Abstract
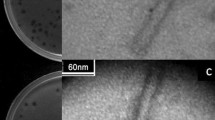
This study was designed to investigate the possibility of using bacteriophages for the detection of viable Shigella boydii in food products. A Shigella bacteriophage belonging to a member of the Siphoviridae family was isolated from swine fecal samples. The free bacteriophages were highly stable against pH 4.0 to 9.0 and temperature change (z-value = 17.1 °C). The bacteriophage amplification assay was able to selectively detect S. boydii in a bacterial mixture of Escherichia coli O157:H7, Listeria monocytogenes, and Salmonella Typhimurium. The number of S. boydii bacteriophages enumerated by the amplification assay was highly correlated with the number of viable S. boydii in single (r = 0.987) and mixed (r = 0.969) cultures. The bacteriophage-based detection of S. boydii was highly reproducible in lettuce (6.3 log CFU/ml and 4.9 log PFU/ml) and cooked chicken breasts (6.1 log CFU/ml and 6.0 log PFU/ml). These results suggest that the bacteriophage amplification assay can be used as an alternative method for rapid, selective, and cost-effective detection of S. boydii in food products, and provides useful information for designing a quick and simple detection kit.




Similar content being viewed by others
References
Bielke L, Higgins S, Donoghue A, Donoghue D, Hargis BM (2007) Salmonella host range of bacteriophages that infect multiple genera. Poult Sci 86:2536–2540
Botsaris G, Slana I, Liapi M, Dodd C, Economides C, Rees C, Pavlik I (2010) Rapid detection methods for viable Mycobacterium avium subspecies paratuberculosis in milk and cheese. Int J Food Microbiol 141:S87–S90
de Siqueira RS, Dodd CER, Rees CED (2006) Evaluation of the natural virucidal activity of teas for use in the phage amplification assay. Int J Food Microbiol 111:259–262
Gracias KS, McKillip JL (2004) A review of conventional detection and enumeration methods for pathogenic bacteria in food. Can J Microbiol 50:883–890
Javed MA, Poshtiban S, Arutyunov D, Evoy S, Szymanski CM (2013) Bacteriophage receptor binding protein based assays for the simultaneous detection of Campylobacter jejuni and Campylobacter coli. PLoS ONE 8:1–10
Li Y, Cao B, Liu B, Liu D, Gao Q, Peng X, Wu J, Bastin DA, Feng L, Wang L (2009) Molecular detection of all 34 distinct O-antigen forms of Shigella. J Med Microbiol 58:69–81
Lleo MM, Bonato B, Tafi MC, Signoretto C, Pruzzo C, Canepari P (2005) Molecular vs culture methods for the detection of bacterial faecal indicators in groundwater for human use. Lett Appl Microbiol 40:289–294
Mandal PK, Biswas AK, Choi K, Pal UK (2011) Methods for rapid detection of foodborne pathogens: an overview. Am J Food Technol 6:87–102
McNerney R, Wilson SM, Sidhu AM, Harley VS, Al Suwaidi Z, Nye PM, Parish T, Stoker NG (1998) Inactivation of mycobacteriophage D29 using ferrous ammonium sulphate as a tool for the detection of viable Mycobacterium smegmatis and M. tuberculosis. Res Microbiol 149:487–495
Mishra CK, Choi TJ, Kang SC (2012) Isolation and characterization of a bacteriophage F20 virulent to Enterobacter aerogenes. J Gen Virol 93:2310–2314
Mokhtari W, Nsaibia S, Majouri D, Ben Hassen A, Gharbi A, Aouni M (2012) Detection and characterization of Shigella species isolated from food and human stool samples in Nabeul, Tunisia, by molecular methods and culture techniques. J Appl Microbiol 113:209–222
Oliveira IC, Almeida RCC, Hofer E, Almeida PF (2012) Bacteriophage amplification assay for detection of Listeria spp. using virucidal laser treatment. Braz J Microbiol 43:1128–1136
Park DJ, Drobniewski FA, Meyer A, Wilson SM (2003) Use of a phage-based assay for phenotypic detection of Mycobacteria directly from sputum. J Clin Microbiol 41:680–688
Riahi R, Mach KE, Mohan R, Liao JC, Wong PK (2011) Molecular detection of bacterial pathogens using microparticle enhanced double-stranded DNA probes. Anal Chem 83:6349–6354
Smartt A, Xu T, Jegier P, Carswell J, Blount S, Sayler G, Ripp S (2012) Pathogen detection using engineered bacteriophages. Anal Bioanal Chem 402:3127–3146
Stewart J, Denyer N, Linley D (1998) The specific and sensitive detection of bacterial pathogens within 4 h using bacteriophage amplification. J Appl Microbiol 84:777–783
Thiem VD, Sethabutr O, von Seidlein L, Van Tung T, Canh DG, Chien BT, Tho LH, Lee H, Houng H-S, Hale TL, Clemens JD, Mason C, Trach DD (2004) Detection of Shigella by a PCR assay targeting the ipaH gene suggests increased prevalence of Shigellosis in Nha Trang, Vietnam. J Clin Microbiol 42:2031–2035
Wang S-J, Chen JH (2012) A rapid and specific PCR method for the detection of Shigella spp. in spiked samples. J Food Drug Anal 20:59–65
Wyckoff EE, Duncan D, Torres AG, Mills M, Maase K, Payne SM (1998) Structure of the Shigella dysenteriae haem transport locus and its phylogenetic distribution in enteric bacteria. Mol Microbiol 28:1139–1152
Acknowledgments
This study was supported by a 2015 Research Grant from Kangwon National University (Grant No. 520150317). The authors also thank Ms. Myeong-Seon Jung of the Korea Basic Science Institute (KBSI, Chuncheon, Korea) for her technical assistance in the analytical field using transmission electron microscopy (TEM).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jung, LS., Ahn, J. Evaluation of bacteriophage amplification assay for rapid detection of Shigella boydii in food systems. Ann Microbiol 66, 883–888 (2016). https://doi.org/10.1007/s13213-015-1178-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13213-015-1178-y