Abstract
Sensitive, effective, and quantitative analysis of infectious pathogens is an important task for the prevention of human health threats. Herein, we present an advanced approach to producing gene-encapsulated microdroplets for quantitative analysis using a micropatterned metal mold and injection molding technique with an automatically operated system. An injection molded microdroplet generation device was successfully fabricated with a minimum channel width of 30 μm and optimized to produce 100 μm diameter droplets. The optimized microchannel design and flow rate also enable the production of stable numbers of microdroplets (~ 16,000 droplets). To verify the applicability of our device and system to droplet-based digital PCR analysis, Escherichia coli (E. coli) O157:H7 was selected as a model bacterial pathogen, and the stx2 gene was amplified in the microdroplets. The generated microdroplets exhibit both chemical and mechanical stability, and our results are similar to those obtained by a commercially available method. Accordingly, the usefulness of the microdroplet generative device and system is confirmed as a simple, fast, and reliable tool for the quantitative molecular analysis of infectious diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
After the outbreak of COVID-19 and other infectious diseases, it has become urgent to develop precise, reliable, and accurate diagnostic methods [1,2,3,4,5]. For this purpose, molecular diagnostics based on the nucleic acid assay, especially using digital nucleic acid analysis, has generated great interest due to its high accuracy, specificity, and absolute quantification compared to traditional culture- and/or immune-based diagnostics [6,7,8,9,10]. The digital nucleic acid assay includes compartmentalization, amplification, and fluorescence analysis [11, 12]. In detail, a solution containing nucleic acid is divided and randomly distributed into a large number of micro-volume compartments. The uniformly sized partitions act as independent reactors for amplification and enable the quantification of target nucleic acids simply by detecting and counting the fluorescent signals from each partition. In addition, the positive and negative partition ratio provide the copy number of the target nucleic acids. Due to these unique characteristics, digital nucleic acid assay is widely applied in various fields, such as biological research and clinical diagnostics.
Many prior research efforts have been undertaken to obtain both devices and systems that can offer reliable, mass producible, simple, and rapid paths to compartmentalization [13, 14]. So far, several methods including solid microchambers/wells, water-in-oil emulsion, and printing have been employed [15,16,17]. The water-in-oil emulsion-assisted microdroplet-based method has become the most effective way to meet the numerous requirements of digital nucleic acid amplification and analysis [18,19,20]. However, such methods still suffer from various technical issues including the need for ways to mass-produce PDMS-free, cost effective devices and the need for automated machines to operate the devices. Recently, MEMS technology has been under development as an option to achieve a precisely micropatterned (< 50 μm) metal mold that is suitable for injection molding for the mass production of plastic-based microfluidic devices that are easy to incorporate into an automated system [21, 22]. Moreover, the plastic device offers many benefits and features such as reliability, reproducibility, accuracy, ease-to-use, and cost-effectiveness [23]. Based on these advantages, these devices are being widely used for diverse digital nucleic acid amplification and analysis applications.
In addition to the devices themselves, an automated system must be developed to satisfy numerous requirements such as precise controllability and large-scale droplet production. The conventional methods are simply an injection of surfactant/oil mixtures and buffers using syringe pumps for controlling both flow rates. However, these methods require relatively time-consuming processes to inject a solution into the syringes, manual connection of all tubes to devices, and difficulties in droplet recovery. Therefore, it is vital to develop an automated system to generate microdroplets.
In this study, we developed a mass producible device to generate microdroplets and an automated system for the microdroplet-based digital nucleic acid analysis using MEMS-based micromolds and an injection molding method. To fabricate the device, an injection molding technique with polycarbonate was used, and the device was sealed using thin adhesive-coated polyethylene terephthalate (PET) and a hot sealing plate. The buffer and surfactant/oil mixture enabled the generation of 10, 000 individual microdroplets with the assistance of an automated system. To simplify the microdroplet generation system, it simply operates by a vacuum pump and valve using negative pressure control. The generated microdroplets were subjected to digital nucleic acid analysis after the amplification processes. To confirm the performance, an infectious foodborne pathogen, Escherichia coli (E. coli) O157:H7, was used as a realistic model. Moreover, the device and system were compared with a commercially available instrument (QX200 droplet digital PCR system, Bio-Rad, USA). The micropatterned device and automated droplet generation system demonstrated excellent and precise droplet generation performance with the benefits of simple fabrication, high sensitivity, and reliability when used with actual pathogenic bacteria.
2 Materials and Methods
2.1 Design of Micro-injection Molded Droplet Generation Chip
Figure 1 shows a schematic illustration of the droplet generation microchip. The droplet microchip (size: 88.00 × 23.25 × 7.05 mm) is composed of eight droplet generation units, each of which consists of an oil inlet, a sample inlet, and a droplet outlet on the top (Fig. 1A) and a droplet generation channel on the bottom (Fig. 1B). Figure 1C displays the details of the droplet generation channel. An oil inlet is connected to 26.23-mm-long dual oil channels for supplying an oil phase solution. The oil channels meet the 16.50-mm-long PCR reagent channel that serves to deliver the PCR mixture to the droplet. The generated droplets flow through the droplet channel toward the droplet outlet and vacuum port for downstream analysis. Figure 1D shows a photomicrograph of the droplet generation microchip.
A Schematic illustration of the top view of the micro-injection molded droplet generation chip with eight droplet generation units, each of which includes an oil reservoir, a PCR reagent reservoir, and a droplet outlet/vacuum port. B Bottom view of the micro-injection molded droplet generation chip. Eight units of droplet generation channels were engraved on the backside of the microchip, and a laminating film was used to cover the channels to finalize the channel fabrication. C Enlarged view of the droplet generation channel. The droplet generation channel consists of an oil inlet that is connected to two oil channels, a PCR reagent inlet with a PCR reagent channel and a droplet outlet/vacuum port with a droplet channel. D An actual digital image of the droplet generation microchip
2.2 Fabrication of the Droplet Generation Chip
A nickel (Ni) plate with a diameter of 150 mm and a thickness of 1 mm was purchased and used for Ni electroplating. The micropatterned Ni plate was prepared using a combination of photolithography and electroplating. To fabricate the micropattern, photoresist (THB 151 N, JSR Micro, USA) was dispersed on the plate, spin-coated at 1,000 rpm for 40 s, and baked at 120 °C for 5 min. After soft bake processing, the wafer was exposed to the UV, developed, and then rinsed with distilled water. For the nickel deposition, an electroplating solution was employed in a bath at a temperature of 55 °C and 0.243 Å for 125 min. After the nickel deposition, the photoresist was finally removed and the micropatterned Ni was further diced to be used as a metal mold.
To fabricate the device, polycarbonate (PC) pellets were purchased from Lotte Chemical (Republic of Korea) and placed in a dehumidifier (Purpose VHM5-LC, HANSE Co.) at 70 °C to remove any remaining moisture. The molten PC was injected into the micromold at a speed of 100–150 mm s−1 and a pressure of 500–1500 bar to replicate the shape of the mold [24]. The mold temperature was set to ~ 150 °C, which is below the glass transition temperature of the PC. In addition, the device was fabricated using an injection molding packing pressure ranging from 300 to 1500 bar that was applied for 20 s.
2.3 Microfluidic Droplet Preparation with the Automatic Droplet Generation System
Figure 3 shows the droplet generation sequence. Prior to loading a droplet generation microchip into the automatic droplet generation system (Bio T&S, Daejeon, Korea) for droplet preparation, 20 μL of PCR mixture was injected into the sample inlet, while 70 μL of QX200™ droplet generation oil for EvaGreen (Bio-Rad) was added in the oil inlet. The microchip was placed into the automatic droplet generation system, and the lid of the generator was lowered down toward the microchip. Droplets were produced by applying a − 80 kPa vacuum to the vacuum port of the droplet generation microchip for 2 min. To confirm the stable production of droplets using our system, the generated droplets were observed with an optical microscope to measure the size of droplets, and the number of droplets was counted with a QX200™ droplet reader. To validate the droplet generation performance for the PCR amplification, a control experiment was carried out with a QX200™ droplet generator (Bio-Rad). PCR mixture of 20 μL and 70 μL of QX200™ droplet generation oil for EvaGreen (Bio-Rad) were loaded into the sample well and oil well, respectively, of the DG8 cartridge (Bio-Rad). Droplets were generated using a QX200™ droplet generator. The droplets, which were produced using both our system and QX200™ droplet generator, were recovered using a pipette and transferred into a 96-well plate (Bio-Rad) for performing a digital PCR.
2.4 Preparation of Cell and Droplet PCR Reagent
E.coli O157:H7(ATCC 43,894) was provided by the Korea Research Institute of Bioscience and Biotechnology and was cultured in Luria–Bertani (LB) broth for 18 h at 37 °C in a shaking incubator. The genomic DNA (gDNA) of the bacteria was extracted using a G-spin™ total DNA extraction kit (iNtRON Biotechnology). After measuring the gDNA concentration using a NanoDrop 1000 spectrophotometer (Thermo Fisher Scientific), a tenfold serial dilution using 10 ng/μL of gDNA was carried out until the gDNA concentration reached 1 pg/μL for a limit of detection test. Each 20 μL volume of the PCR mixture consists of 10 μL of 2 × QX200™ ddPCR™ EvaGreen® Supermix (Bio-Rad), 1 μL of extracted gDNA of E. coli O157:H7, 1.25 μL (10 μM) of forward and reverse primers, and 6.5 μL of diethylpyrocarbonate (DEPC)-treated water (Enzynomics). The sequence of the forward and reverse primer was synthesized as 5′-GAC CCG GCA CAA GCA TAA GC-3′ and (reverse primer) 5′-CCA CCT GCA GCA ACA AGA GG-3′, respectively [25]. After preparing droplets using PCR reagent, the resulting droplets were transferred to a 96-well plate (Bio-Rad) for target gene amplification. PCR amplification was conducted using a T100 thermal cycler (Bio-Rad) under the conditions of initial denaturation at 95 °C for 5 min, 40 thermal cycles of 95 °C for 30 s, 55 °C for 30 s, and signal stabilization at 4 °C for 5 min and 90 °C for 5 min (2 °C/s ramp rate) as specified in the QX200™ ddPCR™ EvaGreen® Supermix protocol provided by the manufacturer (Bio-Rad).
3 Results and Discussion
3.1 Design of the Micro-injection Molded Droplet Generation Device
Our proposed droplet generation chip is manufactured by a micro-injection molding method, which enables the pattern of a microsize channel to be formed on the droplet generation chip. A high-resolution micromolding technique is required because a flow-focusing microchannel part needs to be fabricated with the following specifications: two oil channels for continuous phase injection, each of which is 140 μm wide, a microdroplet formation channel with a width of 110 μm, and a 30-μm-wide PCR reagent channel. The device has eight chambers holding oil, PCR reagent, and generated droplets, respectively. The device is made of clear polycarbonate and sealed with adhesive-coated PET film to prevent any leakages of oils or reagents. The device is composed of eight sets of microchannels and chambers. Each set has one oil chamber with two channels to produce shear and one PCR reagent chamber with one channel. The width and height of the microchannels are 100 and 50 μm, respectively, while the droplet formation channels are 30 μm wide. The device is precisely designed to produce microdroplets with a uniform size and shape using the flow-focusing technique. To optimize droplet generation using the flow-focusing technique, the device has two oil channels and one PCR reagent channel. Once the PCR reagent flows through the orifice, the surfactant/oil mixture produces a shear force to disconnect the PCR reagent from the microdroplet.
A schematic illustration of the complete fabrication process, including both the conventional photolithography steps and our novel injection molding steps, is shown in Fig. 2. To fabricate a microfluidic patterned Ni mold, a thin layer of photoresist was applied over the planar Ni wafer. After deposition of Ni, the photoresist-coated Ni wafer was simply exposed to UV light and transferred with as-prepared mask designs onto the target Ni wafer. After developing the photoresist, the microfluidic channel patterns remained. Once the obtained microstructures were formed by the Ni electroplating process, the microfluidic channel patterns subsequently deposited with Ni metals and the microstructure was constructed on the Ni plate. After removing the remaining photoresist and dicing the Ni wafer, the wafer was loaded into the injection mold machine, and then the molten raw polymers were injected into the mold cavity and replicated the mirror image of the Ni wafer to produce the polymeric device. Once the polymeric microstructure was produced, it was carefully released from the mold, and the device was sealed by laminating with adhesive polymer film.
Schematic of the micro-injection molded droplet generation chip fabrication process. A Prepare a nickel wafer. B Spin-coat photoresist and perform a UV exposure with a droplet generation microchip mask. C Develop the photoresist. D Electroplate nickel on the exposed areas. E Strip the photoresist. F Dice the wafer to fabricate a microchip mold. G Injection mold the micro-injection molded droplet generation chip using a polycarbonate. H Laminate a PET film on the microchannel side
3.2 Performance of the Droplet Generation Device and System
Figure 3 illustrates the overall droplet generation processes. An automated droplet generation system was developed using negative pressure with the assistance of a vacuum pump and three-way valve (Fig. 3A). The vacuum pump and valve allowed the negative pressure to pull both oil and PCR reactant based on the droplet generation sequence as shown in Fig. 3B. Before operating the system, the surfactant/oil mixture and PCR reagents were loaded into the oil and reactor chambers, respectively. When the three-way valve was closed and the vacuum pump was turned on, the negative pressure started to build inside the channels. Once the desired negative pressure was reached, the valve was opened in the direction between the pump and chamber to pull in both oil and PCR reagent. When the valve was opened, all the oil and reagent passed rapidly through the orifice inside the device. Subsequently, a shear force was applied to the PCR reagent, and uniform microdroplets were formed and collected inside the product chamber. Finally, when the reaction was over, the valve was completely opened to balance the pressure between the environment and the chambers.
The sequence described above was conducted using embedded firmware and circuit boards inside of the system. Each process was calculated based on the loaded volumes, microfluidic channels, and flow rates of both oil and PCR reagents. Once the droplet has been generated, each recovery chamber contains a uniform microdroplet.
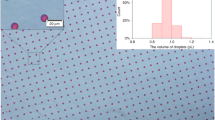
The shape and size of the generated droplets were carefully investigated using optical microscopy at different magnifications. All the droplets had a spherical shape, and there were ~ 16, 000 droplets with a diameter of 100 μm as shown in Fig. 4A, C. Interestingly, all eight chambers show microdroplets of almost the same shape and size, and no merged droplets were found in the chambers. The numbers of droplets were also analyzed using a droplet reading system. Moreover, there were no significant droplet merging, irregular droplet size, or air bubble generation during the automated procedure.
A Micrographs at 4 × and 20 × magnification of droplets generated from eight different droplet generation units and B droplets after PCR thermocycling to verify the thermal stability of the generated droplets during the PCR. (Scale bars: 500 μm with 4 × magnification and 200 μm with 20 × magnification). C The total number of generated droplets and average droplet diameter from eight different droplet generation units
In addition, the thermal stability of the generated droplets was verified by observing droplets using optical microscopy after performing PCR thermal cycling as shown in Fig. 4B. Despite the PCR thermal cycling, no significant changes were found in the size or shape of the droplets, which indicates that the generated droplets are reliable enough to use for droplet-based digital PCR.
3.3 Droplet-Based PCR Analysis for Foodborne Pathogen Detection
To show the feasibility of the system, a digital droplet PCR targeting the stx2 gene of E. coli O157:H7 was performed with the droplet device and amplified. Pathogenic bacteria of E. coli O157:H7 are commonly known to cause foodborne illness. To confirm the performance, a serial dilution consisting of four 1:10 dilutions of target DNA was tested using both our developed system and a commercially available system. Figure 5 shows the performance comparison of our developed droplet device and system versus the commercially available droplet digital PCR using different concentrations of the gene of E. coli O157:H7 from 0.001 to 1 ng/µL. At a DNA concentration of 1 ng/µL, both the control system and our system show almost identical values of 80% for the ratio of positive droplets to total droplets. When we loaded about 0.001 ng/µL of DNA into the droplets, the ratio of positive droplets to total droplets was 0.24%, but the agreement between the two systems was still excellent, and even this low amount of DNA can be detected and analyzed. The experiment results from our system were very similar to the results obtained with the commercially available system. Therefore, the developed droplet device and the system have been shown to be reliable.
4 Conclusion
We have demonstrated the successful production of uniform-sized microdroplets using a polymeric microdroplet chip and an automated droplet generation system. The shear stress driven by a rapid flow of both PCR reagent and oil using negative pressure enables the production of reliable and uniform microdroplets without the generation of air bubbles or the merging of droplets. Compared to the existing digital PCR devices and machines, our system offers easy access and similar performance. Moreover, the proposed device and system are cost effective, stable, reliable, and easy to use for digital droplet PCR. The digital PCR performance was also investigated using an infectious pathogen target of the stx2 gene of E. coli O157:H7, and the developed droplet-generating device and system could detect the target pathogenic genes at levels ranging from 0.001 to 1 ng/µL. In addition, the developed system shows excellent performance and is reliable enough to be commercialized. These findings confirm that the digital droplet PCR technique is useful for the development of a highly reliable, reproducible, and absolute quantification analysis for clinical applications.
References
Sun, Y., et al.: Digital PCR assay for the effective detection of COVID-19 patients with SARS-CoV-2 low viral load. J. Virol. Methods 295, 114185 (2021)
Ahmad, S.: A review of COVID-19 (Coronavirus Disease-2019) diagnosis, treatments and prevention. Eurasian J. Med. Oncol. 4, 116–125 (2020)
Tang, Y.-W., Procop, G.W., Persing, D.H.: Molecular diagnostics of infectious diseases. Clin. Chem. 43, 2021–2038 (1997)
Kim, K.H., et al.: Touchable 3D hierarchically structured polyaniline nanoweb for capture and detection of pathogenic bacteria. Nano Converg. 8, 1–11 (2021)
Yoo, H.J., et al.: Discrimination and isolation of the virus from free RNA fragments for the highly sensitive measurement of SARS-CoV-2 abundance on surfaces using a graphene oxide nano surface. Nano Converg. 8, 31 (2021)
Song, H., Chen, D.L., Ismagilov, R.F., Ismagilov, R.F.: Droplet-based microfluidics reactions in droplets in microfluidic channels. Angew Chem. Inter. Ed. 45, 7336–7356 (2006)
Ottesen, E.A., Hong, J.W., Quake, S.R., Leadbetter, J.R.: Microfluidic digital PCR enables multigene analysis of individual environmental bacteria. Science 314, 1464–1467 (2006)
Park, J., et al.: Pushbutton-activated microfluidic dropenser for droplet digital PCR. Biosens. Bioelectron. 181, 113159 (2021)
Jang, M., et al.: Droplet-based digital PCR system for detection of single-cell level of foodborne pathogens. Biochip J. 11, 329–337 (2017)
Aladese, A.D., Jeong, H.H.: Recent developments in 3D printing of droplet-based microfluidics. Biochip J. 15, 313–333 (2021)
Quan, P.L., Sauzade, M., Brouzes, E.: DPCR: a technology review. Sensors 18, 2171 (2018)
Whale, A.S., Huggett, J.F., Tzonev, S.: Fundamentals of multiplexing with digital PCR. Biomol. Detect. Quantif. 10, 15–23 (2016)
Baker, M.: Digital PCR hits its stride. Nat. Methods 9, 541–544 (2012)
Zhang, H., et al.: Digital PCR system development accelerator—a methodology to emulate dPCR results. Sens. Actuators B Chem. 358, 131527 (2022)
Perkins, G., Lu, H., Garlan, F., Taly, V.: Droplet-based digital PCR: application in cancer research. Adv. Clin. Chem. 79, 43–91 (2017)
Choi, Y., et al.: All-in-one pumpless portable genetic analysis microsystem for rapid naked-eye detection. Sens. Actuators B Chem. 344, 130307 (2021)
Sun, Y., et al.: Wet-etched microchamber array digital PCR chip for SARS-CoV-2 virus and ultra-early stage lung cancer quantitative detection. ACS Omega 7, 1819–1826 (2022)
Sreejith, K.R., Ooi, C.H., Jin, J., Dao, D.V., Nguyen, N.T.: Digital polymerase chain reaction technology-recent advances and future perspectives. Lab Chip 18, 3717–3732 (2018)
Tanaka, H., et al.: Hands-off preparation of monodisperse emulsion droplets using a poly(dimethylsiloxane) microfluidic chip for droplet digital PCR. Anal. Chem. 87, 4134–1443 (2015)
Zhao, S., Zhang, Z., Hu, F., Wu, J., Peng, N.: Massive droplet generation for digital PCR via a smart step emulsification chip integrated in a reaction tube. Analyst 146, 1559–1568 (2021)
Ahrberg, C.D., et al.: Plasmonic heating-based portable digital PCR system. Lab Chip 20, 3560–3568 (2020)
Meng, X., Yu, Y., Jin, G.: Numerical simulation and experimental verification of droplet generation in microfluidic digital PCR chip. Micromachines 12, 409 (2021)
Fiorini, G.S., Chiu, D.T.: Disposable microfluidic devices: fabrication, function, and application. Biotechniques 38, 249–446 (2005)
Kim, B.I., et al.: A continuous tilting of micromolds for fabricating polymeric microstructures in microinjection. Lab Chip 13, 4321–4325 (2013)
You, J.B., et al.: Surface-modified mesh filter for direct nucleic acid extraction and its application to gene expression analysis. Adv. Healthc. Mater. 6, 1700642 (2017)
Funding
This work was supported by the Nanomedical Devices Development Project of NNFC in 2022 and the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (2021-DD-RD-0012). This work was also supported by the Technology Innovation Program (20015577) and the K-Sensor Development Program (RS-2022-00154853 and RS-2022-00154855) funded By the Ministry of Trade, Industry & Energy (MOTIE, Korea).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jo, D., Kim, S.Y., Kang, H.W. et al. Micro-injection Molded Droplet Generation System for Digital PCR Application. BioChip J 16, 433–440 (2022). https://doi.org/10.1007/s13206-022-00079-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13206-022-00079-8