Abstract
Root-associated fungi, with the focus on endophytic species, were isolated from healthy Arabidopsis thaliana and Microthlaspi perfoliatum plants collected at different locations in Germany. A large number of fungal taxa were discovered with a small-scale approach. This provides additional evidence that root-associated and endophytic fungi are common in Brassicaceae. The most prevalent genera associated with A. thaliana roots were Trichoderma and Fusarium, while the roots of M. perfoliatum were dominated by different species of Fusarium and Penicillium. Differences in species composition and richness might be due to preferences and life-cycle of the two plant species. Strains of endophyte species that did not have closely related species in GenBank searches and those already known as root endophytes were chosen for preliminary co-cultivation experiments using germinating host plants on agar medium to observe effects on plant growth and health. Under these conditions several fungal isolates had an adverse effect on plant growth and health, especially on Arabidopsis thaliana. Some isolates did not adversely affect biomass during initial plant growth, while they altered the shoot-root ratio in favour of the shoot, especially in Microthlaspi perfoliatum. These strains are promising candidates for future research on endophytes as they might have some effects in Brassicaceae that are similar to mycorrhizal fungi. They are also promising candidates for investigating interactions with their host plants.




Similar content being viewed by others
References
Al-Shebaz A, Beilstein MA, Kellogg EA (2006) Systematics and phylogeny of the Brassicaceae (cruciferae): an overview. Pl Syst Evol 259:89–120
Aly AH, Debbab A, Proksch P (2011) Fungal endophytes: unique plant inhabitants with great promises. Appl Microbiol Biotechnol 90:1829–1845
Barrow JR, Osuna P (2002) Phosphorus solubilization and uptake by dark septate fungi in fourwing saltbush, Atriplexcanescens (pursh) Nutt. J Arid Environ 51:449–459
Berg G, Smalla K (2009) Plant species and soil type cooperatively shape the structure and function of microbial communities in the rhizosphere. FEMS Micobiol Ecol 68:1–13
Bressan M, Roncato M, Bellvert F, Comte G, Haichar F, Achouak W, Berge O (2009) Exogenous glucosinolate produced by Arabidopsis thaliana has an impact on microbes in the rhizosphere and plant roots. ISME J 3:1243–1257
Brundrett MC (2002) Coevolution of roots and mycorrhizas of land plants. New Phytol 154:275–304
Buell CR, Last RL (2010) Twenty-first century plant biology: impacts of the Arabidopsis genome on plant biology and agriculture. Pl Physiol 154:497–500
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL (2009) BLAST+: architecture and applications. BMC Bioinforma 10:421
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17:540–552
Chaparro JM, Sheflin AM, Manter DK, Vivanco JM (2012) Manipulating the soil microbiome to increase soil health and plant fertility. BiolFert Soils 48:489–499
Delaye L, García-Guzmán G, Heil M (2013) Endophytes versus biotrophic and necrotrophic pathogens—are fungal lifestyles evolutionarily stable traits? Fungal Divers 60:125–135
Diagne N, Escoute J, Lartaud M, Verdeil JL, Franche C, Kane A, Bogusz D, Diouf, Duponnois R, Svistoonoff (2011) Uvitex B: a rapid and efficient stain for detection of arbuscular mycorrhizal fungi within plant roots. Mycorrhiza 21:315–321
Diaz C, Saliba-Colombani V, Loudet O, Belluomo P, Moreau L, Daniel-Vedele F, Morot-Gaudry J, Masclaux-Daubresse C (2006) Leaf Yeallowing and Anthocyanin Accumulation are Two Genetically Independent Strategies in Response to Nitrogen Limitation in Arabidopsis thaliana. Pl Cell Physiol 47:74–83
Ernst M, Mendgen KW, Wirsel SGR (2003) Endophytic fungal mutualists: seed-borne stagonospora spp. Enhance reed biomass production in axenic microcosms. MPMI 16:580–587
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
García E, Alonso A, Plates G, Sacristán S (2013) The endophytic mycobiota of Arabidopsis thaliana. Fungal Divers 60:71–89
Hamayun M, Khan SA, Khan AL, Tang D, Hussain J, Ahmad B, Anwar Y, Lee I (2010a) Growth promotion of cucumber by pure cultures of gibberellins-producing Phoma sp. GAH 7. World J Microbiol Biotechnol 26:889–894
Hamayun M, Khan SA, Khan AL, Rehman G, Kim Y, Iqbal I, Hussain J, Sohn E, Lee I (2010b) Gibberellin producing and plant growth promotion from pure cultures of Cladosporium sp. MH-6 isolated from cucumber (Cucumis sativus L.). Mycologia 102:989–995
Hamilton CE, Bauerle TL (2012) A new currency for mutualism? Fungal endophytes alter antioxidant activity in hosts responding to drought. Fungal Divers 54:3949
Hamilton CE, Gundel PE, Helander M, Saikkonen K (2012) Endophytic mediation of reactive oxygen species and antioxidant activity in plants: a review. Fungal Divers 54:110
Harman GE, Howell CR, Viterbo A, Chet I, Lorito M (2004) Trichoderma species – Opportunistic, avirulent plant symbionts. Nat Rev Microbiol 2:43–57
Hashiba T, Narisawa K (2005) The development and endophytic nature of the fungus Heteroconiumchaetospira. FEMS Microbiol Lett 252:191–196
Hyde KD, Soytong K (2008) The fungal endophyte dilemma. Fungal Divers 33:163–173
Ihaka R, Gentleman R (1996) R: A language for data analysis and graphics. J Comput Graph Stat 5:299–314
Ishimoto H, Fukushi Y, Yoshida T, Tahara S (2000) Rhizopus and Fusarium are selected as dominant fungal genera in rhizospheres of Brassicaceae. J Chem Ecol 26:2387–2399
Jumponnen A (2001) Dark septate endophytes-are they mycorrhizal? Mycorrhiza 11:207–211
Jumponnen A, Trappe JM (1998) Dark septate endophytes: a review of facultative biotrophic root-colonizing fungi. New Phytol 140:295–310
Junker C, Draeger S, Schulz B (2012) A fine line-endophytes or pathogens in Arabidopsis thaliana. Fungal Ecol 5:657–663
Kageyama SA, Madyam KG, Jumpponen A (2008) Diversity, Function and Potential Applications of the Root-Associated Endophytes. In: Varma A (Ed.), Mycorrhiza – State of the Art, Genetics and Molecular Biology, Eco-Phyiology, Structure and Systematics (pp. 29–57). Berlin and Heidelberg, Springer
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780
Katoh K, Toh H (2008) Recent developments in the MAFFT multiple sequence alignment program. Briefings Bioinf 9:286–298
Katoh K, Misawa K, Kuma K, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30:3059–3066
Keawchai S, Soytong K, Hyde KD (2009) Mycofungicides and fungal biofertilizers. Fungal Divers 38:25–50
Ko KTW, Stephenson SL, Bahkali AH, Hyde KD (2011) From morphology to molecular biology: can we use sequence data to identify fungal endophytes? Fungal Divers 50:113–120
Maciá-Vicente JG, Jansson H, Abdullah SK, Descals E, Salinas J, Lopez-Llorca LV (2008) Fungal root endophytes from natural vegetation in Mediterranean environments with special reference to Fusarium spp. FEMS Microbiol Ecol 64:90–105
Mandyam KG, Roe J, Jumpponen A (2013) Arabidopsis thaliana model system reveals a continuum of responses to root endophyte colonization. Fungal Biol 117:250–260
May KJ, Ristaino JB (2004) Identity of the mtDNA halplotype(s) of Phytophtora infestans in historical specimens from the Irish Potato Famine. Mycol Res 108:471–479
Mitchell-Olds T (2001) Arabidopsis thaliana and its wild relatives: a model system for ecology and evolution. Trends Ecol Evol 16:693–700
Newton AC, Fitt BDL, Atkins SD, Walters DR, Daniell TJ (2010) Pathogenesis, parasitism and mutualism in the trophic space of microbe-plant interactions. Trends Microbiol 18:365–373
Oberwinkler F, Riess K, Bauer R, Selosse MA, Weiß M, Garnica S, Zuccaro A (2013) Enigmatic Sebacinales. Mycol Prog 12:1–27
Ott M, Zola J, Aluru S, Stamatakis A (2007) Large-scale maximum likelihood-based phylogenetic analysis on the IBM BlueGene/L. In: Proceeding SC ’07—Proceedings of the 2007 ACM/IEEE Conference on Supercomputing
Paparu P, Macleod A, Dubois T, Coyne D, Viljoen A (2009) Efficacy of chemical and fluorescent protein markers studying plant colonization by endophytic non-pathogenic Fusarium oxysporum isolates. BioControl 54:709–722
Peršoh D (2013) Factors shaping community structure of endophytic fungi–evidence from the Pinus-Viscum-system. Fungal Divers 60:55–69
Peškan-Berghöfer T, Shahollari B, Giong PH, Hehl S, Merkert C, Blanke V, Kost G, Varma A, Oelmüller R (2004) Associations of Piriformospora indica with Arabidopsis thaliana represents a novel system to study beneficial plant-microbe interactions and involves early plant protein modifications in the endoplasmatic reticulum and at the plasma membrane. Physiol Plant 122:465–477
Pham GH, Kumari R, Singh A, Malla R, Prasad R, Sachdev M, Kaldorf M, Buscot F, Oelmüller R, Hampp R, Saxena AK, Rexer KH, Kost G, Varma A (2004) Axenic culture of symbiotic fungus Piriformospora indica. In: Varma A, Abbot L, Werner D, Hampp R (eds) Plant surface microbiology. Springer, Berlin, pp 593–612
Porras-Alfaro A, Bayman P (2011) Hidden fungi, emergent properties: endophytes and microbiomes. Phytopathol 49:291–315
Price MN, Dehal PS, Arkin AP (2010) FastTree 2–approximately maximum-likelihood trees for large alignments. PLoS One 5:e9490
Regvar M, Vogel K, Irgel N, Wraber T, Hildebrandt U, Wilde P, Bothe H (2003) Colonization of pennycresses (Thlaspi spp.) of the Brassicaceae by arbuscular mycorrhizal fungi. JPl Physiol 160:615–626
Rich TCC, Lambrick CR, Kitchen C, Kitchen MAR (1998) Conserving Britain’s biodiversity. I: Thlaspi perfoliatum L. (Brassiaceae), cotswold pennycress. Biodiv Cons 7:915–926
Rodriguez JR, Redman RS, Henson JM (2004) The role of fungal symbiosis in the adaptation of plants to high stress environments. Mitig Adapt Strateg Glob Chang 9:261–272
Rodriguez RJ, Henson J, Van Volkenburgh E, Hoy M, Wright L, Beckwith F, Kim Y, Redman RS (2008) Stress tolerance in plants via habitat-adapted symbiosis. ISME J 2:404–416
Rodriguez RJ, White JF, Arnold AE, Redman RS (2009) Fungal endophytes: diversity and functional roles. New Phytol 182:314–330
Ronquist F, Teslenko M, Pvd M, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542
Scervino JM, Mesa MP, Mónica ID, Reechi M, Moreno NS, Goderas A (2010) Soil fungal isolates produce different organic acid patterns involved in phosphate salts solubilization. Biol Fert Soils 46:755–763
Schreiner RP, Koide RT (1993) Mustards, mustard oils and mycorrhizas. New Phytol 123:107–113
Schulz B, Boyle C (2005) The endophytic continuum. Mycol Res 109:661–686
Schulz B, Wanke U, Draeger S, Aust HJ (1993) Endophytes from herbaceous plants and shrubs: effectiveness of surface sterilization methods. Mycol Res 97:1447–1450
Schulz B, Sucker J, Aust HJ, Krohn K, Ludewig K, Jones PG, Döring D (1995) Biologically active secondary metabolites of endophytic Pezicula species. Mycol Res 99:1007–1015
Selosse MA, Dubois MP, Alvarez N (2009) Do Sebacinales commonly associate with plant roots as endophytes? Mycol Res 113:1062–1069
Sherameti I, Shahollari B, Venus Y, Altschmied L, Varma A,Oelmüller R (2005) The endophytic fungus Piriformospora indica stimulates the expression of nitrate reductase and the starch-degrading enzyme glucan-water dikinase in tobacco and Arabidopsis roots through a homeodomain transcription factor that binds to a conserved motif in their promotors. J Biol Chem 280:26241–26247
Sirrenberg A, Göbel C, Grond S, Czempinski N, Ratzinger A, Karlovsky P, Santos P, Feussner I, Pawlowski (2007) Piriformospora indica affects plant growth by auxin production. Physiologia Plantarum 131:581–589
Stamatakis A (2006) RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22:2688–2690
Stein E, Molitor A, Kogel K, Waller F (2008) Systemic resistance in Arabidopsis conferred by the mycorrhizal fungus Piriformospora indica requires jasmonic acid signaling and the cytoplasmic function of NPR1. Pl Cell Physiol 49:1747–1751
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: Molecular Evolutionary Genetics Analysis Using Maximum Likelihood, Evolutionary Distance, and Maximum Parsimony Methods. Molecular Biology and Evolution 28:2731–2739
Väre H, Vestberg M, Eurola S (1992) Mycorrhiza and root-associated fungi in Spitsbergen. Mycorrhiza 1:93–104
Varma A, Bakshi M, Lou B, Hartmann A, Oelmüller R (2012) Piriformospora indica: a novel plant growth-promoting mycorrhizal fungus. Agric Res 1:117–131
Wada KC, Takeno K (2010) Stress-induced flowering. Pl Signal Behavior 5:944–947
Wang B, Qiu YL (2006) Phylogentic distribution and evolution of mycorrhizas in land plants. Mycorrhiza 16:299–363
Warwick SI, Mummenhoff K, Sauder CA, Koch MA, Al-Shebaz IA (2010) Closing the gaps: phylogenetic relationships in the Brassicaceae based on DNA sequence data of nuclear ribosomal ITS region. Pl Syst Evol 285:209–232
Weiß M, Sýkorova Z, Garnica S, Riess K, Martos F, Krause C, Oberwinkler F, Bauer R, Redecker D (2011) Sebacinales everywhere: previously overlooked ubiquitous fungal endophytes. PLoS ONE 6:1–7
White TJ, Bruns TD, Lee SB, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ and White TJ (Eds.), PCR Protocols: a Guide to Methods and Applications (pp. 315–322). New York, Academic Press
Acknowledgments
This study has been funded by the LOEWE program of the government of Hesse in the framework of the Integrative Fungal Research Cluster (IPF) and the Biodiversity and Climate Research Centre (BiK-F). We are grateful for the constructive criticism of the reviewers, which helped to improve this manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
13225_2014_289_MOESM1_ESM.png
Figures S1–S14. Phylogenetic trees for the different fungal orders and Pythium inferred using Maximum Likelihood. Strains from Microthlaspi perfoliatum are highlighted in green, those from Arabidopsis thaliana are highlighted in yellow. Numbers at branches denote bootstrap support from 1000 bootstrap replicates. Fig. S1 Capnopodiales, Fig. S2. Diaportales, Fig. S3. Eurotiales, Fig. S4. Glomerellales, Fig. S5. Helotiales, Fig. S6. Hypocreales, Fig. S7. Microascales, Fig. S8. Mortierellales, Fig. S9. Mucorales, Fig. S10. Phyllochorales, Fig. S11. Pleosporales, Fig. S12. Pythium, Fig. S13. Sordariales, Fig. S14. Xylariales. (TRE 0 kb) (PNG 542 kb)
Figure S15
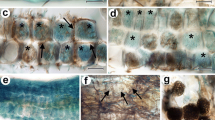
Microthlaspi perfoliatum roots colonized by different fungal species. a Primary rootinfected with Pyrenochaeta sp. with intracellular hyphae (arrows). Overlay of bright field-image and fluorescence image. Scale bar: 50 μm. b Loose hyphal network around a primary root infected by Colletotrichum aff. destructivum. Fluorescence image. Scale bar: 100 μm. c Dense network of subepidermal and intercellular hyphae (arrows) of Pyrenochaeta sp.near the root apex. Fluorescence image. Scale bar: 50 μm. d Hyphal mantle and sporulation (arrows) of Ilyonectria radicicola agg.on the surface of a primary root. Fluorescence image. Scale bar: 100 μm. e Bright-field image of D. Scale bar: 100 μm. f Superficial sporulation by Hypocreales sp. 1 near the root apex. Fluorescence image. Scale bar: 5 μm. (JPEG 929 kb)
Figure S16
Fluorescence microscopy images of Arabidopsis thaliana roots colonized by different fungal species. a Primary root with superficial hyphae and sporulation (arrows) by Cladosporium cladosporioides aggregate. Scale bar: 50 μm. b Spores of Fusarium aff. tricinctum on root hair (arrow). Scale bar: 10 μm. c Subepidermal hyphal structures of Microdochium bolleyi in primary root (arrow). Scale bar: 20 μm. d Superficial hyphae of Microdochium bolleyi on primary root (arrows). Scale bar: 50 μm. (JPEG 442 kb)
ESM 16
(TRE 1 kb)
ESM 17
(TRE 0 kb)
ESM 18
(TRE 0 kb)
ESM 19
(TRE 3 kb)
ESM 20
(TRE 4 kb)
ESM 21
(TRE 0 kb)
ESM 22
(TRE 0 kb)
ESM 23
(TRE 0 kb)
ESM 24
(TRE 0 kb)
ESM 25
(TRE 6 kb)
ESM 26
(TRE 7 kb)
ESM 27
(TRE 0 kb)
ESM 28
(TRE 0 kb)
ESM 29
(TRE 2 kb)
ESM 30
(TRE 3 kb)
ESM 31
(TRE 1 kb)
ESM 32
(TRE 1 kb)
ESM 33
(TRE 0 kb)
ESM 34
(TRE 0 kb)
ESM 35
(TRE 2 kb)
ESM 36
(TRE 2 kb)
ESM 37
(TRE 0 kb)
ESM 38
(TRE 0 kb)
ESM 39
(TRE 0 kb)
ESM 40
(TRE 0 kb)
ESM 41
(TRE 0 kb)
ESM 42
(TRE 0 kb)
ESM 43
(DOCX 16 kb)
Rights and permissions
About this article
Cite this article
Keim, J., Mishra, B., Sharma, R. et al. Root-associated fungi of Arabidopsis thaliana and Microthlaspi perfoliatum . Fungal Diversity 66, 99–111 (2014). https://doi.org/10.1007/s13225-014-0289-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13225-014-0289-2