Abstract
This work was based on the study of a defense-related protein involved in actinorhizal symbiosis established between Casuarina glauca and a nitrogen fixing bacterium from the Frankia genus, namely a class III chitinase (CgCHI3). In order to understand its role during the symbiotic process, recombinant CgCHI3 was overexpressed in Escherichia coli and subsequently characterized. In this context, the recombinant protein showed endochitinase, β-1,3-glucanase, β-1,4 glucanase and lysozyme activity, as well as the capacity to inhibit the growth of non-symbiotic bacteria (E. coli strains BL21 and K12, Paracoccus denitrificans and Bacillus subtilis). It exhibited no antifungal activity against Colletotrichum gloeosporioides, Botrytis cinerea, Trichoderma viride and Fusarium oxysporum and did not affect the growth of the symbiotic bacteria Frankia and Rhizobium, neither the performance of Frankia nodulation factors. These results suggest that recombinant CgCHI3 is a multifunctional protein involved in the infection process, specifically in the lysis of the cell wall during intracellular penetration, or in the formation of the infection thread, or any other modification of the cells in order to accommodate the symbiotic bacteria.
Similar content being viewed by others
1 Introduction
The superfamily of glycoside hydrolases (also named glycosyl hydrolases or glycosidases) is composed of 81 enzyme families that hydrolyze a wide range of substrates (Lombard et al. 2014). Among them, chitinases (EC 3.2.1.14) catalyze the hydrolytic cleavage of β-1,4 glycosidic bonds present in chitin, an insoluble derivative of glucose with sizes ranging from 20 to 90 kDa, and the second most abundant natural polysaccharide on earth after cellulose (Bhattacharya et al. 2007). Aside from being the major component of the exoskeleton of arthropods and nematodes, chitin is also the major component of the cell wall of bacteria, algae and higher fungi such as Basidiomycetes, Ascomycetes and Deuteromycetes. Up to 60 % of the cell wall of these organisms can be made of chitin (Bhattacharya et al. 2007).
Although chitin is not present in virus, plants and mammals, chitinolytic activity was found in these organisms. Since many chitinases are induced in response to pathogen infection, it is believed that they are mainly involved in the defense against fungi containing chitin in their cell wall. The lysozyme activity, which in some cases is observed together with chitinase activity, has been interpreted as a defense mechanism of the plant against possible bacterial attacks.
Plants have different classes of chitinases which can be distinguished by their primary structure, the catalytic mechanism and the sensitivity to inhibitors (Kasprzewska 2003; Taira 2010; Kesari et al. 2015; Islam and Datta 2015). Individually, all classes of chitinases exhibit different substrate specificities and reaction mechanisms. For example, in the case of class III chitinases from tobacco, proteins show a considerable degree of activity of lysozyme as well as chitinolytic activity, while class VI has only chitinase activity (Brunner et al. 1998).
Plant chitinases are present in all organs and tissues, both in the vacuole and apoplast (Kasprzewska 2003). The involvement of chitinase as a mechanism of active or passive defense against pathogens has been reported and verified experimentally. In general, rapid induction and accumulation of chitinases correlates with resistance to attack by pathogens (Samac and Shah 1991; Meier et al. 1993; Gerhardt et al. 1997). However, chitinases may also be involved in plant developmental processes such as cell elongation (Zhong et al. 2002) embryogenesis (Fukamizo 2000), cold tolerance, acting as antifreeze proteins (Yeh et al. 2000; de los Reyes et al. 2001; Yu and Griffith 2001) and storage (Peumans et al. 2002). In the case of symbioses with nitrogen fixing bacteria, it is thought that chitinases are involved in the auto regulation mechanism controlling the number and formation of nodules (Vasse et al. 1993; Staehelin et al. 1995; Goormachtig et al. 1998; Kim and An 2002; Fortunato et al. 2007). Moreover, they were identified as part of the nodule protection machinery against pathogens (Staehelin et al. 2000; Salzer et al. 2000; Kim et al. 2005; Santos et al. 2010; Ribeiro et al. 2011).
In a previous study CgChi3, the gene encoding a class III chitinase, has been isolated from Casuarina glauca root nodules induced by nitrogen-fixing bacteria Frankia (Fortunato et al. 2007). The transcriptional activity of this gene is specifically associated with the symbiotic process, particularly with bacterial nodule infection (Fortunato 2006; Fortunato et al. 2007). The fact that CgChi3 is expressed specifically in nodules makes it a promising symbiotic marker candidate for the elucidation of the role of defense-related genes in symbiosis as well as for the understanding of the mechanisms of evolution of symbiosis as a reminiscent of pathogenesis. In the present manuscript, we describe the cloning, overexpression, purification of the recombinant protein class III chitinase from C. glauca and its functional characterization in order to contribute to the clarification of its role during symbiosis between C. glauca and Frankia Thr strain.
2 Materials and methods
2.1 Biological material
Casuarina (C.) glauca plants were grown hydroponically from seeds provided by B & T World Seeds, Paguignan, 34210 Aigues-Vives, France (http://b-and-t-world-seeds.com) as described by Zhong et al. (2010). For C. glauca inoculation, Frankia Thr strain was used as described in Fortunato et al. (2007).
2.2 Bacterial strains, plasmid and fungal cultures
Cloning of the coding region (ORF-Open Reading Frame) of CgChi3 gene was done into the pJET1.2/blunt vector (Fermentas, Lithuania) and the subcloning into the pET-28b vector (Novagen, Germany). Escherichia coli DH5α and BL21 (DE3) (Novagen, Germany) cells with expression plasmids were grown aerobically in Luria Bertani (LB) broth or on LB agar plates at 37 °C, supplemented with kanamycin (30 μg/mL) for selection of transformants. Phytopathogenic fungal species used in this study included Botrytis cinerea (gray mold of Jatropha curcas), Colletotrichum gloeosporioides (Anthracnose of Manihot esculenta), Fusarium oxysporum (Solanum lycopersicum) and Trichoderma viride (Solanum lycopersicum). Ectomycorrhizal species included Suillus bovinus, Amanita muscari, Hebeloma crustiliforme and endomycorrhizal species included Glomus intraradices. Phytopathogenic and ectomycorrhizal fungi were grown on Potato Dextrose Agar medium (PDA). Endomycorrhizal fungus was grown on Minimal Salts Medium (MSM). Bacterial strains included, symbiotic- (Frankia Thr, ACN14a and CCI3 and Sinorhizobium medicae 55 Mp) and non-symbiotic (E. coli K12 and BL21 (DE3), Paracoccus denitrificans and Bacillus subtilis 168 wild type) bacteria. Frankia Thr culture was grown in Bacterial Alkaline Phosphatase (BAP-PCM) medium at 26 °C in the dark as described by Schwencke (1991) and Sinorhizobium was grown in yeast extract mannitol agar medium.
2.3 Construction of CgCHI3 expression system
Based on the cDNA full-size sequence, gene-specific primers covering the coding sequence of C. glauca chitinase III gene (CgChi3, GenBank accession number EF134410) were designed and used to amplify the corresponding cDNA-ORF by PCR in an iQ5 thermalcycler (BIO-RAD, USA).
For subcloning, specific primers were designed for the same region containing the NdeI and SalI restriction sites (underlined sequences): forward primer (CgChi3 fwd) 5’–CCATCATATGGCATTTTGTACAACTCTTCC–3’ and reverse primer (CgChi3 rev) 5’–CCATGTCGACGACATGGCCCTTAATGG–3’. PCR mix (50 μL) contained 50 ng of primer, 0.45 μg of template DNA, 2.5 μL 10× BD Advantage 2 SA PCR buffer, 2.5 μL 50× BD Advantage 2 polymerase (Clontech, San Jose, CA) mix, and 10 mM dNTPs. The amplification program included an initial denaturation step (94 °C for 30 sec) followed by 35 cycles of 30 sec at 94 °C, 30 sec at 55 °C and 30 sec at 72 °C, followed by an extension step at 72 °C for 5 min. The amplified fragments were purified using the High Pure PCR Product Purification Kit (Roche Applied Science, Germany) and ligated into the cloning vector pJET1.2/blunt, using the CloneJETTM PCR Cloning Kit (Fermentas, Lithuania), according to the manufacturer’s instructions. The ligation product was used to transform competent E. coli DH5α cells, plated on LB agar medium containing kanamycin (30 μg/mL) and incubated at 37 °C overnight.
Selected clone (pJET1.2-Cgchi3-ORF) was used to inoculate 5 mL of LB cultures containing kanamycin (30 μg/mL), grown at 37 °C for 18 h at 220 rpm orbital shaking for isolation of plasmid DNA using the Illustra PlasmidPrep Midi Flow kit (GE Healthcare Life Science, Germany). The plasmids, pJET1.2-Cgchi3-ORF and pET-28b, were subsequently quantified and hydrolyzed with the restriction endonucleases NdeI and SalI (Fermentas, Lithuania) according to instructions of the manufacturer. The fragment corresponding to CgChi3-ORF gene was purified from agarose gel using the High Pure PCR Product Purification kit (Roche Diagnostics GmbH, Germany) and quantified spectrophotometrically.
For the construction of the overexpression vector the purified fragment was ligated to the linearized expression vector in a 3:1 (insert:vector) ratio using the enzyme T4 DNA ligase (Fermentas, Lithuania), following the instructions of the manufacturer. The ligation product was used to transform competent E. coli DH5α cells as described above. Selection of positive clones was performed by colony-PCR using the amplification conditions described before. Plasmid DNA was isolated and sequenced using specific primers for the pET-28b plasmid, in order to confirm the cloned sequence.
2.4 Recombinant protein overexpression in Escherichia coli BL21 (DE3) cells and purification
One of the positive clones (pET28b-CgCHI3-ORF) was used to transform E. coli BL21 (DE3) competent cells by heat shock following the protocol of the manufacturer.
For overexpression of recombinant protein, a 5 mL LB medium culture containing kanamycin was inoculated with one E. coli BL21 (DE3) transformant and grown for 8 h at 37 °C, at 210 rpm. The pre-inoculum was diluted 100× in 50 mL of LB-kanamycin medium and incubated for 16 h under the same conditions. The overnight culture was diluted 100× in 1 L of LB medium containing antibiotic and incubated at 37 °C with stirring at 210 rpm until OD600 nm between 0.5 and 0.6. At this time, protein expression was induced with 0.1 mM of isopropyl β-D-1-thiogalactopyranoside (IPTG). After 3 h the culture was centrifuged at 5,700 g for 30 min at 4 °C to collect overexpression cells. The pellet was suspended in 10 mM Tris–HCl pH 7.6. The overexpression of recombinant protein was assessed by SDS-PAGE (12.5 %) as described by Laemmli (1970) in a vertical system (BIO-RAD, USA) at a constant current of 120 V for 1 h 30 min.
After three freeze-thaw cycles of the above obtained suspension, DNase I from bovine pancreas (Sigma, Spain) was added to the cell suspension before cell disruption in a French Pressure Cell Press (Thermo Electron Corporation, USA) at a pressure of 1,700 psi. After the addition of 10 mM of benzamidine hydrochloride (Sigma, Spain) and 1 mM of phenylmethanesulfonyl fluoride (Sigma, Spain), the mixture was centrifuged at 5,700 g for 20 min at 4 °C. The resulting cellular suspension was then ultracentrifuged (Beckman Optima LE-80 K com rotor 70 Ti, USA) at 225,072 g for 1 h at 4 °C, in order to separate the soluble fraction from the membrane fraction. Pellets were resuspended in lysis buffer (50 mM Tris–HCl, pH 7.8, 1 mM EDTA) and frozen at −80 °C.
As CgCHI3 recombinant protein was found to be in the pellet from the low speed centrifugation, solubilization from inclusion bodies was done using the procedure described by (Kirubakaran and Sakthivel 2007) with a slight modification. The addition of β-mercaptoethanol (2.5 mM) was replaced by a redox buffer composed of GSH (3 mM / 0.3 mM GSSG) (Sigma, Spain). Membrane fraction was centrifuged at 13,500 g for 10 min at 4 °C and the pellet containing the inclusion bodies was washed in three volumes of wash buffer (10 mM Tris–HCl, pH 7.5, 300 mM NaCl, 1 mM EDTA, 1 % Triton X-100 and 1 M urea), kept at room temperature for 5 min, and centrifuged as above. The detergent was removed by ressuspending and washing the pellet using sterile distilled water 5 times and centrifuged as above. For protein renaturation, the pellet was suspended in a 10× volume of 50 mM Tris–HCl, pH 7.8, 5 mM MgCl2, 0.3 mM GSSG, 3 mM GSH, 0.1 % (w/v) Tween 20 (denaturation–renaturation buffer; DR buffer) containing 8 M urea and incubated at room temperature for 1 h. The mixture was centrifuged at 10,000 g for 20 min at 4 °C. The supernatant was dialyzed gradually overnight against DR buffer containing progressively lower concentrations of urea down to 0 mM.
The presence of the protein on the soluble fraction was analyzed on a 12.5 % denaturing polyacrylamide gel, followed by Western Blotting by the positive reaction with an antibody against Class III chitinase of Lupinus albus (Regalado et al. 2000) and a monoclonal antibody against histidine (Sigma, Spain) that recognizes the affinity tag of the recombinant protein. After electrophoresis, proteins were electrotransferred to a PVDF membrane (GE Healthcare Life Sciences, Germany) following the protocol described by Regalado and Ricardo 1996).
2.5 Purification from inclusion bodies
Purification of the recombinant protein was performed on ÄKTA Prime Plus Liquid Chromatography system (GE Healthcare Life Science, Germany) at 4 °C. The solubilized and refolded protein solution was centrifuged at 5,850 g for 20 min at 4 °C. The supernatant was filtered and applied into a 1 mL HisTrap FF ™ column (GE Healthcare Life Science) equilibrated with buffer A composed by 50 mM Tris–HCl pH 7.8, 5 mM MgCl2, 2.5 mM β-mercaptoethanol, 0.1 % (w/v) Tween 20, 20 mM imidazol. After washing the column with buffer A, adsorbed proteins were eluted with buffer B (50 mM Tris–HCl pH 7.8, 5 mM MgCl2, 2.5 mM β-mercaptoetanol, 0.1 % (w/v) Tween 20, 500 mM imidazol) and collected in 1 mL fractions at a flow rate of 1 mL/min.
Protein purity was assessed by SDS-PAGE (12.5 %). The concentration of dialyzed protein was determined by the bicinchoninic acid (BCA) assay method (Sigma, Spain).
2.6 Functional analysis of recombinant protein
2.6.1 Enzymatic activities assays
The CgCHI3 chitinolytic activity was tested using the polymeric substrate, carboxymethyl chitin Remazol Brilliant Violet (CM-chitin-RBV, Loewe Biochemica, Germany), a soluble derivative of chitin, as described by Wirth and Wolf (1992). The enzyme, 0.1 mL of enzyme at a concentration of 1 μg/mL, was pre-incubated at 37 °C with 0.2 mL of 0.2 M sodium acetate buffer pH 5.4. The reaction was started by the addition of 0.1 mL of CM-chitin-RBV (2 mg/mL) and after specific reaction times, stopped by the addition of 0.1 mL of 2 N HCl. The non-degraded material was precipitated on ice for 10 min and then centrifuged at 10,000 g for 5 min. The absorbance of the supernatant was measured in a Evolution 300 UV–vis Spectrophotomer (Thermo Scientific, USA) at 550 nm against water. Three independent measurements were performed and average values used. The specific activity was defined as U/mg, with activity unit (U) as the ΔAbs550 nm/min, calculated as the initial velocity in accordance with an exponential first order increase. The exochitinase activity was assessed using the chitinase assay kit from Sigma that tests the chitobiosidase and β-N-acetylglucosaminidase enzymatic activities using 4-nitrophenyl N,N’-diacetyl-β-D-chitobioside and 4-nitrophenyl N-acetyl-β-D-glucosaminide, respectively, as substrates.
Lysozyme activity of the purified enzyme was measured by the rate of lysis of Micrococcus lysodeikticus cell walls (Sigma, Germany) as described by Shugar (1952). The reaction contained 2.9 mL of cell suspension (9 mg of dried M. lysodeikticus cells walls in 30 mL of 0.1 M potassium phosphate buffer, pH 7.0) and 0.1 mL of enzyme solution at different concentrations and monitored continuously as decrease in light scattering at 450 nm for 30 min at 25 °C in a Evolution 300 UV–vis Spectrophotomer (Thermo Scientific, USA). One unit of enzymatic activity was considered equal to a decrease in turbidity of 0.001 per minute at 450 nm.
β-1,3-glucanase activity was determined using the polymeric substrate, carboxymethyl-curdlan chitin Remazol Brilliant Blue (CM-Curdlan-RBB, Loewe Biochemica, Germany) a soluble derivative of chitin, as described by Wirth and Wolf (1992). The enzymatic assay was identical to the chitinolytic test, but using a stock solution of substrate at 4 mg/mL. The increase of optical density was measured at 600 nm against a blank assay prepared without enzyme solution. Three independent measurements were performed and average values used. The activity as for the chitinase one, with activity unit (U) as the ΔAbs600 nm/min.
β-1,4-endoglucanase (cellulase) activity was analyzed using sodium salts of carboxymethyl cellulose (CMC) in 0.2 % (w/v) 50 mM potassium phosphate-citric acid (PCA) pH 5.2 as enzyme substrate, according to Chen et al. (2004): 0.2 mL of substrate was mixed with 0.2 mL of enzyme solution (0.54 mg/mL) and 0.4 mL of PCA buffer and incubated at 30 °C for different reaction times. To measure the production of reducing sugar units, 0.8 mL of distilled water and 1 mL of BCA reagent were added to 0.2 mL of the previous reaction mixture at 100 °C for 15 min (Waffenschmidt and Jaenicke 1987). Samples were then cooled at room temperature for 10 min and absorbance was measured at 570 nm.
2.6.2 Antifungal activity
The disk diffusion method was used to study the antifungal activity (Elhamid et al. 2010). The assay was carried out in 9 cm diameter Petri plates containing PDA medium. The mycelium of several phytopathogenic filamentous fungi, Botrytis cinerea, Ccirolletotrichum gloeosporioides, Furasium oxysporum and Trichoderma viride were inoculated at the center of the plates; when the colony diameter reached 3–4 cm, different amounts of chitinase protein (0.01 mg and 0.0025 mg) were applied directly into the Petri plate or in a filter paper followed by incubation at 25 ± 2 °C. After 3 days, the growth was observed. Positive controls with water and DR buffer was used and all preparations of enzyme, buffer and water were sterilized by filtration. Each assay was performed in triplicate.
2.6.3 Antibacterial activity
The same approach described above was performed. In the present assay 3 species of pathogenic bacterial strains were used: E. coli K12 and BL21 (DE3), Paracoccus denitrificans and Bacillus subtilis 168 wild type. Each bacterial strain was grown in 5 mL LB for 16 h at 37 °C with vigorous agitation. 0.1 mL of the cell cultures were spread onto LB agar plates and different amounts of chitinase protein (0.01 mg and 0.0025 mg) were applied directly in the plate or in filter paper followed by incubation at 37 °C; growth was observed after 24 h.
2.6.4 CgCHI3 effect on Frankia and rhizobium growth
100 μL of 0.1 M sodium acetate pH 5.4 and 100 μL of pure enzyme (0.1 mg/mL) were added to 800 μL of the supernatant cultures of the two Frankia strains ACN14a and CcI3. As a negative control we used 400 μL Bacterial Alkaline Phosphatase (BAP) buffer, 50 μL of 0.1 M sodium acetate pH 5.4 and 50 μL of pure enzyme; as a positive control we used the supernatant of the culture medium of each bacterium. Reactions were incubated at 37 °C for 1 h and 24 h, respectively. Enzyme was inactivated for 10 min at 70 °C, 1:10 dilutions were made in all assays at a final volume of 1 mL and plant roots were inoculated with 100 mL of the reaction. Observation of results was performed daily under the microscope in order to verify changes in roots morphology (Tisa et al. 2013).
3 Results and discussion
In this study, a Casuarina glauca gene (GenBank: EF134410) encoding chitinase (CgChi3) was overexpressed in E. coli. Recombinant CgCHI3 produced has a His-tagged protein with theoretical biochemical parameters, determined by ExpasyParamTool tool (http://web.expasy.org/protparam/), of 31.9 kDa of molecular mass and a pI of 6.8. Although slightly high for an acidic protein, the pI value is similar to the ones for proteins isolated from Cornus florida (Cardwell and McDaniel 1998) and green beans (Ye and Ng 2005). Proteins of higher plants and algae typically exhibit pI values between 3.0 and 10 (Jollès and Muzzarelli 1999).
Amino acid sequence analysis revealed that CgCHi3 is a class III chitinase belonging to family 18 (Figure S1 Supplementary material shows a comparative analysis of the primary structure of CHI3). In addition, and based on its amino acid sequence it was possible to classify CgCHI3 as a protein from the PR-8 family of the pathogenesis-related (PR) proteins group. Mauch and Staehelin proposed a model for the defense mechanism of plants that involves the synergistic action of β-1,3-glucanases (that are members of the PR-2 family) and chitinases, in which the first would act in an early stage of infection, releasing β-1,3-glucans from fungal hyphae which then activate the expression of defense gene, in particular those encoding chitinases that would completely degrade the cell walls of pathogens leading to lysis (Mauch and Staehelin 1989).
3.1 Cloning and overexpression
The production of recombinant CgCHI3 using the overexpression plasmid pET28b-CgCHI3-ORF in E. coli BL21 (DE3) cells was achieved at 15 °C in LB medium with kanamycin (30 μg/mL) after induction of gene expression with 0.1 mM IPTG for 3 h. Cellular fractionation showed that the protein was predominantly produced as inclusion bodies. Attempts to produce the protein in the soluble form was unsuccessful. Protein aggregates were denatured with 8 M urea and renaturated slowly removing the denaturing agent by dialysis. Purity of protein preparations obtained by solubilization and refolding of inclusion bodies was verified by SDS-PAGE (Fig. 1). Analysis of gels revealed a high intensity band around 32 kDa (32.5 ± 2.2 kDa), characteristic of chitinases. Proteins from higher plants and algae have molecular masses around 30 kDa, while in mollusks, vertebrates and some fungi and bacteria may vary between 40 and 120 kDa (Jollès and Muzzarelli 1999). Besides the major band at ∼32 kDa, SDS-PAGE gels of all protein preparations also showed the presence of minor bands with higher molecular masses that were attributed to stable oligomeric forms of the protein since purification of the refolded protein by affinity chromatography using a HisTrap FF column did produce protein preparations with the same degree of purity and identical electrophoretic pattern.
SDS-PAGE analysis of recombinant C. glauca chitinase obtained from solubilization of inclusion bodies. M – Low Molecular Weight Marker (LMW NZYTech, Portugal); 1 – Recombinant CgCHI3 containing solution obtained after renaturation of inclusion bodies. The box indicates the position of the band corresponding to chitinase
3.2 Functional analysis of recombinant protein
In order to evaluate the role of CgCHI3 in the symbiotic process with Frankia different catalytic activities were tested.
3.2.1 Enzymatic activities assays
CgCHI3 showed chitinolytic activity for the substrate CM-Chitin-RBV at pH 5.4 and 37 ° C, with a specific activity of 23.5 ± 4.2 U/mg (Fig. 2a), a value of the same order of magnitude of those reported for chitinases from barley seeds, recombinant and native, which have a specific activity of 12 ± 0.7 U/mg for the same substrate (Andersen et al. 1997). Moreover, CgCHI3 has a kinetic profile similar to the one obtained by Saborowski et al. (1993) in the optimization of the activity assay used to assess chitinase activity. In fact, the change in absorbance at 550 nm as a function of reaction time is linear up to 0.5 absorbance units at a rate of 7.3 × 10−3 U compared with 4.8 × 10−3 U for CgCHI3. The activity of endochitinase was confirmed when testing the exochitinase activity of CgCHI3 since the values of specific activities obtained for the later were always less than 1 % of the value obtained for the positive control from the Sigma kit (Tronsmo and Harman 1993; Duo-Chuan et al. 2005).
Enzymatic activities assays of CgCHI3. a Chitinase activity using CM-Chitin-RVB as substrate at pH 5.4 and 37 °C; b Lysozyme activity using a Micrococcus lysodeikticus cell culture at pH 7.0, 25 °C; c β-1,3-glucanase activity using CM-curdlan-RBB as substrate at pH 5.4 and 37 °C; d β-1,4-glucanase activity (carboxymethyl cellulase) using carboxymethyl cellulose at pH 5.2, 30 °C. Each experimental point represents the mean of three replicates
CgCHI3 also exhibited lysozyme activity with a specific activity equal to 115 ± 30 U/mg, which is a value similar to that described in the literature for homologous enzymes, in particular for latex wild-cotton chitinase (Ipomoea carnea) that has a specific activity between 69 and 80 U/mg (Patel et al. 2009; 2010). The dependence of the rate of lysis of Micrococcus lysodeikticus cells with CgCHI3 concentration is represented in Fig. 2b. Many endochitinases belonging to family 18, from plants and microorganisms such as Pseudomonas aeruginosa, Choanephora cucurbinatum and Phascolomyces articulosus, are described as bifunctional enzymes exhibiting lysozyme and chitinolytic activities (Wang et al. 2005; Jekbl et al. 1991; Bokma et al. 2002). Lysozymes and chitinases represent an important group of enzymes that hydrolyze polysaccharides. While chitinase acts on chitin, a linear polymer of N-acetylglucosamine linked by β-1,4 bonds present in the cell walls of insects, crustaceans, and fungi, lysozyme hydrolyzes the peptidoglycan of bacterial cell walls composed of alternating residues of N-acetylglucosamine and N-acetylmuramic acid linked by β-1,4 bonds. The chemical similarities between the two substrates explain the lysozyme activity of some chitinases. Despite this bifunctionality, the two proteins have amino acid sequences with low homology (Wohlkönig et al. 2010).
The ability of CgCHI3 to hydrolyze the substrate CM-curdlan-RBB and thus infer about its β-1,3-endoglucanase activity was evaluated from Fig. 2c resulting in the determination of a specific activity of 0.75 ± 0.2 U/mg. To our knowledge this is the first chitinase that shows this type of activity. β-1,3-glucanases are defense proteins from the PR-2 family, being the most abundant glucanases in plant cell walls. They hydrolyze β-1,3 glycosidic bonds of glucans present in the cell wall of most fungi, and are divided into three classes (I-III). Class I glucanases are basic proteins of approximately 33 kDa and are located in the vacuoles; classes II and III include acidic and extracellular proteins with molecular masses approximately equal to 36 kDa. Plant β-1,3-glucanases are induced in response to pathogenic infection and other stress factors such as salicylic acid, abscisic acid, wound or drought are also involved in different physiological processes, such as sporogenesis, fertilization, maturation or germination (Sinha et al. 2014). In addition to these functions, numerous studies have demonstrated in vitro antifungal capacity of glucanases belonging to classes I and II (Theis and Stahl 2004).
The activity of CgCHI3 towards the substrate carboxymethyl cellulose to evaluate β-1,4-endoglucanase (cellulase) activity was also tested. A specific activity of 73 ± 18 U/mg was determined (Fig. 2d). Cellulose is the main component of plant cell walls, constituting the most abundant polysaccharide on earth and a major source of carbon in the biosphere. It is a linear polymer of D-glucose residues linked by β-1,4 bonds. This polysaccharide can be degraded by the synergistic action of three classes of cellulases, including β-1,4-endoglucanases, cellobiohydrolases and β-glucosidase. Again, this is the first report of a chitinase with β-1,4-glucanase activity. The fact that an enzyme, like CgCHI3, presenting activity of chitinase, glucanase, and cellulase is unique and may correspond to a more effective defense against pathogens or in an optimized system involved in the symbiosis process. Enzymatic plasticity is often associated with promiscuity of enzymes, despite their specificity. The ability to recognize and metabolize various substrates may represent an evolution towards a better catalytic efficiency.
3.2.2 Antifungal activity
The importance of chitinases as components of a plant defense mechanism has been the subject of several studies (Graham and Sticklen 1994; Carstens et al. 2003). Chitin, the natural substrate of chitinase, was not found in plants, but is synthesized by organisms associated with plants, in particular fungi. As mentioned earlier, analysis of the amino acid sequence indicated that CgCHI3 belongs to family 18, class III of acidic endochitinases. In fact, as discussed above, the recombinant protein described in this work presents chitinolytic activity. Therefore, we have further evaluated its antifungal potential. No inhibition of hyphae growth was observed in any of the phytopathogenic fungi tested (data not shown). The lack of antifungal activity was also observed in the acidic chitinase class III from tamarind seeds (Rao and Gowda 2008), the acidic chitinase from grain-pea (Cicer arietinumm L.) (Vogelsang and Barz 1993) and in four enzymes isolated from Streptomyces coelicolor (Kawase et al. 2006). Although it would seem obvious to associate the antifungal activity of chitinases to chitin hydrolysis, which would lead to the weakening of cell walls and subsequent lysis of pathogen cells, the inhibition of fungal growth mechanism cannot be explained solely by the chitinolytic activity since, as mentioned above, there are various chitinases that hydrolyze chitin but have no activity against fungi. Furthermore, it must be considered that CgCHI3 is a multifunctional enzyme whose expression is closely associated with the symbiotic processes (Fortunato et al. 2007; Tromas et al. 2012). The results seem to confirm these observations.
3.2.3 Antibacterial activity
Some chitinases have higher activity for bacterial peptidoglycans than for chitin, and thus are also classified as lysozymes (Brunner et al. 1998). Lysozymes are defined as enzymes with specific hydrolytic activity directed against peptidoglycan, components from bacterial cell wall (murein) in both Gram-negative and Gram-positive bacteria. The in vitro bacteriolytic activity from different lysozymes in a wide variety of organisms is well known (During 1993). Furthermore, lysozymes have been detected in various plant species (Howard and Glazer 1967; Glazer et al. 1969; Bernasconi et al. 1987; Audy et al. 1988; Kombrink et al. 1988; Trudel et al. 1989; Stintzi et al. 1993). Thus, the antimicrobial activity of CgCHI3 was evaluated. Bacterial growth inhibition was observed, demonstrating that bacterial species are sensitive to this enzyme (Fig. 3 and Figure S2 of Supplementary material). These results are in line with the in-suspension lysozyme activity of CgCHI3 demonstrated above (Fig. 2b).
Antibacterial activity of recombinant CgCHI3. Plates containing LB agar inoculated with the bacterial species in the presence of CgCHI3 at 37 °C. A representative assay with E. coli K12 cells is presented. a – Pure enzyme; b – Diluted enzyme 1:4; c and d – Positive growth controls in which the enzyme solution was replaced by buffer and water, respectively
3.2.4 CgCHI3 effect on Frankia and Rhizobium growth
In the legume-rhizobia symbiosis, chitinases are involved in the degradation of the chitin backbone of nodulation (Nod) factors (Goormachtig et al. 1998). Since CgCHI3 is an early noduline, specific from nodules (Fortunato et al. 2007), we tested the possibility of its involvement in the interaction with symbiotic bacteria Sinorhizobium medicae 55 Mp and Frankia ACN14a and CCI3. Contrary to legume chitinases, the presence of CgCHI3 did not affect the growth of symbiotic bacteria (data not shown). Likewise, the pre-treatment of Frankia Nod factors with CgCHI3 produced no changes in the early stages of infection, i.e., adhesion and curling of Alnus glutinosa root hairs. This observation suggests that CgCHI3 does not interact directly with the Frankia Nod factors. In fact, unlike rhizobia Nod factors, the presence of chitin backbones in Frankia factors has not been reported.
3.3 Discussion of the function of CgCHI3 in symbiosis
According to Fortunato et al. 2007, CgChi3 gene has specific transcriptional activity in nodules, mostly in the infection zone of the nodule. Based on these observations, the authors pointed out three possible roles for the product of this gene during the symbiotic process: i) recognition of the bacteria; ii) infection process; and/or iii) nodule formation. For the above mentioned reasons, the involvement of CgCHI3 in bacterial recognition process appears to be unlikely. In turn, the location of CgCHI3 in the apoplast in conjunction with the cellulase and glucanase activities, suggest that this enzyme could be involved in the intracellular penetration process of bacteria participating in structural changes, such as digestion of cell wall (cellulase) and/or formation of the infection thread involving the mycrosymbiont (glucanase). In fact, several authors have indicated the involvement of β-1,4-glucanases (cellulases) on the metabolism of the cell wall of higher plants, including cellulose biosynthesis and degradation and modification of other wall polysaccharides containing β-1,4 glucosyl residues (Buchanan et al. 2012). In the case of β-1,3-glucanase activity, it may be inferred that CgCHI3 may be involved in cell wall remodeling cleaving the β-1,3 glucan bonds of callose which is deposited in response to Frankia infection (Nowicki et al. 2012). Although the obtained results do not allow speculating on the involvement of CgCHI3 in nodule formation process, this hypothesis cannot be excluded.
Based on the obtained results, CgCHI3 proved to be a multifunctional enzyme, with chitinolytic, lysozyme, glucanase and cellulase activities. To our knowledge this is the first report of a chitinase with β-1,3 and β-1,4-glucanase activity. Besides its importance in the symbiotic process, this protein may also be appreciated in view of its potential biotechnological applications, for example in the paper, textile, beverages, feed or detergents industry. Additionally it might have interest for the ethanol production from plant waste.
References
Andersen MD, Jensen A, Robertus JD, Leaha R, Skriver K (1997) Heterologous expression and characterization of wild-type and mutant forms of a 26 kDa endochitinase from barley (Hordeum vulgare L.). Biochem J 322:815–822
Audy P, Benhamou N, Trudel J, Asselin A (1988) Immunocytochemical localization of a wheat germ lysozyme in wheat embryo and coleoptile cells and cytochemical study of its interaction with the cell wall. Plant Physiol 88:1317–1322
Bernasconi P, Locher R, Pilet PE, Jollès J, Jollès P (1987) Purification and N-terminal amino-acid sequence of a basic lysozyme from Parthenocissus quinquifolia cultured in vitro. Biochim Biophys Acta 915:254–260
Bhattacharya D, Nagpure A, Gupta RK (2007) Bacterial chitinases: properties and potential. Crit Rev Biotechnol 27:21–28
Bokma E, Rozeboom HJ, Sibbald M, Dijkstra BW, Beintema JJ (2002) Expression and characterization of active site mutants of hevamine, a chitinase from the rubber tree Hevea brasiliensis. Eur J Biochem 269:893–901
Brunner F, Stintzi A, Friting B, Legrand M (1998) Substrate specificities of tobacco chitinases. Plant J 14:225–234
Buchanan M, Burton RA, Dhugga KS, Rafalski AJ, Tingey SV, Shirley NJ, Fincher GB (2012) Endo-(1,4)-β-glucanase gene families in the grasses: temporal and spatial cotranscription of orthologous genes. BMC Plant Biol 12:235
Cardwell NA, McDaniel GL (1998) Comparison of chitinases from dogwood anthracnose resistant and susceptible Cornus species. HortSci 33:298–301
Carstens M, Vivier MA, Pretorius IS (2003) The Saccharomyces cerevisiae chitinase, encoded by the CTS1-2 gene, confers antifungal activity against Botrytis cinerea to transgenic tobacco. Transgenic Res 12:497–508
Chen PJ, Wei TC, Chang YT, Lin LP (2004) Purification and characterization of carboxymethyl cellulase from Sinorhizobium fredii. Bot Bull Acad Sinica 45:111–118
de los Reyes BG, Taliaferro CM, Anderson MP, Melcher U, McMaugh S (2001) Induced expression of the class II chitinase gene during cold acclimation and dehydration of bermudagrass (Cynodon sp.). Theor Appl Genet 103:297–306
Duo-Chuan LI et al (2005) Purification and partial characterization of two chitinases from the mycoparasitic fungus Talaromyces flavus. Mycopathologia 159:223–229
During K (1993) Can lysozymes mediate antibacterial resistance in plants? Plant Mol Biol 23:209–214
Elhamid AMI, Makboul HE, Sedik MZ, Ibrahin IMA (2010) Cloning, expression and antifungal activity of an endochitinase gene derived from barley (Hordeum vulgare). Res J Agr Biol Scis 6:356–363
Fortunato ASC (2006) Involvement of defense-related genes during Casuarina glauca nodulation. PhD thesis Universidade Nova de Lisboa.
Fortunato A, Santos P, Graça I, Gouveia MM, Martins SM, Ricardo CPP, Pawlowski K, Ribeiro A (2007) Isolation and characterization of cgchi3, a nodule-specific gene from Casuarina glauca encoding a class III chitinase. Physiol Plant 130:418–426
Fukamizo T (2000) Chitinolytic enzymes: catalysis, substrate binding, and their application. Curr Protein Pept Sci 1:105–124
Gerhardt LBD, Sachetto-Martins G, Contarini MG, Sandroni M, Ferreira RD, de Lima MV, Cordeiro MC, de Oliveira DE, Margis-Pinheiro M (1997) Arabidopsis thaliana class IV chitinase is early induced during the interaction with Xanthomonas campestris. FEBS Lett 419:69–75
Glazer AN, Barel AO, Howard JB, Brown DM (1969) Isolation and characterization of fig lysozyme. J Biol Chem 244:3583–3589
Goormachtig S, Lievens S, van de Velde W, van Montagu M, Holsters M (1998) Srchi13, a novel early nodulin from Sesbania rostrata, is related to acidic class III chitinases. Plant Cell 10:905–915
Graham LS, Sticklen MB (1994) Plant chitinases. Can J Bot 72:1057–1083
Howard JB, Glazer AN (1967) Studies on the physicochemical and enzymatic properties of Papaya lysozyme. J Biol Chem 242:5715–5723
Islam R, Datta B (2015) Diversity of chitinases and their industrial potential. Int J Appl Res 1:55–60
Jekbl PA, Hartmann JBH, Beintema JJ (1991) The primary structure of hevamine, an enzyme with lysozyme/chitinase activity from Hevea brasiliensis latex. Eur J Biochem 200:123–130
Jollès P, Muzzarelli RAA (1999) Chitin and Chitinases. Birkhauser Verlag; Basel, Switzerland.
Kasprzewska A (2003) Plant chitinases – regulation and functions. Cell Mol Biol Lett 8:809–824
Kawase T, Yokokawa S, Saito A, Fujii T, Nikaidou N, Miyashita K, Watanabe T (2006) Comparison of enzymatic and antifungal properties between family 18 and 19 chitinases from Streptomyces coelicolor A3(2). Biosci Biotechnol Biochem 70:988–998
Kesari P, Patil DN, Kumar P, Tomar S, Sharma AK, Kumar P (2015) Structural and functional evolution of chitinase-like proteins from plants. Proteomics 15:1693–1705
Kim HB, An CS (2002) Differential expression patterns of an acidic chitinase and a basic chitinase in the root nodule of Elaeagnus umbellata. Mol Plant Microbe Interact 15:209–215
Kim YJ, Kim HB, Baek HE, Heu S, An CS (2005) Constitutive expression of two endochitinases from root nodules of Elaeagnus umbellata confers resistance on transgenic Arabidopsis plants against the fungal pathogen Botrytis cinerea. J Plant Biol 48:39–46
Kirubakaran I, Sakthivel N (2007) Heterologous expression of new antifungal chitinase from wheat. Protein Expr Purif 56:100–109
Kombrink E, Schroder M, Hahlbrock K (1988) Several pathogenesis-related proteins in potato are 1,3-β-glucanases and chitinases. Proc Natl Acad Sci 85:6750–6754
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227(5259):680–685
Lombard V, Golaconda Ramulu H, Drula E, Coutinho PM, Henrissat B (2014) The Carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res 42:D490–D495
Mauch F, Staehelin LA (1989) Functional implications of the subcellular localization of ethylene-induced chitinase and [beta]-1,3-glucanase in bean leaves. Plant Cell 1:447–457
Meier BM, Shaw N, Slusarenko AJ (1993) Spatial and temporal accumulation of defense gene transcripts in bean (Phaseolus vulgaris) leaves in relation to bacteria-induced hypersensitive cell death. Mol Plant Microbe Interact 6:453–466
Nowicki M, Lichocka M, Nowakowska M, Kłosinska U, Kozik EU (2012) A simple dual stain for detailed investigations of plant-fungal pathogen interactions. Veg Crop Res Bull 77:61–74
Patel AK, Singh VK, Yadav RP, Moir AJG, Jagannadham MV (2009) ICChI, a glycosylated chitinase from the latex of Ipomoea carnea. Phytochemistry 70:1210–1216
Patel AK, Singh VK, Yadav RP, Moir AJG, Jagannadham MV (2010) Purification and characterization of a new chitinase from latex of Ipomoea carnea. Process Biochem 45:675–681
Peumans WJ, Proost P, Swennen RL, van Damme EJ (2002) The abundant class III chitinase homolog in young developing banana fruits behaves as a transient vegetative storage protein and most probably serves as an important supply of amino acids for the synthesis of ripening- associated proteins. Plant Physiol 130:1063–1072
Rao DH, Gowda LR (2008) Abundant class III acidic chitinase homologue in tamarind (Tamarindus indica) seed serves as the major storage protein. J Agricult Food Chem 56:2175–82
Regalado AP, Ricardo CPP (1996) Study of the intercellular fluid of healthy Lupinus albus organs: presence of a chitinase and a thaumatin-like protein. Plant Physiol 110:227–232
Regalado AP, Pinheiro C, Vidal S, Chaves I, Ricardo CPP, Pousada CR (2000) The Lupinus albus class-III chitinase gene, IF3, is constitutively expressed in vegetative organs and developing seeds. Planta 210:543–550
Ribeiro A, Graça I, Pawlowski K, Santos P (2011) Actinorhizal plant defence-related genes in response to symbiotic Frankia. Funct Plant Biol 38:639–644
Saborowski R, Buchholz F, Vetter RAH, Wirth SJ, Wolf GA (1993) A soluble, dye-labelled chitin derivative adapted for the assay of krill chitinase. Comp Biochem Physiol 105:673–678
Salzer P, Bonanomi A, Beyer K, Vögeli R, Aeschbacher RA, Lange J, Wiemken A, Kim D, Cook DR, Boller T (2000) Differential expression of eight chitinase genes in Medicago truncatula roots during mycorrhiza formation, nodulation and pathogen infection. Mol Plant Microbe Interact 13:763–777
Samac DA, Shah DM (1991) Developmental and pathogen-induced activation of the Arabidopsis acidic chitinase promoter. Plant Cell 3:1063–1072
Santos P, Fortunato A, Graca I, Martins S, Gouveia M, Auguy F, Bogusz D, Ricardo CPP, Pawlowski K, Ribeiro A (2010) Characterization of four defense-related genes up-regulated in root nodules of Casuarina glauca. Symbiosis 50:27–35
Schwencke J (1991) Rapid, exponential growth and increased biomass yield of some Frankia strains in buffered and stirred mineral medium (BAP) with phosphatidyl choline. Plant Soil 137:37–41
Shugar D (1952) The measurement of lysozyme activity and the ultraviolet inactivation of lysozyme. Biochim Biophys Acta 8:302–309
Sinha M, Singh RP, Kushwaha GS, Iqbal N, Singh A, Kaushik S, Kaur P, Sharma S, Singh TP (2014) Current overview of allergens of plant pathogenesis related protein families. The Scientific World J 2014: ID 543195
Staehelin C, Schultze M, Kondorosi E, Kondorosi A (1995) Lipochitooligosaccharide nodulation signals from Rhizobium meliloti induce their rapid degradation by the host plant alfalfa. Plant Physiol 108:1607–1614
Staehelin C, Schultze M, Tokuyasu K, Poinsot V, Promé JC, Kondorosi E, Kondorosi A (2000) N-deacetylation of Sinorhizobium meliloti Nod factors increases their stability in the Medicago sativa rhizosphere and decreases their biological activity. Mol Plant Microbe Interact 13:72–79
Stintzi A, Geoffroy D, Bersuder D, Fritig B, Legrand M (1993) cDNA cloning and expression studies of tobacco class III chitinases-lysozymes. In: Fritig B, Legrand M (eds) Developments in plant pathology, vol 2. Kluwer, Dordrecht, London, pp 312–315
Taira T (2010) Structures and antifungal activity of plant chitinases. J Appl Glycosci 57:167–176
Theis T, Stahl U (2004) Antifungal protein: target, mechanism and prospective applications. Cell Mol Life Sci 61:437–455
Tisa LS, Beauchemin N, Gtari M, Sen A, Wall LG (2013) What stories can the Frankia genomes start to tell us? J Biosci 38:1–8
Tromas A, Parizot B, Diagne N, Champion A, Hocher V et al (2012) Heart of endosymbioses: transcriptomics reveals a conserved genetic program among arbuscular mycorrhizal, actinorhizal and legume-rhizobial symbioses. PLoS ONE 7, e44742
Tronsmo A, Harman GE (1993) Detection and quantification of N-acetyl-beta-D-glucosaminidase, chitobiosidase, and endochitinase in solutions and on gels. Anal Biochem 208:74–79
Trudel J, Aidy P, Asselin A (1989) Electrophoretic forms of chitinase activity in Xanthi-nc tobacco, healthy and infected with tobacco mosaic virus. Mol Plant Microbe Interact 2:315–324
Vasse J, de Billy F, Truchet G (1993) Abortion of infection during the Rhizobium meliloti–alfalfa symbiotic interactions is accompanied by a hypersensitive reaction. Plant J 4:555–566
Vogelsang R, Barz W (1993) Purification, characterization and differential hormonal regulation of a beta-1,3-glucanase and two chitinases from chickpea (Cicer arietinum L.). Planta 189:60–69
Waffenschmidt S, Jaenicke L (1987) Assay of reducing sugars in the nanomole range with 2,2’-bicinchoninate. Anal Biochem 165:337–340
Wang S, Wu J, Rao P, Ng TB, Ye X (2005) A chitinase with antifungal activity from the mung bean. Protein Expr Purif 40:230−236
Wirth SJ, Wolf GA (1992) Micro-plate colorimetric assay for Endo-acting cellulase, xylanase, chitinase, 1,3-β-glucanase and amylase extracted from forest soil horizons. Soil Biol Biochem 24:511–519
Wohlkönig A, Huet J, Looze Y, Wintjens R (2010) Structural relationships in the lysozyme superfamily: significant evidence for glycoside hydrolase signature motifs. PLoS ONE 5, e15388
Ye X, Ng TB (2005) A chitinase with antifungal activity from the mung bean. Protein Express Purif 40:230–236
Yeh S, Moffatt BA, Griffith M, Xiong F, Yang DSC, Wiseman SB, Sarhan F, Danyluk J, Xue YQ, Hew CL et al (2000) Chitinase genes responsive to cold encode antifreeze proteins in winter cereals. Plant Physiol 124:1251–1263
Yu XM, Griffith M (2001) Winter rye antifreeze activity increases in response to cold and drought, but not abscisic acid. Physiol Plant 112:78–86
Zhong R, Kays SJ, Schroeder BP, Ye ZH (2002) Mutation of a chitinase-like gene causes ectopic deposition of lignin, aberrant cell shapes, and overproduction of ethylene. Plant Cell 14:165–179
Zhong C, Zhang I, Chen Y, Jiang Q, Chen Z, Liang J, Pinyopusarerk K, Franche C, Bogusz D (2010) Casuarina research and applications in China. Symbiosis 50:107–114
Acknowledgments
This work was supported by Fundação para a Ciência e Tecnologia under the scope of the projects PTDC/AGR-FOR/4218/2012 and grant SFRH/ BD/41589/2007, co-financiated by PIDDAC and Fundo Social Europeu. UCIBIO-Research Unit on Applied Molecular Biosciences is financed by Portuguese funds through FCT/MEC (UID/Multi/04378/2013) and co-financed by FEDER under the PT2020 Partnership Agreement. We also thank Dr. Philippe Normand from Lyon University, Dr. Matthieu Joosten from Wageningen University and Dr. Isabel Videira Castro from INIAV, for their valuable contributions to the work.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
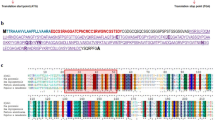
Comparative analysis of the primary structure of CgCHI3 with other proteins identifies as Class III of the chitinase family 18 of glycosyl hydrolases, carried out with the CLC Sequence Viewer software version 7.0.2 (www.clcbio.com). Accession numbers of proteins available in the NCBI database: acidic endochitinase from Vitis vinifera (XP_002279147.1), acidic endochitinase-like from Vitis vinifera (XP_002279205.1), precursor of hevamine-A from Theobroma cacao (XP_007021467.1), chitinase from Populus trichocarpa (XP_002301383.1), putative precursor of hevamine-A from Ricinus communis (XP_002511934.1), Glycine max hevamine-A (XP_003516317.1), acidic endochitinase-like from Cicer arietinum (XP_004490140.1), chitinase class III from Medicago truncatula (AAQ21404.1), hypothetical hevamine-A from Citrus sinensis (XP_006444927.1) and acidic endochitinase from Morus notabilis (EXB52705.1). The catalytic motif is highlighted D145-X-X-D148-X-D150-X-E152. (GIF 337 kb)
Fig. S2
Antibacterial activity of CgCHI3. Plates containing LB agar inoculated with the bacterial species in the presence of CgCHI3 at 37 °C. 1 - E. coli BL21 (DE3), 2 - Paracoccus denitrificans, 3 - Bacillus subtilis. a – Pure enzyme; b – Diluted enzyme 1:4 ; c and d – Positive growth controls in which the enzyme solution was replaced by buffer and water, respectively. (JPEG 28.6 kb)
Rights and permissions
About this article
Cite this article
Graça, I., Liang, J., Guilherme, M. et al. Cloning, overexpression and functional characterization of a class III chitinase from Casuarina glauca nodules. Symbiosis 70, 139–148 (2016). https://doi.org/10.1007/s13199-016-0403-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13199-016-0403-1