Abstract
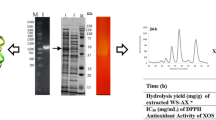
Advanced techniques of enzyme production and purification have become prerequisite due to their diverse industrial applications. There is an utmost requirement for screening of new strains capable of synthesising industrially useful enzymes. The present study reports the production and profiling of extracellular proteins expressed by the newly isolated strain of a filamentous fungus, Aspergillus oryzae LC1. The extracellular enzyme production was done by submerged fermentation using Mendel’s and Sternberg’s medium (MSM), and its optimisation was done using one factor at a time (OFAT). The presence of xylanase was confirmed by sodium dodecyl sulphate-polyacrylamide gel electrophoresis (SDS-PAGE) and zymography. In addition, the profiling of extracellular proteome of Aspergillus oryzae LC1 was carried out by liquid chromatography coupled tandem mass spectrometry (LC-MS/MS). In this study, media optimisation showed 5.7-fold increase in xylanase activity. The multiple bands observed in zymography revealed the presence of various forms of xylanase. A total of 73 proteins were identified in LC-MS/MS analysis. Functional classification showed that the hydrolytic enzymes consisted of 48% glycoside hydrolase, 11% proteases, 1% polysaccharide lyase and esterase’s, 9% oxidoreductases and 30% other proteins. A total of 26 families of glycosidic hydrolase were detected with other protein families such as serine peptidase, S, LysM, G-D-S-L, M35, carboxyl esterase (CE1), pectate lyase (PL) and oxidoreductases. Among the huge diversity of synergistically acting biomass cleaving enzymes, endo-1, 4-β xylanase with isoforms: xyn F1, xyn B, β xylanase and xyn 11A belonging to GH10 family covered the major portion of the total percentage of identified proteins. As per our knowledge, this is the first report of extracellular proteome analysis of Aspergillus oryzae LC1 suggesting its capability for recombinant expression and evaluation in hemicellulose deconstruction applications.






Similar content being viewed by others
Abbreviations
- MSM:
-
Mendel’s and Sternberg’s Medium
- OFAT:
-
One Factor at a Time
- SDS-PAGE:
-
Sodium Dodecyl Sulphate-Polyacrylamide Gel Electrophoresis
- LC-MS/MS:
-
Liquid chromatogrpahy coupled tandem mass spectrometry
- CE1:
-
Carboxy esterase
- PL:
-
Pectate lyase
- SCP:
-
Single cell protein
- CzDB:
-
Czapek Dox broth
- BSMwb:
-
Basal Salt Media with wheat bran
- DNSA:
-
Di-nitro Salicylic Acid
- TCA:
-
Trichloroacetic acid
- ACN:
-
Acetonitrile
References
Bakri Y, Masson M, Thonart P (2010) Isolation and identification of two new fungal strains for xylanase production. Appl Biochem Biotechnol 162(6):1626–1634. https://doi.org/10.1007/s12010-010-8944-x
Bhardwaj N, Chanda K, Kumar B, Prasad HK, Sharma GD, Verma P (2017) Statistical optimization of nutritional and physical parameters for xylanase production from newly isolated Aspergillus oryzae LC1 and its application in the hydrolysis of lignocellulosic agro-residues. BioResour 12(4):8519–8538. https://doi.org/10.15376/biores.12.4.8519-8538
Biely P, Cziszárová M, Agger JW, Li XL, Puchart V, Vršanská M, Westereng B (2014) Trichoderma reesei CE16 acetyl esterase and its role in enzymatic degradation of acetylated hemicellulose. Biochem Biophys Acta (BBA)-Gen Subj 1840(1):516–525. https://doi.org/10.1016/j.bbagen.2013.10.008
Braaksma M, Martens-Uzunova ES, Punt PJ, Schaap PJ (2010) An inventory of the Aspergillus niger secretome by combining in silico predictions with shotgun proteomics data. BMC Genomics 11(1):584. https://doi.org/10.1186/1471-2164-11-584
Bule MV, Chaudhary I, Gao AH, Chen S (2016) Effects of extracellular proteome on wheat straw pretreatment during solid-state fermentation of Phlebia radiata ATCC 64658. Int Biodeteriorat Biodegradat 109:36–44. https://doi.org/10.1016/j.ibiod.2015.12.002
Champer J, Ito JI, Clemons KV, Stevens DA, Kalkum M (2016) Proteomic analysis of pathogenic fungi reveals highly expressed conserved cell wall proteins. J Fungi 2(1):6. https://doi.org/10.3390/jof2010006
Chaturvedi V, Verma P (2013) An overview of key pretreatment processes employed for bioconversion of lignocellulosic biomass into biofuels and value added products. 3 Biotech 3(5):415–431. https://doi.org/10.1007/s13205-013-0167-8
Chutani P, Sharma KK (2015) Biochemical evaluation of xylanases from various filamentous fungi and their application for the deinking of ozone treated newspaper pulp. Carbohydr Polym 127:54–63. https://doi.org/10.1016/j.carbpol.2015.03.053
Dhillon GS, Kaur S, Brar SK, Gassara F, Verma M (2012) Improved xylanase production using apple pomace waste by Aspergillus niger in koji fermentation. Eng Life Sci 12(2):198–208. https://doi.org/10.1002/elsc.201100102
Duck N, Carr B, Koziel M, Carozzi N, Berg B (2003) Methods for enzymatic hydrolysis of lignocellulose:Google Patents. https://patents.google.com/patent/US20040005674A1/en
Eugenio LI, Méndez-Líter JA, Nieto-Domínguez M, Alonso L, Gil-Muñoz J, Barriuso J, Prieto A, Martínez MJ (2017) Differential β-glucosidase expression as a function of carbon source availability in Talaromyces amestolkiae: a genomic and proteomic approach. Biotechnol Biofuels 10(1):161. https://doi.org/10.1186/s13068-017-0844-7
Ferreira V, Da Silva R, Silva D, Gomes E (2010) Production of pectate lyase by Penicillium viridicatum RFC3 in solid-state and submerged fermentation. Internat J Microbiol 2010:1–8. https://doi.org/10.1155/2010/276590
Fic E, Kedracka-Krok S, Jankowska U, Pirog A, Dziedzicka-Wasylewska M (2010) Comparison of protein precipitation methods for various rat brain structures prior to proteomic analysis. Electrophoresis 31(21):3573–3579. https://doi.org/10.1002/elps.201000197
Fukuda M, Watanabe S, Yoshida S, Itoh H, Itoh Y, Kamio Y, Kaneko J (2010) Cell surface xylanases of the glycoside hydrolase family 10 are essential for xylan utilization by Paenibacillus sp. W-61 as generators of xylo-oligosaccharide inducers for the xylanase genes. J Bacteriol 192(8):2210–2219. https://doi.org/10.1128/JB.01406-09
Ghio S, Insani EM, Piccinni FE, Talia PM, Grasso DH, Campos E (2016) GH10 XynA is the main xylanase identified in the crude enzymatic extract of Paenibacillus sp. A59 when grown on xylan or lignocellulosic biomass. Microbiol Res 186:16–26. https://doi.org/10.1016/j.micres.2016.02.006
Ghose T, Bisaria VS (1987) Measurement of hemicellulase activities: part I xylanases. Pure Appl Chem 59(12):1739–1751. https://doi.org/10.1351/pac198759121739
Girard V, Dieryckx C, Job C, Job D (2013) Secretomes: the fungal strike force. Proteom 13(3–4):597–608. https://doi.org/10.1002/pmic.201200282
Giridhar PV, Chandra T (2010) Production of novel halo-alkali-thermo-stable xylanase by a newly isolated moderately halophilic and alkali-tolerant Gracilibacillus sp. TSCPVG. Process Biochem 45(10):1730–1737. https://doi.org/10.1016/j.procbio.2010.07.012
Grosse-Holz F, Kelly S, Blaskowski S, Kaschani F, Kaiser M, van der Hoorn RA (2018) The transcriptome, extracellular proteome and active secretome of agroinfiltrated Nicotiana benthamiana uncover a large, diverse protease repertoire. Plant Biotechnol J 16(5):1068–1084. https://doi.org/10.1111/pbi.12852
Hashemi M, Mousavi SM, Razavi SH, Shojaosadati SA (2013) Comparison of submerged and solid state fermentation systems effects on the catalytic activity of Bacillus sp. KR-8104 α-amylase at different pH and temperatures. Ind Crops Prod 43:661–667. https://doi.org/10.1016/j.indcrop.2012.08.002
Irfan M, Nadeem M, Syed Q (2014) One-factor-at-a-time (OFAT) optimization of xylanase production from Trichoderma viride-IR05 in solid-state fermentation. J Radiat Res Appl Sci 7(3):317–326. https://doi.org/10.1016/j.jrras.2014.04.004
Jeon J, Mok HJ, Choi Y, Park SC, Jo H, Her J, Han JK, Kim YK, Kim KP, Ban C (2017) Proteomic analysis of extracellular vesicles derived from Propionibacterium acnes. Proteom Clinic Appl 11(1–2):1600040. https://doi.org/10.1002/prca.201600040
Korwar AM, Vannuruswamy G, Jagadeesha prasad MG, Jayaramaiah RH, Bhat S, Regin BS, Balasubramanyam M (2015) Development of diagnostic fragment ion library for glycated peptides of human serum albumin: targeted quantification in prediabetic, diabetic, and micro albuminuria plasma by parallel reaction monitoring, SWATH, and MSE. Mol Cell Proteomics 14(8):2150–2159. https://doi.org/10.1074/mcp.M115.050518
Kumar V, Satyanarayana T (2011) Applicability of thermo-alkali-stable and cellulase-free xylanase from a novel thermo-halo-alkaliphilic Bacillus halodurans in producing xylooligosaccharides. Biotechnol Lett 33(11):2279–2285. https://doi.org/10.1007/s10529-011-0698-1
Kumar V, Satyanarayana T (2013) Biochemical and thermodynamic characteristics of thermo-alkali-stable xylanase from a novel polyextremophilic Bacillus halodurans TSEV1. Extremophiles 17(5):797–808. https://doi.org/10.1007/s00792-013-0565-1
Kumar NV, Rani ME, Gunaseeli R, Kannan ND (2018) Paper pulp modification and deinking efficiency of cellulase-xylanase complex from Escherichia coli SD5. Int J Biol Macromol 111:289–295. https://doi.org/10.1016/j.ijbiomac.2017.12.126
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227(5259):680–685. https://doi.org/10.1038/227680a0
Liao H, Sun S, Wang P, Bi W, Tan S, Wei Z, Shen Q (2014) A new acidophilic endo-β-1, 4-xylanase from Penicillium oxalicum: cloning, purification, and insights into the influence of metal ions on xylanase activity. J Indus Microbiol Biotechnol 41(7):1071–1083. https://doi.org/10.1007/s10295-014-1453-0
Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with the Folin phenol reagent. J Biol Chem 193(1):265–275 http://www.jbc.org/content/193/1/265.long
Miller GL (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugar. Analy Chem 31(3):426–428. https://doi.org/10.1021/ac60147a030
Nagar S, Gupta VK, Kumar D, Kumar L, Kuhad RC (2010) Production and optimization of cellulase-free, alkali-stable xylanase by Bacillus pumilus SV-85S in submerged fermentation. J Ind Microbiol Biotechnol 37(1):71–83. https://doi.org/10.1007/s10295-009-0650-8
Nagar S, Mittal A, Kumar D, Gupta VK (2012) Production of alkali tolerant cellulase free xylanase in high levels by Bacillus pumilus SV-205. Int J Biol Macromol 50(2):414–420. https://doi.org/10.1007/s10295-009-0650-8
Pandya JJ, Gupte A (2012) Production of xylanase under solid-state fermentation by Aspergillus tubingensis JP-1 and its application. Bioprocess Biosyst Eng 35(5):769–779. https://doi.org/10.1007/s00449-011-0657-1
Park Y, Kang S, Lee J, Hong S, Kim S (2002) Xylanase production in solid state fermentation by Aspergillus niger mutant using statistical experimental designs. Appl Microbiol Biotechnol 58(6):761–766. https://doi.org/10.1007/s00253-002-0965-0
Sánchez-Herrera LM, Ramos-Valdivia AC, De La Torre M, Salgado LM, Ponce-Noyola T (2007) Differential expression of cellulases and xylanases by Cellulomonas flavigena grown on different carbon sources. App Microbiol Biotechnol 77(3):589–595. https://doi.org/10.1007/s00253-007-1190-7
Shah AR, Madamwar D (2005) Xylanase production by a newly isolated Aspergillus foetidus strain and its characterization. Process Biochem 40(5):1763–1771. https://doi.org/10.1016/j.procbio.2004.06.041
Silva L, Terrasan CRF, Carmona EC (2015) Purification and characterization of xylanases from Trichoderma inhamatum. Electron J Biotechnol 18(4):307–313. https://doi.org/10.1016/j.ejbt.2015.06.001
Singh S, Tiwari R, Renuse S, Pranaw K, Nain L (2015) Proteomic analysis of Streptomyces sp. ssr-198 grown on paddy straw. J Basic Microbio 55(6):790–797. https://doi.org/10.1002/jobm.201400639
Straathof AJ, Panke S, Schmid A (2002) The production of fine chemicals by bio transformations. Curr Opin Biotechnol 13(6):548–556. https://doi.org/10.1016/S0958-1669(02)00360-9
Tallapragada P, Venkatesh K (2011) Isolation, identification and optimization of xylanase enzyme produced by Aspergillus niger under submerged fermentation. J Microbiol Biotechnol Res 1(4):137–147 https://www.jmbronline.com/index.php/JMBR/article/view/66
Tiwari R, Singh S, Singh N, Adak A, Rana S, Sharma A, Nain L (2014) Unwrapping the hydrolytic system of the phytopathogenic fungus Phoma exigua by secretome analysis. Process Biochem 49(10):1630–1636. https://doi.org/10.1016/j.procbio.2014.06.023
Umsza-Guez MA, Díaz AB, Ory Id, Blandino A, Gomes E, Caro I (2011) Xylanase production by Aspergillus awamori under solid state fermentation conditions on tomato pomace. Brazil J Microbiol 42(4):1585–1597. https://doi.org/10.1590/S1517-83822011000400046
Watanabe J, Tanaka H, Mogi Y, Yamazaki T, Suzuki K, Watanabe T, Akita O (2011) Loss of Aspergillus oryzae amyR function indirectly affects hemicellulolytic and cellulolytic enzyme production. J Biosci Bioeng 111(4):408–413. https://doi.org/10.1016/j.jbiosc.2010.12.006
Acknowledgements
The authors are thankful to the Department of Biotechnology, Government of India, for providing the financial support (Grant No. BT/304/NE/TBP/2012).
Funding
This study was funded by the Department of Biotechnology, Government of India, for providing the financial support (Grant No. BT/304/NE/TBP/2012).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Research involving human participants and/or animals
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed consent
Informed consent is not required in this study.
Rights and permissions
About this article
Cite this article
Bhardwaj, N., Verma, V.K., Chaturvedi, V. et al. GH10 XynF1 and Xyn11A: the predominant xylanase identified in the profiling of extracellular proteome of Aspergillus oryzae LC1. Ann Microbiol 68, 731–742 (2018). https://doi.org/10.1007/s13213-018-1378-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13213-018-1378-3