Abstract
Purpose
Emergent cancer cells likely secrete factors that inhibit anti-tumor immunity. To identify such factors, we applied a functional assay with proteomics to an immunotherapy resistant syngeneic mouse melanoma model. Four secreted factors were identified that potentially mediate immunosuppression and could become targets for novel immunotherapies. We tested for consistent clinical correlates in existing human data and verified in vivo whether knocking out tumor cell production of these factors improved immune-mediated control of tumor growth.
Methods
Existing human data was analyzed for clinical correlates. A CRISPR/Cas9 approach to generate knockout cell lines and a kinetic analysis leveraging a Markov Chain Monte Carlo (MCMC) approach quantified the various knockouts’ effect on cells’ intrinsic growth rate. Flow cytometry was used to characterize differences in immune infiltration.
Results
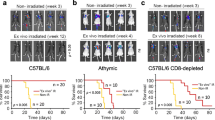
While all four gene products were produced by malignant melanocytes, only increased CCN4 expression was associated with reduced survival in primary melanoma patients. In immunocompetent C57BL/6 mice the CCN4 knockout increased survival while the other knockouts had no effect. This survival advantage was lost when the CCN4 knockout cells were injected into immunocompromised hosts, indicating that the effect of CCN4 may be immune mediated. Parameter estimation from the MCMC analysis shows that CCN4 was the only knockout tested that decreased the net tumor growth rate in immunocompetent mice. Flow cytometry showed an increase in NK cell infiltration in CCN4 knockout tumors.
Conclusions
The results suggest that CCN4 is a mediator of immunosuppression in the melanoma tumor microenvironment and a potential collateral immunotherapy target.






Similar content being viewed by others
Data Availability
Bulk tissue transcriptomics profiling data using Illumina RNA sequencing was accessed from the cutaneous melanoma (SKCM) arm of the Cancer Genome Atlas. Data were downloaded from TCGA data commons using the “TCGAbiolinks” (V2.14.1) package in R (V3.6.3). Gene expression data were expressed in counts. The single cell RNAseq datasets used in the analysis for this article are available in Gene Expression Omnibus repository with the following GEO accession numbers: GSE115978 and GSE72056
Code Availability
The code used in the Markov Chain Monte Carlo analysis can be obtained from the following GitHub repository: https://github.com/arcoolbaugh/B16-In-Vivo-Screen
Abbreviations
- IL-12:
-
Interleukin 12
- DNMT3A:
-
DNA methyltransferase 3
- PTPN11:
-
Protein Tyrosine Phosphatase Non-Receptor Type 11
- CCN4:
-
Cellular Communication Network Factor 4 or Wnt-inducible signaling pathway protein-1
- SPARC:
-
Secreted Protein Acidic and Rich in Cysteine
- ATCC:
-
American Tissue Culture Collection
- NSG:
-
NOD-scid IL2R γnull immunodeficient mice
- WVU:
-
West Virginia University
- SKCM:
-
Skin cutaneous melanoma
- TCGA:
-
The Cancer Genome Atlas
- scRNAseq:
-
Single cell RNA sequencing
- MCMC:
-
Markov Chain Monte Carlo
- WT:
-
Wildtype
- KO:
-
Knock out
References
Tumeh, P. C., et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature. 515(7528):568–571, 2014. https://doi.org/10.1038/nature13954.
Ganesan, S., and J. Mehnert. Biomarkers for response to immune checkpoint blockade. Annu. Rev. Cancer Biol. 4:331–351, 2020. https://doi.org/10.1146/annurev-cancerbio-030419-033604.
Garris, C. S., et al. Successful anti-PD-1 cancer immunotherapy requires T cell-dendritic cell crosstalk involving the cytokines IFN-γ and IL-12. Immunity. 49(6):1148-1161.e7, 2018. https://doi.org/10.1016/j.immuni.2018.09.024.
Kulkarni, Y. M., E. Chambers, A. J. R. McGray, J. S. Ware, J. L. Bramson, and D. J. Klinke. A quantitative systems approach to identify paracrine mechanisms that locally suppress immune response to Interleukin-12 in the B16 melanoma model. Integr. Biol. (United Kingdom). 4(8):925–936, 2012. https://doi.org/10.1039/c2ib20053h.
Wu, Y., W. Deng, E. C. McGinley, and D. J. Klinke. Melanoma exosomes deliver a complex biological payload that upregulates PTPN11 to suppress T lymphocyte function. Pigment Cell Melanoma Res. 30(2):203–218, 2017. https://doi.org/10.1111/pcmr.12564.
Bland, C. L., C. N. Byrne-Hoffman, A. Fernandez, S. L. Rellick, W. Deng, and D. J. Klinke. Exosomes derived from B16F0 melanoma cells alter the transcriptome of cytotoxic T cells that impacts mitochondrial respiration. FEBS J. 285(6):1033–1050, 2018. https://doi.org/10.1111/febs.14396.
Klinke, D. J., A. Fernandez, W. Deng, A. Razazan, H. Latifizadeh, and A. C. Pirkey. Data-driven learning how oncogenic gene expression locally alters heterocellular networks. Nat. Commun. 13(1):1986, 2022. https://doi.org/10.1038/s41467-022-29636-3.
Deng, W., A. Fernandez, S. L. McLaughlin, and D. J. Klinke. WNT1-inducible signaling pathway protein 1 (WISP1/CCN4) stimulates melanoma invasion and metastasis by promoting the epithelial mesenchymal transition. J. Biol. Chem. 294(14):5261–5280, 2019. https://doi.org/10.1074/jbc.RA118.006122.
Jerby-Arnon, L., et al. A cancer cell program promotes T cell exclusion and resistance to checkpoint blockade. Cell. 175(4):984-997.e24, 2018. https://doi.org/10.1016/j.cell.2018.09.006.
Tirosh, I., et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science (1979). 352(6282):189–196, 2016. https://doi.org/10.1126/science.aad0501.
Klinke, D. J. In silico model-based inference: a contemporary approach for hypothesis testing in network biology. Biotechnol. Prog. 30(6):1247–1261, 2014. https://doi.org/10.1002/btpr.1982.
Deng, W., A. Fernandez, S. L. McLaughlin, and D. J. Klinke. Cell Communication Network Factor 4 (CCN4/WISP1) shifts melanoma cells from a fragile proliferative state to a resilient metastatic state. Cell Mol. Bioeng. 13(1):45–60, 2020. https://doi.org/10.1007/s12195-019-00602-2.
Overwijk, W. W., and N. P. Restifo. B16 as a mouse model for human melanoma. Curr. Protoc. Immunol. 2000. https://doi.org/10.1002/0471142735.im2001s39.
Byrne-Hoffman, C. N., W. Deng, O. McGrath, P. Wang, Y. Rojanasakul, and D. J. Klinke. Interleukin-12 elicits a non-canonical response in B16 melanoma cells to enhance survival. Cell Commun. Signal. 2020. https://doi.org/10.1186/s12964-020-00547-4.
Jia, H., et al. The tumor cell-secreted matricellular protein WISP 1 drives pro-metastatic collagen linearization. EMBO J. 38(16):1–15, 2019. https://doi.org/10.15252/embj.2018101302.
Wang, Q. Y., Y. J. Feng, and R. Ji. High expression of WISP1 promotes metastasis and predicts poor prognosis in hepatocellular carcinoma. Eur. Rev. Med. Pharmacol. Sci. 24(20):10445–10451, 2020. https://doi.org/10.26355/eurrev_202010_23396.
Fernandez, A., et al. Cell communication network factor 4 promotes tumor-induced immunosuppression in melanoma. EMBO Rep. 2022. https://doi.org/10.15252/embr.202154127.
Hou, C. H., C. H. Tang, C. J. Hsu, S. M. Hou, and J. F. Liu. CCN4 induces IL-6 production through αvβ5 receptor, PI3K, Akt, and NF-κB singling pathway in human synovial fibroblasts. Arthritis Res. Ther. 15(1):R19, 2013. https://doi.org/10.1186/ar4151.
Ono, M., et al. CCN4/WISP1 controls cutaneous wound healing by modulating proliferation, migration and ECM expression in dermal fibroblasts via α5β1 and TNFα. Matrix Biol. 68–69:533–546, 2018. https://doi.org/10.1016/j.matbio.2018.01.004.
Tao, W., et al. Dual Role of WISP1 in maintaining glioma stem cells and tumor-supportive macrophages in glioblastoma. Nat. Commun. 2020. https://doi.org/10.1038/s41467-020-16827-z.
Wu, S. Z., et al. A single-cell and spatially resolved atlas of human breast cancers. Nat. Genet. 53(9):1334–1347, 2021. https://doi.org/10.1038/s41588-021-00911-1.
Chen, B., and W. Chan. The de novo DNA methyltransferase DNMT3A in development and cancer. Epigenetics. 9(5):669–677, 2014. https://doi.org/10.4161/epi.28324.
Robertson, K. D., et al. The human DNA methyltransferases (DNMTs) 1, 3a and 3b: coordinate mRNA expression in normal tissues and overexpression in tumors. Nucleic Acids Res. 27(11):2291–2298, 1999. https://doi.org/10.1093/nar/27.11.2291.
Zhang, W., and J. Xu. DNA methyltransferases and their roles in tumorigenesis. Biomark. Res. 5(1):1–8, 2017. https://doi.org/10.1186/s40364-017-0081-z.
Deng, T., Y. Kuang, L. Wang, J. Li, Z. Wang, and J. Fei. An essential role for DNA methyltransferase 3a in melanoma tumorigenesis. Biochem. Biophys. Res. Commun. 387(3):611–616, 2009. https://doi.org/10.1016/j.bbrc.2009.07.093.
Moufarrij, S., et al. Combining DNMT and HDAC6 inhibitors increases anti-tumor immune signaling and decreases tumor burden in ovarian cancer. Sci. Rep. 2020. https://doi.org/10.1038/s41598-020-60409-4.
Segovia, C., et al. Inhibition of a G9a/DNMT network triggers immune-mediated bladder cancer regression. Nat. Med. 25(7):1073–1081, 2019. https://doi.org/10.1038/s41591-019-0499-y.
Hill, K. S., et al. PTPN11 plays oncogenic roles and is a therapeutic target for BRAF wild-type melanomas. Mol. Cancer Res. 17(2):583–593, 2019. https://doi.org/10.1158/1541-7786.MCR-18-0777.
Girotti, M. R., et al. SPARC promotes cathepsin B-mediated melanoma invasiveness through a collagen i/α2Β1 integrin axis. J. Invest. Dermatol. 131(12):2438–2447, 2011. https://doi.org/10.1038/jid.2011.239.
Jiang, Y., and H. Zhan. Communication between EMT and PD-L1 signaling: new insights into tumor immune evasion. Cancer Lett. 468(2019):72–81, 2020. https://doi.org/10.1016/j.canlet.2019.10.013.
Haber, C. L., et al. SPARC modulates the proliferation of stromal but not melanoma cells unless endogenous SPARC expression is downregulated. Int. J. Cancer. 122(7):1465–1475, 2008. https://doi.org/10.1002/ijc.23216.
Prada, F., L. G. Benedetti, A. I. Bravo, M. J. Alvarez, C. Carbone, and O. L. Podhajcer. SPARC endogenous level, rather than fibroblast-produced SPARC or stroma reorganization induced by SPARC, is responsible for melanoma cell growth. J. Invest. Dermatol. 127(11):2618–2628, 2007. https://doi.org/10.1038/sj.jid.5700962.
Manguso, R. T., et al. In vivo CRISPR screening identifies Ptpn2 as a cancer immunotherapy target. Nature. 547(7664):413–418, 2017. https://doi.org/10.1038/nature23270.
Masoudi, M., M. Seki, R. Yazdanparast, N. Yachie, and H. Aburatani. A genome-scale CRISPR/Cas9 knockout screening reveals SH3D21 as a sensitizer for gemcitabine. Sci. Rep. 9(1):1–9, 2019. https://doi.org/10.1038/s41598-019-55893-2.
Acknowledgements
Figure 4 was created with Biorender.com, for which the subscription was provided by the Cellular and Molecular Biology and Biomedical Engineering NIH T32 Program (GM133369) at West Virginia University. We would also like to thank Audry Fernandez for her assistance with flow cytometry experiments. This work was supported by grants received by David. J. Klinke II from the National Science Foundation (NSF CBET-1644932) and National Cancer Institute (NCI 1R01CA193473). The content is solely the responsibility of the authors and does not necessarily represent the official views of the NSF or NCI. We also used equipment from the WVU Flow Cytometry & Single Cell Core Facility (RRID: SCR_017738). The core was supported by National Institute of Health Grants GM121322, GM103434, OD016165, and GM104942.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Animal Studies
All procedures involving animals were approved by the West Virginia University (WVU) Institutional Animal Care and Use Committee and performed at the WVU Animal Facility ((IACUC Protocol #1604002138).
Additional information
Associate Editor Matthew Lazzara oversaw the review of this article.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
12195_2023_787_MOESM1_ESM.tif
Supplementary file1 (TIF 17743 kb) Supplemental Figure S1: Western blot images confirming the knockout of (A) DNMT3A, (B) PTPN11, (C) SPARC, and (D) CCN4 in B16F0 cells.
12195_2023_787_MOESM2_ESM.tif
Supplementary file2 (TIF 13841 kb) Supplemental Figure S2: Representative images of diagnostics for Markov Chain Monte Carlo estimates of the posterior distribution of initial tumor bolus size in a single mouse. (A) The posterior distribution of the parameter for each of the three independent chains overlap, indicating agreement between the chains. (B) New steps in the Markov Chain were proposed with increased or decreased risk to achieve a desired acceptance fraction of test point of 0.20. (C) The full-length traces of three independent chains (colored in black, blue, and red, respectively) with over disperse starting points. (D) The Gelman-Rubin potential scale reduction factor (PSRF) was used to assess convergence of the Markov Chains to the posterior distribution in each parameter where a value of less than 1.2 indicated that the chains have converged.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pirkey, A.C., Deng, W., Norman, D. et al. Head-to-Head Comparison of CCN4, DNMT3A, PTPN11, and SPARC as Suppressors of Anti-tumor Immunity. Cel. Mol. Bioeng. 16, 431–442 (2023). https://doi.org/10.1007/s12195-023-00787-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12195-023-00787-7