Abstract
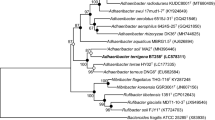
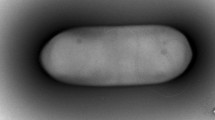
A rod-shaped, white color colony with lobate architectures, strain h2T was isolated from a moderately acidic soil on Jeju Island, Republic of Korea. Comparative analysis of the 16S rRNA gene sequence showed that the strain h2T is closely related to Paenibacillus relictisesami DSM 25385T (97.4%, 16S rRNA gene sequence similarity), Paenibacillus azoreducens KACC 11244T (97.2%), and Paenibacillus cookii LMG 18419T (97.0%). DNA-DNA hybridization indicated that the strain h2T has relatively low levels of DNA-DNA relatedness with respect to P. relictisesami DSM 25385T (10.2%) and P. azoreducens KACC 11244T (13.7%). Additionally, the genomic DNA G + C content of h2T is 51.5 mol%. The isolated strain grew at pH 4.0–9.0 (optimum, pH 6.0–7.0) and 0–5% (w/v) NaCl (optimum, 0%) and a temperature of 15–45°C (optimum 35°C). The quinones in the strain are MK-6 and MK-7, and the predominant fatty acid is C15:0 anteiso (32.1%) followed by C17:0 anteiso (26.5%), and C16:0 iso (21.0%). Based on its phenotypic properties, genotypic distinctiveness, and chemotaxonomic features, strain h2T is proposed as a novel species in the genus Paenibacillus, for which the name Paenibacillus albilobatus sp. nov. is proposed (= KCCM 43269T = JCM 32395T = LMG 30408T). The type strain of Paenibacillus albilobatus is h2T.
Similar content being viewed by others
References
Ash, C., Farrow, J.A., Dorsch, M., Stackebrandt, E., and Collins, M.D. 1991. Comparative analysis of Bacillus anthracis, Bacillus cereus, and related species on the basis of reverse transcriptase sequencing of 16S rRNA. Int. J. Syst. Bacteriol. 41, 343–346.
Ash, C., Priest, F.G., and Collins, M.D. 1993. Molecular identification of rRNA group 3 bacilli (Ash, Farrow, Wallbanks and Collins) using a PCR probe test. Proposal for the creation of a new genus Paenibacillus. Antonie van Leeuwenhoek 64, 253–260.
Bae, J.Y., Kim, K.Y., Kim, J.H., Lee, K., Cho, J.C., and Cha, C.J. 2010. Paenibacillus aestuarii sp. nov., isolated from an estuarine wetland. Int. J. Syst. Evol. Microbiol. 60, 644–647.
Berge, O., Guinebretiere, M.H., Achouak, W., Normand, P., and Heulin, T. 2002. Paenibacillus graminis sp. nov. and Paenibacillus odorifer sp. nov., isolated from plant roots, soil and food. Int. J. Syst. Evol. Microbiol. 52, 607–616.
Ezaki, T., Adnan, S., and Miyake, M. 1990. [Quantitative microdilution plate hybridization to determine genetic relatedness among bacterial strains]. Nihon Saikingaku Zasshi 45, 851–857.
Felsenstein, J. 1981. Evolutionary trees from DNA sequences: a maximum likelihood approach. J. Mol. Evol. 17, 368–376.
Fischer, M. and Thatte, B. 2010. Revisiting an equivalence between maximum parsimony and maximum likelihood methods in phylogenetics. Bull. Math. Biol. 72, 208–220.
González, J.M. and Saiz-Jimenez, C. 2002. A fluorimetric method for the estimation of G + C mol% content in microorganisms by thermal denaturation temperature. Environ. Microbiol. 4, 770–773.
Hu, H.Y., Fujie, K., and Urano, K. 1999. Development of a novel solid phase extraction method for the analysis of bacterial quinones in activated sludge with a higher reliability. J. Biosci. Bioeng. 87, 378–382.
Kimura, M. 1980. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 16, 111–120.
Koh, H.W., Hong, H., Min, U.G., Kang, M.S., Kim, S.G., Na, J.G., Rhee, S.K., and Park, S.J. 2015a. Rhodanobacter aciditrophus sp. nov., an acidophilic bacterium isolated from mine wastewater. Int. J. Syst. Evol. Microbiol. 65, 4574–4579.
Koh, H.W., Rani, S., Kim, S.J., Moon, E., Nam, S.W., Rhee, S.K., and Park, S.J. 2017. Halomonas aestuarii sp. nov., a moderately halophilic bacterium isolated from a tidal flat. Int. J. Syst. Evol. Microbiol. 67, 4298–4303.
Koh, H.W., Song, H.S., Song, U., Yim, K.J., Roh, S.W., and Park, S.J. 2015b. Halolamina sediminis sp. nov., an extremely halophilic archaeon isolated from solar salt. Int. J. Syst. Evol. Microbiol. 65, 2479–2484.
Kumar, S., Stecher, G., and Tamura, K. 2016. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874.
Logan, N.A., De Clerck, E., Lebbe, L., Verhelst, A., Goris, J., Forsyth, G., Rodriguez-Diaz, M., Heyndrickx, M., and De Vos, P. 2004. Paenibacillus cineris sp. nov. and Paenibacillus cookii sp. nov., from Antarctic volcanic soils and a gelatin-processing plant. Int. J. Syst. Evol. Microbiol. 54, 1071–1076.
Meehan, C., Bjourson, A.J., and McMullan, G. 2001. Paenibacillus azoreducens sp. nov., a synthetic azo dye decolorizing bacterium from industrial wastewater. Int. J. Syst. Evol. Microbiol. 51, 1681–1685.
Menendez, E., Flores-Felix, J.D., Mulas, R., Andres, F.G., Fernandez-Pascual, M., Peix, A., and Velazquez, E. 2017. Paenibacillus tritici sp. nov., isolated from wheat roots. Int. J. Syst. Evol. Microbiol. 67, 2312–2316.
Morales, P., Sendra, J.M., and Perez-Gonzalez, J.A. 1995. Purification and characterization of an arabinofuranosidase from Bacillus polymyxa expressed in Bacillus subtilis. Appl. Microbiol. Biotechnol. 44, 112–117.
Ottow, J.C. 1972. Pectinolytic-, ureolytic-, and lecithinolytic activity as a diagnostic aid in the identification of species classified in the genus Bacillus Cohn. Zentralbl. Bakteriol. Parasitenkd. Infektionskr. Hyg. 127, 301–312.
Park, S.J., Cha, I.T., Kim, S.J., Shin, K.S., Hong, Y., Roh, D.H., and Rhee, S.K. 2012. Salinisphaera orenii sp. nov., isolated from a solar saltern. Int. J. Syst. Evol. Microbiol. 62, 1877–1883.
Rani, S., Koh, H.W., Kim, H., Rhee, S.K., and Park, S.J. 2017. Marinobacter salinus sp. nov., a moderately halophilic bacterium isolated from a tidal flat environment. Int. J. Syst. Evol. Microbiol. 67, 205–211.
Saitou, N. and Nei, M. 1987. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425.
Shida, O., Takagi, H., Kadowaki, K., Nakamura, L.K., and Komagata, K. 1997. Transfer of Bacillus alginolyticus, Bacillus chondroitinus, Bacillus curdlanolyticus, Bacillus glucanolyticus, Bacillus kobensis, and Bacillus thiaminolyticus to the genus Paenibacillus and emended description of the genus Paenibacillus. Int. J. Syst. Bacteriol. 47, 289–298.
Shimoyama, T., Johari, N.B., Tsuruya, A., Nair, A., and Nakayama, T. 2014. Paenibacillus relictisesami sp. nov., isolated from sesame oil cake. Int. J. Syst. Evol. Microbiol. 64, 1534–1539.
Tang, Q.Y., Yang, N., Wang, J., Xie, Y.Q., Ren, B., Zhou, Y.G., Gu, M.Y., Mao, J., Li, W.J., Shi, Y.H., et al. 2011. Paenibacillus algorifonticola sp. nov., isolated from a cold spring. Int. J. Syst. Evol. Microbiol. 61, 2167–2172.
Thompson, J.D., Gibson, T.J., Plewniak, F., Jeanmougin, F., and Higgins, D.G. 1997. The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 25, 4876–4882.
Wang, X.M., Ma, S., Yang, S.Y., Peng, R., Zheng, Y., and Yang, H. 2016. Paenibacillus nasutitermitis sp. nov., isolated from a termite gut. Int. J. Syst. Evol. Microbiol. 66, 901–905.
Weisburg, W.G., Barns, S.M., Pelletier, D.A., and Lane, D.J. 1991. 16S ribosomal DNA amplification for phylogenetic study. J. Bacteriol. 173, 697–703.
Xie, J.B., Du, Z., Bai, L., Tian, C., Zhang, Y., Xie, J.Y., Wang, T., Liu, X., Chen, X., Cheng, Q., et al. 2014. Comparative genomic analysis of N2-fixing and non-N2-fixing Paenibacillus spp.: organization, evolution and expression of the nitrogen fixation genes. PLoS Genet. 10, e1004231.
Yarza, P., Yilmaz, P., Pruesse, E., Glockner, F.O., Ludwig, W., Schleifer, K.H., Whitman, W.B., Euzeby, J., Amann, R., and Rossello-Mora, R. 2014. Uniting the classification of cultured and uncultured bacteria and archaea using 16S rRNA gene sequences. Nat. Rev. Microbiol. 12, 635–645.
Yoon, S.H., Ha, S.M., Kwon, S., Lim, J., Kim, Y., Seo, H., and Chun, J. 2017. Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int. J. Syst. Evol. Microbiol. 67, 1613–1617.
Yoon, J.H., Kang, S.J., Yeo, S.H., and Oh, T.K. 2005. Paenibacillus alkaliterrae sp. nov., isolated from an alkaline soil in Korea. Int. J. Syst. Evol. Microbiol. 55, 2339–2344.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplemental material for this article may be found at http://www.springerlink.com/content/120956.
The GenBank/EMBL/DDBJ accession number for the 16S rRNA gene sequence of strain h2T is MF565846.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Lee, JW., Kim, YE., Kang, MS. et al. Paenibacillus albilobatus sp. nov., isolated from acidic soil on Jeju Island. J Microbiol. 56, 393–398 (2018). https://doi.org/10.1007/s12275-018-8158-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-018-8158-4