Abstract
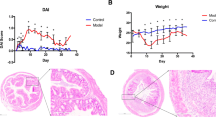
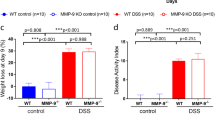
An abnormal composition of the gut microbiota is believed to be associated with the pathogenesis of inflammatory bowel disease (IBD). We utilized terminal restriction fragment length polymorphism (T-RFLP) analysis to quantify faecal bacterial communities from rats with experimental colitis. Male Sprague Dawley rats (n=10/group) ingested 2% dextran sulfate sodium (DSS) or water for up to 7 days. Rats were killed and colonic tissues collected for histological analysis. Damage severity score in the distal colon was significantly greater (P<0.001) following DSS consumption compared to controls. T-RFLP faecal bacterial profiles generated with either MspI or CfoI revealed a significant difference (P<0.001) in community composition between healthy and colitic rats, with bacterial composition in healthy rats more variable than in rats with colitis. Operational taxonomic units (OTU: taxonomically related groups of bacteria) associated with either the healthy or colitic state were identified. OTU (116, 226, 360, and 948; CfoI) and (118 and 188; MspI) were strongly associated with untreated healthy rats, while OTU (94, 98, 174, and 384; CfoI) and (94 and 914; MspI) were predominantly associated with DSS-treated colitic rats. Phylogenetic OTU assignment suggested that Bacteroidales and Lactobacillus sp. were predominantly associated with the colitic and healthy rats, respectively. These results show that faecal bacterial profiling is a rapid, sensitive and non-invasive tool for detecting and identifying changes in gut microbiota associated with colitis. Restoring microbial homeostasis by targeting colitis-associated OTU through specific microbiological interventions could form the basis of novel therapeutic strategies for IBD.
Similar content being viewed by others
References
Andoh, A., Sakata, S., Koizumi, Y., Mitsuyama, K., Fujiyama, Y., and Benno, Y. 2007. Terminal restriction fragment length polymorphism analysis of the diversity of fecal microbiota in patients with ulcerative colitis. Inflamm. Bowel Dis.13, 955–962.
Bibiloni, R., Fedorak, R.N., Tannock, G.W., Madsen, K.L., Gionchetti, P., Campieri, M., De Simone, C., and Sartor, R.B. 2005. VSL#3 probiotic-mixture induces remission in patients with active ulcerative colitis. Am. J. Gastroenterol.100, 1539–1546.
Bjerketorp, J., Chiang, A.N.T., Hjort, K., Rosenquist, M., Liu, W.T., and Jansson, J.K. 2008. Rapid lab-on-a-chip profiling of human gut bacteria. J. Microbiol. Methods72, 82–90.
Bousvaros, A., Morley, F.A., Pensabene, L., and Cucchiara, S. 2008. Research and clinical challenges in pediatric inflammatory bowel disease. Dig. Liver. Dis.40, 32–38.
Bray, J.R. and Curtis, K.R. 1957. An ordination of the upland forest communities of southern Wisconsin. Ecol. Monogr.27, 325–349.
Clarke, K.R. 1993. Non-parametric multivariate analyses of changes in community structure. Austr. J. Ecol.18, 117–143.
Clarke, K.R. and Warwick, R.M. 2001. Changes in marine communities: an approach to statistical analysis and interpretation. 2nd ed. Primer F Ltd., Plymouth, UK.
Dethlefsen, L., Mcfall-Ngai, M., and Relman, D.A. 2007. An ecological and evolutionary perspectives on human microbe mutualism and disease. Nature449, 811–818.
Dicksved, J., Floistrup, H., Bergstrom, A., Rosenquist, M., Pershagen, G., Scheynius, A., Roos, S., Alm, J.S., Engstrand, L., Braun-Fahrlander, C., andet al. 2007. Molecular fingerprinting of the fecal microbiota of children raised according to different lifestyles. Appl. Environ. Microbiol.73, 2284–2289.
Eckburg, P.B., Bik, E.M., Bernstein, C.N., Purdom, E., Dethlefsen, L., Sargent, M., Gill, S.R., Nelson, K.E., and Relman, D.A. 2005. Diversity of the human intestinal microbial flora. Science308, 1635–1638.
Egert, M., Marhan, S., Wagner, B., Scheu, S., and Friedrich, M.W. 2004. Molecular profiling of 16S rRNA genes reveals diet-related differences of microbial communities in soil, gut, and casts of Lumbricus terrestris L. (Oligochaeta: Lumbricidae). FEMS Microbiol. Ecol.48, 187–197.
Fiocchi, C. 1998. Inflammatory bowel disease: etiology and pathogenesis. Gastroenterol.155, 182–205.
Frank, D.N., Amand, A.L.S., Feldman, R.A., Boedeker, E.C., Harpaz, N., and Pace, N.R. 2007. Molecular phylogenetic characterization of microbial community imbalance in human inflammatory bowel diseases. Proc. Natl. Acad. Sci. USA104, 13780–13785.
Geier, M.S., Butler, R.N., Giffard, P.M., and Howarth, G.S. 2007. Lactobacillus fermentum BR11, a potential new probiotic, alleviates symptoms of colitis induced by dextran sulfate sodium (DSS) in rats. Int. J. Food Microbiol.114, 267–274.
Gorbach, S.L., Nahas, L., Plaut, A.G., Weinstein, L., Patterson, J.F., and Levitan, R. 1968. Studies of intestinal microflora. V. Faecal microbial ecology in ulcerative colitis and regional enteritis: relationship to severity of disease and chemotherapy. Gastroenterol.54, 575–587.
Hartley, M.G., Hudson, M.J., Swarbrick, E.T., Hill, M.J., Gent, A.E., Hellier, M.D., and Grace, R.H. 1992. The rectal mucosa associated microflora in patients with ulcerative colitis. J. Med. Microbiol.36, 33–39.
Heimesaat, M.M., Fischer, A., Siegmund, B., Kupz, A., Niebergall, J., Fuchs, D., Jahn, H.K., Freudenberg, M., Loddenkemper, C., Batra, A., andet al. 2007. Shifts towards proinflammatory intestinal bacteria aggravate acute murine colitis via toll like receptors 2 and 4. PLoS One2, e662.
Kent, A.D., Smith, D.J., Benson, B.J., and Triplett, E.W. 2003. A web-based phylogenetic assignment tool for analysis of terminal restriction fragment length polymorphism profiles of microbial communities. Appl. Environ. Microbiol.69, 6768–6776.
Kruskal, J.B. 1964. Multidimensional scaling by optimizing a goodness of fit to a nonmetric hypothesis. Psychometrics29, 1–28.
Lane, D.J. 1991. 16S/23S rRNA sequencing, pp. 115–175. In Stackebrandt, E. and Goodfellow, M. (eds.), Nucleic acid techniques in bacterial systematics. John Wiley and Sons Inc., New York, USA.
Lee, H.K., Kim, H.R., Mengoni, A., and Lee, D.H. 2008. Modified T-RFLP methods for taxonomic interpretation of T-RF. J. Microbiol. Biotechnol.18, 624–630.
Li, F., Hullar, M.A.J., and Lampe, J.W. 2007. Optimization of terminal restriction fragment polymorphism (TRFLP) analysis of human gut microbiota. J. Microbiol. Methods68, 303–311.
Lupp, C., Robertson, M.L., Wickham, M.E., Sekirov, I., Champion, O.L., Gaynor, E.C., and Finlay, B.B. 2007. Host mediated inflammation disrupts the intestinal microbiota and promotes the overgrowth of Enterobacteriaceae. Cell. Host Microbe 2, 119–129.
Mai, V., Braden, C.R., Heckendorf, J., Pironis, B., and Hirshon, J.M. 2006. Monitoring of stool microbiota in subjects with diarrhea indicates distortions in composition. J. Clin. Microbiol.44, 4550–4552.
Manichanh, C., Rigottier-Gois, L., Bonnaud, E., Gloux, K., Pelletier, E., Frangeul, L., Nalin, R., Jarrin, C., Chardon, P., Marteau, P., andet al. 2006. Reduced diversity of faecal microbiota in Crohn’s disease revealed by a metagenomic approach. Gut55, 205–211.
Marteau, P., Seksik, P., and Shanahan, F. 2003. Manipulation of the bacterial flora in inflammatory bowel disease. Best Pract. Res. Clin. Gastroenterol.17, 47–61.
Munkholm, P., Langholz, E., Davidsen, M., and Binder, V. 1995. Disease activity courses in a regional cohort of Crohn’s disease patients. Scand. J. Gastroenterol.30, 699–706.
Murthy, S.N., Cooper, H.S., Shim, H., Shah, R.S., Ibrahim, S.A., and Sedergran, D.J. 1993. Treatment of dextran sulphate sodium induced murine colitis by intracolonic cyclosporine. Dig. Dis. Sci.38, 1722–1734.
Muyzer, G., Teske, A., Wirsen, C.O., and Jannasch, H.W. 1995. Phylogenetic relationships of Thiomicrospira species and their identification in deep sea hydrothermal vent samples by denaturing gradient gel electrophoresis of 16S rDNA fragments. Arch. Microbiol.164, 165–172.
Nagashima, K., Hisada, T., Sato, M., and Mochizuki, J. 2003. Application of new primer-enzyme combination to terminal restriction fragment length polymorphism profiling of bacterial populations in human feces. Appl. Environ. Microbiol.69, 1251–1262.
Scanlan, P.D., Shanahan, F., O’Mahony, C., and Marchesi, J.R. 2006. Culture independent analyses of temporal variation of the dominant fecal microbiota and targeted bacterial subgroups in Crohn’s disease. J. Clin. Microbiol.44, 3980–3988.
Sellon, R.K., Tonkonogy, S., Schultz, M., Dieleman, L.A., Grenther, W., Balish, E., Rennick, D.M., and Sartor, R.B. 1998. Resident enteric bacteria are necessary for development of spontaneous colitis and immune system activation in interleukin 10 deficient mice. Infect. Immun.66, 5224–5231.
Sepehri, S., Kotlowski, R., Bernstein, C.N., and Krause, D.O. 2007. Microbial diversity of inflamed and noninflamed gut biopsy tissues in inflammatory bowel disease. Inflamm. Bowel. Dis.13, 675–683.
Shepard, R.N. 1962. The analysis of proximities: multidimensional scaling with an unknown distance function: parts I and II. Psychometrics27, 125–140.
Shyu, C., Soule, T., Bent, S.J., Foster, J.A., and Forney, L.J. 2007. MiCA: A web-based tool for the analysis of microbial communities based on terminal-restriction fragment length polymorphisms of 16S and 18S rRNA Genes. J. Microb. Ecol.53, 562–570.
Steed, H., Macfarlane, G.T., and Macfarlane, S. 2008. Prebiotics, synbiotics and inflammatory bowel disease. Mol. Nutr. Food Res.52, 898–905.
Stirling, G.R., Griffin, D., Ophel-Keller, K., McKay, A., Hartley, D., Curran, J., Stirling, A.M., Monsour, D., Winch, J., and Hardie, B. 2004. Combining an initial risk assessment process with DNA assays to improve prediction of soilborne diseases caused by root-knot nematode (Meloidogyne spp.) and Fusarrium oxysporum f. sp. lycopersici in the Queensland tomato industry. Aus. Plant Pathol.33, 285–293.
Strober, W., Fuss, I., and Mannon, P. 2007. The fundamental basis of inflammatory bowel disease. J. Clin. Invest.117, 514–521.
Swidsinski, A., Loening-Baucke, V., Lochs, H., and Hale, L.P. 2005. Spatial organization of bacterial flora in normal and inflamed intestine: a fluorescence in situ hybridization study in mice. World J. Gastroenterol.11, 1131–1140.
Takaishi, H., Matsuki, T., Nakazawa, A., Takada, T., Kado, S., Asahara, T., Kamada, N., Sakuraba, A., Yajima, T., Higuchi, H., andet al. 2008. Imbalance in intestinal microflora constitution could be involved in the pathogenesis of inflammatory bowel disease. Int. J. Med. Microbiol.298, 463–472.
Tamboli, C.P., Neut, C., Desreumaux, P., and Colombel, J.F. 2004. Dysbiosis in inflammatory bowel disease. Gut53, 1–4.
Tannock, G.W. 2007. What immunologists should know about bacterial communities of the human bowel. Semin. Immunol.19, 94–105.
Taurog, J.D., Richardson, J.A., Croft, J.T., Simmons, W.A., Zhou, M., Fernandez-Sueiro, J.L., Balish, E., and Hammer, R.E. 1994. The germfree state prevents development of gut and joint inflammatory disease in HLA-B27 transgenic rats. J. Exp. Med.180, 2359–2364.
Tomas, F.M., Knowles, S.E., Owens, P.C., Read, L.C., Chandler, C.S., Gargosky, S.E., and Ballard, F.J. 1991. Increased weight gain, nitrogen retention and muscle protein synthesis following treatment of diabetic rats with insulin like growth factor (IGF)-1 and des (1–3) IGF-1. Biochem. J.276, 547–554.
Torok, V.A., Hughes, R.J., Ophel-Keller, K., Ali, M., and MacAlpine, R. 2009. Influence of different litter materials on cecal microbiota colonization in broiler chickens. Poult. Sci.88, 2474–2481.
Torok, V.A., Ophel-Keller, K., Loo, M., and Hughes, R.J. 2008. Application of methods for identifying broiler chicken gut bacterial species linked with increased energy metabolism. Appl. Environ. Microbiol.74, 783–791.
Widmer, F., Hartmann, M., Frey, B., and Kolliker, R. 2006. A novel strategy to extract specific phylogenetic sequence information from community T-RFLP. J. Microbiol. Methods66, 512–520.
Xenoulis, P.G., Palculict, B., Allenspach, K., Steiner, J.M., Van House, A.M., and Suchodolski, J.S. 2008. Molecular phylogenetic characterization of microbial communities imbalances in the small intestine of dogs with inflammatory bowel disease. FEMS Microbiol. Ecol.66, 579–589.
Yazbeck, R., Howarth, G.S., Geier, M.S., Demuth, H.U., and Abbott, C.A. 2008. Inhibiting dipeptidyl peptidase activity partially ameliorates colitis in mice. Front. Biosci.13, 6850–6858.
Zoetendal, E.G., von Wright, A., Vilpponen-Salmela, T., Ben-Amor, K., Akkermans, A.D.L., and de Vos, W.M. 2002. Mucosa-associated bacteria in the human gastrointestinal tract are uniformly distributed along the colon and differ from the community recovered from faces. Appl. Environ. Microbiol.68, 3401–3407.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplemental material for this article may be found at http://www.springerlink.com/content/120956.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Samanta, A.K., Torok, V.A., Percy, N.J. et al. Microbial fingerprinting detects unique bacterial communities in the faecal microbiota of rats with experimentally-induced colitis. J Microbiol. 50, 218–225 (2012). https://doi.org/10.1007/s12275-012-1362-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-012-1362-8