H, Wang S (2021) Study on the prognosis predictive model of COVID-19 patients based on CT radiomics. Sci Rep 11:11591. https://doi.org/10.1038/s41598-021-90991-0
Wu Z, Li L, Jin R, Liang L, Hu Z, Tao L, Han Y, Feng W, Zhou D, Li W, Lu Q, Liu W, Fang L, Huang J, Gu Y, Li H, Guo X (2021) Texture feature-based machine learning classifier could assist in the diagnosis of COVID-19. Eur J Radiol 137:109602. https://doi.org/10.1016/j.ejrad.2021.109602
Li T, Wei W, Cheng L, Zhao S, Xu C, Zhang X, Zeng Y, Gu J (2021) Computer-aided diagnosis of COVID-19 CT scans based on spatiotemporal information fusion. J Healthc Eng 2021:1–11. https://doi.org/10.1155/2021/6649591
Jin C, Chen W, Cao Y, Xu Z, Tan Z, Zhang X, Deng L, Zheng C, Zhou J, Shi H, Feng J (2020) Development and evaluation of an artificial intelligence system for COVID-19 diagnosis. Nat Commun 11:5088. https://doi.org/10.1038/s41467-020-18685-1
Yao Q, Xiao L, Liu P, Zhou SK (2021) Label-free segmentation of COVID-19 lesions in lung CT. IEEE Trans Med Imaging 40:2808–2819. https://doi.org/10.1109/TMI.2021.3066161
Cardobi N, Benetti G, Cardano G, Arena C, Micheletto C, Cavedon C, Montemezzi S (2021) CT radiomic models to distinguish COVID-19 pneumonia from other interstitial pneumonias. Radiol Med (Torino) 126:1037–1043. https://doi.org/10.1007/s11547-021-01370-8
Muhammad G, Shamim Hossain M (2021) COVID-19 and non-COVID-19 classification using multi-layers fusion from lung ultrasound images. Inf Fusion 72:80–88. https://doi.org/10.1016/j.inffus.2021.02.013
Wang S, Dong D, Li L, Li H, Bai Y, Hu Y, Huang Y, Yu X, Liu S, Qiu X, Lu L, Wang M, Zha Y, Tian J (2021) A deep learning radiomics model to identify poor outcome in COVID-19 patients with underlying health conditions: a multicenter study. IEEE J Biomed Health Inform 25:2353–2362. https://doi.org/10.1109/JBHI.2021.3076086
Pan F, Li L, Liu B, Ye T, Li L, Liu D, Ding Z, Chen G, Liang B, Yang L, Zheng C (2021) A novel deep learning-based quantification of serial chest computed tomography in Coronavirus Disease 2019 (COVID-19). Sci Rep 11:417. https://doi.org/10.1038/s41598-020-80261-w
Zhao X, Zhang P, Song F, Fan G, Sun Y, Wang Y, Tian Z, Zhang L, Zhang G (2021) D2A U-Net: automatic segmentation of COVID-19 CT slices based on dual attention and hybrid dilated convolution. Comput Biol Med 135:104526. https://doi.org/10.1016/j.compbiomed.2021.104526
Rahimzadeh M, Attar A, Sakhaei SM (2021) A fully automated deep learning-based network for detecting COVID-19 from a new and large lung CT scan dataset. Biomed Signal Process Control 68:102588. https://doi.org/10.1016/j.bspc.2021.102588
Zhang B, Ni-jia-Ti M, Yan R, An N, Chen L, Liu S, Chen L, Chen Q, Li M, Chen Z, You J, Dong Y, Xiong Z, Zhang S (2021) CT-based radiomics for predicting the rapid progression of coronavirus disease 2019 (COVID-19) pneumonia lesions. Br J Radiol 94:20201007. https://doi.org/10.1259/bjr.20201007
Gao K, Su J, Jiang Z, Zeng L-L, Feng Z, Shen H, Rong P, Xu X, Qin J, Yang Y, Wang W, Hu D (2021) Dual-branch combination network (DCN): towards accurate diagnosis and lesion segmentation of COVID-19 using CT images. Med Image Anal 67:101836. https://doi.org/10.1016/j.media.2020.101836
Heidari M, Mirniaharikandehei S, Khuzani AZ, Danala G, Qiu Y, Zheng B (2020) Improving the performance of CNN to predict the likelihood of COVID-19 using chest X-ray images with preprocessing algorithms. Int J Med Inf 144:104284. https://doi.org/10.1016/j.ijmedinf.2020.104284
Zhang K, Liu X, Shen J, Li Z, Sang Y, Wu X, Zha Y, Liang W, Wang C, Wang K, Ye L, Gao M, Zhou Z, Li L, Wang J, Yang Z, Cai H, Xu J, Yang L, Cai W, Xu W, Wu S, Zhang W, Jiang S, Zheng L, Zhang X, Wang L, Lu L, Li J, Yin H, Wang W, Li O, Zhang C, Liang L, Wu T, Deng R, Wei K, Zhou Y, Chen T, Lau JY-N, Fok M, He J, Lin T, Li W, Wang G (2020) Clinically applicable AI system for accurate diagnosis, quantitative measurements, and prognosis of COVID-19 pneumonia using computed tomography. Cell 181:1423–1433. https://doi.org/10.1016/j.cell.2020.04.045
Awasthi N, Dayal A, Cenkeramaddi LR, Yalavarthy PK (2021) Mini-COVIDNet: efficient lightweight deep neural network for ultrasound based point-of-care detection of COVID-19. IEEE Trans Ultrason Ferroelectr Freq Control 68:2023–2037. https://doi.org/10.1109/TUFFC.2021.3068190
Dastider AG, Sadik F, Fattah SA (2021) An integrated autoencoder-based hybrid CNN-LSTM model for COVID-19 severity prediction from lung ultrasound. Comput Biol Med 132:104296. https://doi.org/10.1016/j.compbiomed.2021.104296
Li L, Qin L, Xu Z, Yin Y, Wang X, Kong B, Bai J, Lu Y, Fang Z, Song Q, Cao K, Liu D, Wang G, Xu Q, Fang X, Zhang S, Xia J, Xia J (2020) Using artificial intelligence to detect COVID-19 and community-acquired pneumonia based on pulmonary CT: evaluation of the diagnostic accuracy. Radiology 296:E65–E71. https://doi.org/10.1148/radiol.2020200905
Wang D, Mo J, Zhou G, Xu L, Liu Y (2020) An efficient mixture of deep and machine learning models for COVID-19 diagnosis in chest X-ray images. PLoS ONE 15:e0242535. https://doi.org/10.1371/journal.pone.0242535
Yan T, Wong PK, Ren H, Wang H, Wang J, Li Y (2020) Automatic distinction between COVID-19 and common pneumonia using multi-scale convolutional neural network on chest CT scans. Chaos Solitons Fractals 140:110153. https://doi.org/10.1016/j.chaos.2020.110153
Wang Z, Liu Q, Dou Q (2020) Contrastive cross-site learning with redesigned net for COVID-19 CT classification. IEEE J Biomed Health Inform 24:2806–2813. https://doi.org/10.1109/JBHI.2020.3023246
Yang D, Xu Z, Li W, Myronenko A, Roth HR, Harmon S, Xu S, Turkbey B, Turkbey E, Wang X, Zhu W, Carrafiello G, Patella F, Cariati M, Obinata H, Mori H, Tamura K, An P, Wood BJ, Xu D (2021) Federated semi-supervised learning for COVID region segmentation in chest CT using multi-national data from China, Italy, Japan. Med Image Anal 70:101992. https://doi.org/10.1016/j.media.2021.101992
Xue W, Cao C, Liu J, Duan Y, Cao H, Wang J, Tao X, Chen Z, Wu M, Zhang J, Sun H, Jin Y, Yang X, Huang R, Xiang F, Song Y, You M, Zhang W, Jiang L, Zhang Z, Kong S, Tian Y, Zhang L, Ni D, Xie M (2021) Modality alignment contrastive learning for severity assessment of COVID-19 from lung ultrasound and clinical information. Med Image Anal 69:101975. https://doi.org/10.1016/j.media.2021.101975
Shaban WM, Rabie AH, Saleh AI, Abo-Elsoud MA (2020) A new COVID-19 patients detection strategy (CPDS) based on hybrid feature selection and enhanced KNN classifier. Knowl Based Syst 205:106270. https://doi.org/10.1016/j.knosys.2020.106270
Bai HX, Wang R, Xiong Z, Hsieh B, Chang K, Halsey K, Tran TML, Choi JW, Wang D-C, Shi L-B, Mei J, Jiang X-L, Pan I, Zeng Q-H, Hu P-F, Li Y-H, Fu F-X, Huang RY, Sebro R, Yu Q-Z, Atalay MK, Liao W-H (2020) Artificial intelligence augmentation of radiologist performance in distinguishing COVID-19 from pneumonia of other origin at chest CT. Radiology 296:E156–E165. https://doi.org/10.1148/radiol.2020201491
Arntfield R, VanBerlo B, Alaifan T, Phelps N, White M, Chaudhary R, Ho J, Wu D (2021) Development of a convolutional neural network to differentiate among the etiology of similar appearing pathological B lines on lung ultrasound: a deep learning study. BMJ Open 11:e045120. https://doi.org/10.1136/bmjopen-2020-045120
Wang J, Bao Y, Wen Y, Lu H, Luo H, Xiang Y, Li X, Liu C, Qian D (2020) Prior-attention residual learning for more discriminative COVID-19 screening in CT images. IEEE Trans Med Imaging 39:2572–2583. https://doi.org/10.1109/TMI.2020.2994908
Arias-Londono JD, Gomez-Garcia JA, Moro-Velazquez L, Godino-Llorente JI (2020) Artificial intelligence applied to chest X-ray images for the automatic detection of COVID-19. A Thoughtful Evaluation Approach. IEEE Access 8:226811–226827. https://doi.org/10.1109/ACCESS.2020.3044858
Chaganti S, Grenier P, Balachandran A, Chabin G, Cohen S, Flohr T, Georgescu B, Grbic S, Liu S, Mellot F, Murray N, Nicolaou S, Parker W, Re T, Sanelli P, Sauter AW, Xu Z, Yoo Y, Ziebandt V, Comaniciu D (2020) Automated quantification of CT patterns associated with COVID-19 from chest CT. Radiol Artif Intell 2:e200048. https://doi.org/10.1148/ryai.2020200048
Wang S, Kang B, Ma J, Zeng X, Xiao M, Guo J, Cai M, Yang J, Li Y, Meng X, Xu B (2021) A deep learning algorithm using CT images to screen for Corona virus disease (COVID-19). Eur Radiol 31:6096–6104. https://doi.org/10.1007/s00330-021-07715-1
Yan Q, Wang B, Gong D, Luo C, Zhao W, Shen J, Ai J, Shi Q, Zhang Y, Jin S, Zhang L, You Z (2021) COVID-19 chest CT image segmentation network by multi-scale fusion and enhancement operations. IEEE Trans Big Data 7:13–24. https://doi.org/10.1109/TBDATA.2021.3056564
Mortani Barbosa EJ, Gefter WB, Ghesu FC, Liu S, Mailhe B, Mansoor A, Grbic S, Vogt S (2021) Automated detection and quantification of COVID-19 airspace disease on chest radiographs: a novel approach achieving expert radiologist-level performance using a deep convolutional neural network trained on digital reconstructed radiographs from computed tomography-derived ground truth. Investig Radiol 56:471–479. https://doi.org/10.1097/RLI.0000000000000763
Tan H-B, Xiong F, Jiang Y-L, Huang W-C, Wang Y, Li H-H, You T, Fu T-T, Lu R, Peng B-W (2020) The study of automatic machine learning base on radiomics of non-focus area in the first chest CT of different clinical types of COVID-19 pneumonia. Sci Rep 10:18926. https://doi.org/10.1038/s41598-020-76141-y
Sharifrazi D, Alizadehsani R, Roshanzamir M, Joloudari JH, Shoeibi A, Jafari M, Hussain S, Sani ZA, Hasanzadeh F, Khozeimeh F, Khosravi A, Nahavandi S, Panahiazar M, Zare A, Islam SMS, Acharya UR (2021) Fusion of convolution neural network, support vector machine and Sobel filter for accurate detection of COVID-19 patients using X-ray images. Biomed Signal Process Control 68:102622. https://doi.org/10.1016/j.bspc.2021.102622
Vidal PL, de Moura J, Novo J, Ortega M (2021) Multi-stage transfer learning for lung segmentation using portable X-ray devices for patients with COVID-19. Expert Syst Appl 173:114677. https://doi.org/10.1016/j.eswa.2021.114677
Born J, Wiedemann N, Cossio M, Buhre C, Brändle G, Leidermann K, Aujayeb A, Moor M, Rieck B, Borgwardt K (2021) Accelerating detection of lung pathologies with explainable ultrasound image analysis. Appl Sci 11:672. https://doi.org/10.3390/app11020672
Duran-Lopez L, Dominguez-Morales JP, Corral-Jaime J, Vicente-Diaz S, Linares-Barranco A (2020) COVID-XNet: a custom deep learning system to diagnose and locate COVID-19 in chest x-ray images. Appl Sci 10:5683. https://doi.org/10.3390/app10165683
Song Y, Zheng S, Li L, Zhang X, Zhang X, Huang Z, Chen J, Wang R, Zhao H, Chong Y, Shen J, Zha Y, Yang Y (2021) Deep learning enables accurate diagnosis of novel coronavirus (COVID-19) with CT images. IEEE/ACM Trans Comput Biol Bioinform 18:2775–2780. https://doi.org/10.1109/TCBB.2021.3065361
Li C, Dong D, Li L, Gong W, Li X, Bai Y, Wang M, Hu Z, Zha Y, Tian J (2020) Classification of severe and critical Covid-19 using deep learning and radiomics. IEEE J Biomed Health Inform 24:3585–3594. https://doi.org/10.1109/JBHI.2020.3036722
Wang L, Lin ZQ, Wong A (2020) COVID-Net: a tailored deep convolutional neural network design for detection of COVID-19 cases from chest X-ray images. Sci Rep 10:19549. https://doi.org/10.1038/s41598-020-76550-z
Qian X, Fu H, Shi W, Chen T, Fu Y, Shan F, Xue X (2020) M3Lung-Sys: a deep learning system for multi-class lung pneumonia screening from CT imaging. IEEE J Biomed Health Inform 24:3539–3550. https://doi.org/10.1109/JBHI.2020.3030853
Li Y, Wei D, Chen J, Cao S, Zhou H, Zhu Y, Wu J, Lan L, Sun W, Qian T, Ma K, Xu H, Zheng Y (2020) Efficient and effective training of COVID-19 classification networks with self-supervised dual-track learning to rank. IEEE J Biomed Health Inform 24:2787–2797. https://doi.org/10.1109/JBHI.2020.3018181
Ouyang X, Huo J, Xia L, Shan F, Liu J, Mo Z, Yan F, Ding Z, Yang Q, Song B, Shi F, Yuan H, Wei Y, Cao X, Gao Y, Wu D, Wang Q, Shen D (2020) Dual-sampling attention network for diagnosis of COVID-19 from community acquired pneumonia. IEEE Trans Med Imaging 39:2595–2605. https://doi.org/10.1109/TMI.2020.2995508
Zhou L, Li Z, Zhou J, Li H, Chen Y, Huang Y, Xie D, Zhao L, Fan M, Hashmi S, Abdelkareem F, Eiada R, Xiao X, Li L, Qiu Z, Gao X (2020) A rapid, accurate and machine-agnostic segmentation and quantification method for CT-based COVID-19 diagnosis. IEEE Trans Med Imaging 39:2638–2652. https://doi.org/10.1109/TMI.2020.3001810
Roy S, Menapace W, Oei S, Luijten B, Fini E, Saltori C, Huijben I, Chennakeshava N, Mento F, Sentelli A, Peschiera E, Trevisan R, Maschietto G, Torri E, Inchingolo R, Smargiassi A, Soldati G, Rota P, Passerini A, van Sloun RJG, Ricci E, Demi L (2020) Deep learning for classification and localization of COVID-19 markers in point-of-care lung ultrasound. IEEE Trans Med Imaging 39:2676–2687. https://doi.org/10.1109/TMI.2020.2994459
Qayyum A, Razzak I, Tanveer M, Kumar A (2021) Depth-wise dense neural network for automatic COVID19 infection detection and diagnosis. Ann Oper Res. https://doi.org/10.1007/s10479-021-04154-5
Liu J, Dong B, Wang S, Cui H, Fan D-P, Ma J, Chen G (2021) COVID-19 lung infection segmentation with a novel two-stage cross-domain transfer learning framework. Med Image Anal 74:102205. https://doi.org/10.1016/j.media.2021.102205
Yang Y, Zhang L, Du M, Bo J, Liu H, Ren L, Li X, Deen MJ (2021) A comparative analysis of eleven neural networks architectures for small datasets of lung images of COVID-19 patients toward improved clinical decisions. Comput Biol Med 139:104887. https://doi.org/10.1016/j.compbiomed.2021.104887
Funding
AG, JD, GB, AN, HM and AT performed this research as a part of employment at the Commonwealth Scientific and Industrial Research Organisation (CSIRO), Australia. FL is supported by an Australian Government Research Training Program scholarship and a CSIRO Top Up Scholarship.
Author information
Authors and Affiliations
Contributions
AG conceived the idea, reviewed studies, performed methodology analysis and wrote the manuscript. FL, JP, GB, AN, GM and JD reviewed studies and reviewed the manuscript. AT performed author analysis and reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gillman, A.G., Lunardo, F., Prinable, J. et al. Automated COVID-19 diagnosis and prognosis with medical imaging and who is publishing: a systematic review. Phys Eng Sci Med 45, 13–29 (2022). https://doi.org/10.1007/s13246-021-01093-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13246-021-01093-0