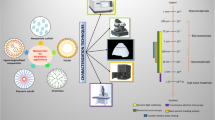
Abstract
The current frontiers in Biology thus in Medicine and Pharmacy are at the nanoscale. Indeed, this is the relevant scale for extracting or synthetizing, visualizing, counting, characterizing and/or modifying nanoparticles. Nanoparticles are highly diverse including: extracellular vesicles (e.g.: exosomes), proteins, viruses and nanovectors or drug delivery systems for instance. To quantify the concentration of nano-sized objects, a growing number of size-tracking instruments is being developed. However, to date, the generated data is only used to provide a concentration measurement. The objective of this study was to determine which sizes of nanoparticles are statistically significant between 2 groups of samples. Using different samples (in silico data; calibrated beads; various biological samples), an approach that statistically compares 2 groups of samples was developed and validated. The proof of concept of the proposed approach was illustrated with applications in the field of Biology, Medicine and Pharmacy using liposomes and extracellular vesicles. For the first time to our knowledge, our results suggest that the presented approach enables comparing different groups of biological samples. It may be extended to situations such as batch 1 versus batch 2; healthy versus disease or non-treated versus treated.




Similar content being viewed by others
Availability of data and materials
The datasets used and analyzed during the current study are available from the corresponding author on reasonable request.
Code availability
Custom code might be available from the corresponding author on reasonable request.
References
Alexis F et al (2008) New frontiers in nanotechnology for cancer treatment. Urol Oncol 26(1):74–85
Allen TM, Cullis PR (2004) Drug delivery systems: entering the mainstream. Science 303(5665):1818–1822
Allison DB et al (2006) Microarray data analysis: from disarray to consolidation and consensus. Nat Rev Genet 7(1):55–65
Bachmann MF, Jennings GT (2010) Vaccine delivery: a matter of size, geometry, kinetics and molecular patterns. Nat Rev Immunol 10(11):787–796
Barile L, Vassalli G (2017) Exosomes: Therapy delivery tools and biomarkers of diseases. Pharmacol Ther 174:63–78
Birmingham A et al (2009) Statistical methods for analysis of high-throughput RNA interference screens. Nat Methods 6(8):569–575
Bolstad BM et al (2003) A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19(2):185–193
Brossa A et al (2019) Alternative Strategies to Inhibit Tumor Vascularization. Int J Mol Sci 20(24):6180
Bullard JH et al (2010) Evaluation of statistical methods for normalization and differential expression in mRNA-Seq experiments. BMC Bioinformatics 11:94
Buzas EI et al (2017) Single particle analysis: Methods for detection of platelet extracellular vesicles in suspension (excluding flow cytometry). Platelets 28(3):249–255
Buzas EI et al (2014) Emerging role of extracellular vesicles in inflammatory diseases. Nat Rev Rheumatol 10(6):356–364
Caradec J et al (2014) Reproducibility and efficiency of serum-derived exosome extraction methods. Clin Biochem 47(13–14):1286–1292
Caraus I et al (2015) Detecting and overcoming systematic bias in high-throughput screening technologies: a comprehensive review of practical issues and methodological solutions. Brief Bioinform 16(6):974–986
Carr BWM (2013) Nanoparticle tracking analysis: a review of applications and usage 2010–2012." NanoSight Ltd.
Carr BWM (2014) Nanoparticle tracking analysis: a review of the first 1,000 reports of applications and usage of NTA. Malvern Instruments Ltd.
Chiang CY, Chen C (2019) Toward characterizing extracellular vesicles at a single-particle level. J Biomed Sci 26(1):9
Coumans FAW et al (2017) Methodological Guidelines to Study Extracellular Vesicles. Circ Res 120(10):1632–1648
Couvreur P et al (2006) Squalenoyl nanomedicines as potential therapeutics. Nano Lett 6(11):2544–2548
Dai Y et al (2021) Unbiased RNA-Seq-driven identification and validation of reference genes for quantitative RT-PCR analyses of pooled cancer exosomes. BMC Genomics 22(1):27
Dill KB, S, (2003) Molecular driving forces: statistical thermodynamics in chemistry and biology. Garland Science, New York
Dobrovolskaia MA, McNeil SE (2007) Immunological properties of engineered nanomaterials. Nat Nanotechnol 2(8):469–478
Draghici S (2002) Statistical intelligence: effective analysis of high-density microarray data. Drug Discov Today 7(11):S55-63
Dragovic RA et al (2011) Sizing and phenotyping of cellular vesicles using nanoparticle tracking analysis. Nanomedicine 7(6):780–788
Ferrari M (2005) Cancer nanotechnology: opportunities and challenges. Nat Rev Cancer 5(3):161–171
Filipe V et al (2010) Critical evaluation of Nanoparticle Tracking Analysis (NTA) by NanoSight for the measurement of nanoparticles and protein aggregates. Pharm Res 27(5):796–810
Foteini P et al (2019) Physicochemical study of the protein-liposome interactions: influence of liposome composition and concentration on protein binding. J Liposome Res 29(4):313–321
Geeurickx E et al (2019) The generation and use of recombinant extracellular vesicles as biological reference material. Nat Commun 10(1):3288
Gobbo J et al. (2016) Restoring Anticancer Immune Response by Targeting Tumor-Derived Exosomes With a HSP70 Peptide Aptamer. J Natl Cancer Inst 108(3).
Goetzl EJ et al (2020) Traumatic brain injury increases plasma astrocyte-derived exosome levels of neurotoxic complement proteins. Faseb J 34(2):3359–3366
Gorji-Bahri G et al. (2021) Validation of common reference genes stability in exosomal mRNA-isolated from liver and breast cancer cell lines. Cell Biol Int.
Gunasekaran PM et al. (2019) For what factors should we normalize urinary extracellular mRNA biomarkers? Biomol Detect Quantif 17:100090.
Harrison SC (2010) Virology. Looking inside adenovirus. Science 329(5995):1026–1027
Heath JR, Davis ME (2008) Nanotechnology and cancer. Annu Rev Med 59:251–265
Iyer AK et al (2006) Exploiting the enhanced permeability and retention effect for tumor targeting. Drug Discov Today 11(17–18):812–818
Kamerkar S et al (2017) Exosomes facilitate therapeutic targeting of oncogenic KRAS in pancreatic cancer. Nature 546(7659):498–503
Keerthikumar S et al (2016) ExoCarta: a web-based compendium of exosomal cargo. J Mol Biol 428(4):688–692
Konkoth A et al (2021) Multifaceted role of extracellular vesicles in atherosclerosis. Atherosclerosis 319:121–131
Koritzinsky EH et al (2019) Circadian variation in the release of small extracellular vesicles can be normalized by vesicle number or TSG101. Am J Physiol Renal Physiol 317(5):F1098–F1110
Koritzinsky EH et al (2017) Quantification of exosomes. J Cell Physiol 232(7):1587–1590
Krzywinski M, Altman N (2014) Points of significance: comparing samples-part I. Nat Methods 11(3):215–216
Livshits MA et al (2015) Isolation of exosomes by differential centrifugation: theoretical analysis of a commonly used protocol. Sci Rep 5:17319
Lombardo D et al (2019) Colloidal stability of liposomes. AIMS Materials Science 6(2):200–213
Malo N et al (2006) Statistical practice in high-throughput screening data analysis. Nat Biotechnol 24(2):167–175
Moghimi SM, Kissel T (2006) Particulate nanomedicines. Adv Drug Deliv Rev 58(14):1451–1455
Momen-Heravi F et al (2012) Alternative methods for characterization of extracellular vesicles. Front Physiol 3:354
O'Loghlen A (2018). Role for extracellular vesicles in the tumour microenvironment. Philos Trans R Soc Lond B Biol Sci 373(1737).
Otake K et al (2021) Quantitative comparison of the mRNA content of human iPSC-derived motor neurons and their extracellular vesicles. FEBS Open Bio 11(2):494–506
Pathan M et al (2017) A novel community driven software for functional enrichment analysis of extracellular vesicles data. J Extracell Vesicles 6(1):1321455
Pellequer Y et al (2003) Methodology for assaying recombinant interleukin-2 associated with liposomes by combined gel exclusion chromatography and fluorescence. J Chromatogr B Analyt Technol Biomed Life Sci 783(1):151–162
Quackenbush J (2001) Computational analysis of microarray data. Nat Rev Genet 2(6):418–427
Quackenbush J (2002) Microarray data normalization and transformation. Nat Genet 32(Suppl):496–501
Raposo G, Stoorvogel W (2013) Extracellular vesicles: exosomes, microvesicles, and friends. J Cell Biol 200(4):373–383
Ries E. (2011). "The Lean Startup: How Today's Entrepreneurs Use Continuous Innovation to Create Radically Successful Businesses. Currency: 336.
Robinson MD, Oshlack A (2010) A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol 11(3):R25
Rodrigues M et al (2019) Rapid lipid-based approach for normalization of quantum-dot-detected biomarker expression on extracellular vesicles in complex biological samples. Nano Lett 19(11):7623–7631
Seigneuric R et al (2016) Tumor exosomes: potential biomarkers and targets in Cancer. J Clin Cell Immunol 7:472
Seigneuric R, Garrido C (2016). "Tumor-derived extracellular vesicles: protocols, models and clinical evidence. Front Oncol
Seigneuric R, Garrido C (2016b) Tumor-derived extracellular vesicles: protocols, models, and clinical evidence. Front Oncol 6:230
Seigneuric R et al. (2011) Targeting cancer with peptide aptamers. Oncotarget.
Seigneuric R et al (2010) From Nanotechnology to Nanomedicine: applications to cancer research. Curr Mol Med 10(7):640–652
Serrano-Pertierra E et al. (2020) Extracellular Vesicles: Current Analytical Techniques for Detection and Quantification. Biomolecules 10(6).
Shtam T et al. (2020) Evaluation of immune and chemical precipitation methods for plasma exosome isolation. PLoS ONE 15(11): e0242732.
Simeone P et al. (2020) Diameters and Fluorescence Calibration for Extracellular Vesicle Analyses by Flow Cytometry. Int J Mol Sci 21(21).
Skog J et al (2008) Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat Cell Biol 10(12):1470–1476
Slonim DK (2002) From patterns to pathways: gene expression data analysis comes of age. Nat Genet 32(Suppl):502–508
Storey J (2002) A direct approach to false discovery rates. J r Statist Soc B 64(3):479–498
Storey JD, Tibshirani R (2003) Statistical significance for genomewide studies. Proc Natl Acad Sci USA 100(16):9440–9445
Streit M, Gehlenborg N (2014) Bar charts and box plots. Nat Methods 11(2):117
Thery C et al. (2006) Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Curr Protoc Cell Biol Chapter 3: Unit 3 22.
Thery C et al (2018) Minimal information for studies of extracellular vesicles 2018 (MISEV2018): a position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. J Extracell Vesicles 7(1):1535750
Thery C et al (2002) Exosomes: composition, biogenesis and function. Nat Rev Immunol 2(8):569–579
Torchilin VP (2005) Recent advances with liposomes as pharmaceutical carriers. Nat Rev Drug Discov 4(2):145–160
Torchilin VP (2010) Passive and active drug targeting: drug delivery to tumors as an example. Handb Exp Pharmacol(197): 3–53.
Valadi H et al (2007) Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol 9(6):654–659
Valkonen S et al (2017) Biological reference materials for extracellular vesicle studies. Eur J Pharm Sci 98:4–16
van der Pol E et al (2013) Innovation in detection of microparticles and exosomes. J Thromb Haemost 11(Suppl 1):36–45
Vestad B et al (2017) Size and concentration analyses of extracellular vesicles by nanoparticle tracking analysis: a variation study. J Extracell Vesicles 6(1):1344087
Wan Y et al (2018) Aptamer-conjugated extracellular nanovesicles for targeted drug delivery. Cancer Res 78(3):798–808
Welsh JA et al (2020) Towards defining reference materials for measuring extracellular vesicle refractive index, epitope abundance, size and concentration. J Extracell Vesicles 9(1):1816641
Whitesides GM (2003) The “right” size in nanobiotechnology. Nat Biotechnol 21(10):1161–1165
Wiles AM et al (2008) An analysis of normalization methods for Drosophila RNAi genomic screens and development of a robust validation scheme. J Biomol Screen 13(8):777–784
Yuan F et al (1995) Vascular permeability in a human tumor xenograft: molecular size dependence and cutoff size. Cancer Res 55(17):3752–3756
Zahran AM et al (2019) Circulating Microparticles in Children With Sickle Cell Anemia in a Tertiary Center in Upper Egypt. Clin Appl Thromb Hemost 25:1076029619828839
Acknowledgements
RS thanks S. de Almeida for technical assistance and discussions and A. Bourredjem for discussions.
Funding
This work was supported by the Région Bourgogne Franche-Comté PARI (grant number 9201AAO050S01716), Ligue contre le Cancer (grant number R18032MM) and Nano2Bio and FEDER (Grant Number BG0005900). No funding sources were involved in the study design, collection, analysis, interpretation of data, writing or in the decision to submit the manuscript for publication.
Author information
Authors and Affiliations
Contributions
YP, GZa, JMR, IS, MCC, GZi, FN and RS contributed to the design of experiments. YP synthetized liposomes. All authors (YP, GZa, JMR, IS, MCC, GZi, FN and RS) contributed to data analysis and manuscript preparation. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflicts of interest/Competing interests
All authors hereby declare that there is no conflict of interest.
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
All authors agreed to publish this version of the article.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Pellequer, Y., Zanetta, G., Rebibou, JM. et al. Development of a new methodology to determine size differences of nanoparticles with nanoparticle tracking analysis. Appl Nanosci 11, 2129–2141 (2021). https://doi.org/10.1007/s13204-021-01932-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13204-021-01932-2