Abstract
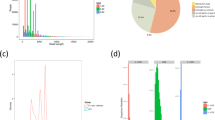
Species of Sargassum are distributed worldwide, and are of great ecological and economic importance in marine ecosystems and bioresources. In this study, transcriptome sequencings of six Sargassum species were performed for the first time using an Illumina platform. For each sample, a total of 2.1–2.5 Gb of nucleotides are collected and assembled into 69 871–116 790 scaffolds, with an average length of 410–550 bp and N50 length of 756–1 462 bp. A total of 20 512–28 684 unigenes of each sample were annotated and compared well with known gene sequences from nr database. Clusters of Orthologous Groups (COG), gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses were also performed for further understanding of gene functions and regulation pathways. Gene expression levels were calculated based on RPKM values and compared among these species, especially for those genes related to carbohydrate metabolism. Cluster analyses indicated that the differences of global gene expression between S. fusiforme, which was nominated as Hizikia fusiformis before, and other five species were not significant. Further phylogenetic analysis of 108 orthologous genes confirmed that S. fusiforme had closer relationship with S. hemiphyllum rather than S. horneri. These transcriptome data provided valuable information for better understanding of genome and gene characteristics of Sargassum algae and benefiting comparative and phylogenetic studies of Phaeophyceae species in future studies.
Similar content being viewed by others
References
Ale M T, Maruyama H, Tamauchi H, et al. 2011. Fucose-containing sulfated polysaccharides from brown seaweeds inhibit proliferation of melanoma cells and induce apoptosis by activation of caspase-3 in vitro. Mar Drugs, 9: 2605–2621
Carpenter E J, Cox J L. 1974. Production of pelagic Sargassum and a blue-green epiphyte in the Western Sargasso Sea. Limnol Oceanogr, 19: 429–436
Cho G Y, Rousseau F, de Reviers B, et al. 2006. Phylogenetic relationships within the Fucales (Phaeophyceae) assessed by the photosystem I coding psaA sequences. Phycologia, 45: 512–519
Cock J M, Sterck L, Rouze P, et al. 2010. The Ectocarpus genome and the independent evolution of multicellularity in brown algae. Nature, 465: 617–621
Conover J T, Sieburth J M. 1964. Effect of Sargassum distribution on its epibiota and antibacterial activity. Botanica Marina, 6: 147–157
Curtis B A, Tanifuji G, Burki F, et al. 2012. Algal genomes reveal evolutionary mosaicism and the fate of nucleomorphs. Nature, 492: 59–65
Deng Yunyan, Yao Jianting, Wang Xiuliang, et al. 2012. Transcriptome sequencing and comparative analysis of Saccharina japonica (Laminariales, Phaeophyceae) under blue light induction. PLoS One, 7: e39704
Dittami S M, Scornet D, Petit J L, et al. 2009. Global expression analysis of the brown alga Ectocarpus siliculosus (Phaeophyceae) reveals large-scale reprogramming of the transcriptome in response to abiotic stress. Genome Biol, 10: R66
Ghangal R, Raghuvanshi S, Chand Sharma P. 2009. Isolation of good quality RNA from a medicinal plant seabuckthorn, rich in secondary metabolites. Plant Physiol Biochem, 47: 1113–1115
Huynh Truonggiang, Yeh Sutuen, Lina Yongchin, et al. 2011. White shrimp Litopenaeus vannamei immersed in seawater containing Sargassum hemiphyllum var. chinense powder and its extract showed increased immunity and resistance against Vibrio alginolyticus and white spot syndrome virus. Fish Shellfish Immunol, 31(2): 286–293
Johnson M T, Carpenter E J, Tian Zhijian, et al. 2012. Evaluating methods for isolating total RNA and predicting the success of sequencing phylogenetically diverse plant transcriptomes. PLoS One, 7: e50226
Li Tianyong, Ren Lei, Zhou Guan, et al. 2012. A suitable method for extracting total RNA from red algae. Transactions of Oceanology and Limnology (in Chinese), 4: 64–71
Makinen V, Salmela L, Ylinen J. 2012. Normalized N50 assembly metric using gap-restricted co-linear chaining. BMC Bioinformatics, 13: 255
Mattio L, Payri C E. 2009. Taxonomic revision of Sargassum species (Fucales, Phaeophyceae) from New Caledonia based on morphological and molecular analyses(1). J Phycol, 45: 1374–1388
Mortazavi A, Williams B A, McCue K, et al. 2008. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods, 5: 621–628
Oak J H, Suh Y, Lee I K. 2002. Phylogenetic relationships of Sargassum subgenus Bactrophycus (Sargassaceae, Phaeophyceae) inferred from rDNA ITS Sequences. Algae, 17: 235–247
Phillips N E, Smith C M, Morden C W. 2005. Testing systematic concepts of Sargassum (Fucales,Phaeophyceae) using portions of the rbcLS operon. Phycol Res, 53: 1–10
Preeprame S, Hayashi K, Lee J B, et al. 2001. A novel antivirally active fucan sulfate derived from an edible brown alga, Sargassum horneri. Chem Pharm Bull (Tokyo), 49: 484–485
Price D C, Chan C X, Yoon H S, et al. 2012. Cyanophora paradoxa genome elucidates origin of photosynthesis in algae and plants. Science, 335: 843–847
Rivera M, Scrosati R. 2006. Population dynamics of Sargassum lapazeanum (Fucales, Phaeophyta) from the Gulf of California, Mexico. Phycologia, 45: 178–189
Southichak B, Nakano K, Nomura M, et al. 2008. Marine macroalga Sargassum horneri as biosorbent for heavy metal removal: roles of calcium in ion exchange mechanism. Water Sci Technol, 58: 697–704
Stamatakis A, Aberer A J, Goll C, et al. 2012. RAxML-Light: a tool for computing terabyte phylogenies. Bioinformatics, 28: 2064–2066
Stiger V, Horiguchi T, Yoshida T, et al. 2003. Phylogenetic relationships within the genus Sargassum (Fucales, Phaeophyceae), inferred from ITS-2 nrDNA, with an emphasis on the taxonomic subdivision of the genus. Phycol Res, 51: 1–10
Tatusov R L, Fedorova N D, Jackson J D, et al. 2003. The COG database: an updated version includes eukaryotes. BMC Bioinformatics, 4: 41
Wang Guoliang, Zhao Ge, Feng Yanbin, et al. 2010. Cloning and comparative studies of seaweed trehalose-6-phosphate synthase genes. Mar Drugs, 8: 2065–2079
Wong C K, Ooi V E, Ang P O. 2000. Protective effects of seaweeds against liver injury caused by carbon tetrachloride in rats. Chemosphere, 41: 173–176
Wong C L, Ng S M, Phang S M. 2007. Use of RAPD in differentiation of selected species of Sargassum (Sargassaceae, Phaeophyta). J Appl Phycol, 19: 771–781
Xu Jia, Aileni M, Abbagani S, et al. 2010. A reliable and efficient method for total rna isolation from various members of spurge family (Euphorbiaceae). Phytochem Anal, 21: 395–398
Yao Jianting, Fu Wandong, Wang Xiuliang, et al. 2008. Improved RNA isolation from Laminaria japonica Aresch (Laminariaceae, Phaeophyta). J Appl Phycol, 21: 233–238
Zdobnov E M, Apweiler R. 2001. InterProScan—an integration platform for the signature-recognition methods in InterPro. Bioinformatics, 17: 847–848
Zeng Chengkui. 2009. Seaweeds in Bohai Sea and Yellow Sea of China (in Chinese). Beijing: Sicence Press, 363–372
Zou Dinghui, Gao Kunshan, Luo Hanjin. 2011. Short- and long-term effects of elevated CO2 on photosynthesis and respiration in the marine macroalga Hizikia Fusiformis (Sargassaceae, Phaeophyta) grown at low and high N supplies. J Phycol, 47: 87–97
Author information
Authors and Affiliations
Corresponding authors
Additional information
Foundation item: The National Natural Science Foundation of China under contract Nos 31140070, 31271397 and 41206116; the algal transcriptome sequencing was supported by 1KP Project (www.onekp.com).
Contributed equally.
Rights and permissions
About this article
Cite this article
Wang, G., Sun, J., Liu, G. et al. Comparative analysis on transcriptome sequencings of six Sargassum species in China. Acta Oceanol. Sin. 33, 37–44 (2014). https://doi.org/10.1007/s13131-014-0439-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13131-014-0439-0