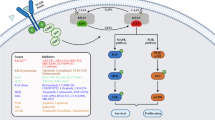
Abstract
Colorectal cancer (CRC) is a malignant tumor initiating from the mucosa of the colorectum. According to the 2020 statistics from the World Health Organization, there are 10.0% CRC cases among all 19.3 million new cancers, followed by lung and breast cancer, and 9.4% CRC cases among all 9.9 million cancer deaths, ranking second. The population of CRC patients in China is large, and its incidence and mortality continue to increase each year. Despite the continuous development of surgical methods, chemotherapy, radiotherapy, targeted therapy and immunotherapy, the overall survival of CRC patients remains low. Past research has suggested that c-myc plays a pivotal role in the development of CRC. A higher expression level of c-Myc is a negative prognostic marker in CRC. However, there are few drugs targeting c-myc directly. Therefore, we focused on discovering the mechanism of c-myc in CRC to provide a reference for a better therapy choice for patients.


Similar content being viewed by others
References
Sung H, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71(3):209–49. https://doi.org/10.3322/caac.21660.
Xia C, et al. Cancer statistics in China and United States, 2022: profiles, trends, and determinants. Chin Med J (Engl). 2022;135(5):584–90. https://doi.org/10.1097/CM9.0000000000002108.
Cheng Y, et al. The effect of type 2 diabetes mellitus on the short-term outcomes and prognosis of stage I–III colorectal cancer: a propensity score matching analysis. Cancer Manag Res. 2022;14:205–14. https://doi.org/10.2147/CMAR.S347242.
Chakedis J, Schmidt CR. Surgical treatment of metastatic colorectal cancer. Surg Oncol Clin N Am. 2018;27(2):377–99. https://doi.org/10.1016/j.soc.2017.11.010.
Gelibter AJ, et al. Adjuvant chemotherapy in resected colon cancer: when, how and how long? Surg Oncol. 2019;30:100–7. https://doi.org/10.1016/j.suronc.2019.06.003.
Hasbullah HH, Musa M. Gene therapy targeting p53 and KRAS for colorectal cancer treatment: a myth or the way forward? Int J Mol Sci. 2021. https://doi.org/10.3390/ijms222111941.
Kishore C, Bhadra P. Current advancements and future perspectives of immunotherapy in colorectal cancer research. Eur J Pharmacol. 2021;893: 173819. https://doi.org/10.1016/j.ejphar.2020.173819.
Peng D, Cheng YX, Cheng Y. Improved overall survival of colorectal cancer under multidisciplinary team: a meta-analysis. Biomed Res Int. 2021;2021:5541613. https://doi.org/10.1155/2021/5541613.
Byrne R, et al. Age-related differences in gene expression in colorectal cancer (CRC). J Clin Oncol. 2018;36:654–654. https://doi.org/10.1200/JCO.2018.36.4_suppl.654.
Caiado H, et al. Evaluation of MGP gene expression in colorectal cancer. Gene. 2019;723: 144120. https://doi.org/10.1016/j.gene.2019.144120.
Cherradi S, et al. Claudin gene expression profiles and clinical value in colorectal tumors classified according to their molecular subtype. Cancer Manag Res. 2019;11:1337–48. https://doi.org/10.2147/CMAR.S188192.
Yang Q, et al. Gene expression profile comparison between colorectal cancer and adjacent normal tissues. Oncol Lett. 2017. https://doi.org/10.3892/ol.2017.6915.
Lee CM, Reddy EP. The v-myc oncogene. Oncogene. 1999;18(19):2997–3003. https://doi.org/10.1038/sj.onc.1202786.
He TC, et al. Identification of c-MYC as a target of the APC pathway. Science. 1998; 281(5382):1509–12; https://doi.org/10.1126/science.281.5382.1509.
Palomero T, et al. NOTCH1 directly regulates c-MYC and activates a feed-forward-loop transcriptional network promoting leukemic cell growth. Proc Natl Acad Sci U S A, 2006. 103(48): p. 18261–6; https://doi.org/10.1073/pnas.0606108103.
Strippoli A, et al. c-MYC expression is a possible keystone in the colorectal cancer resistance to EGFR inhibitors. Cancers. 2020. https://doi.org/10.3390/cancers12030638.
Wang W, et al. Synergistic role of Cul1 and c-Myc: prognostic and predictive biomarkers in colorectal cancer. Oncol Rep. 2017;38(1):245–52. https://doi.org/10.3892/or.2017.5671.
Lee K, et al. Favorable prognosis in colorectal cancer patients with co-expression of c-MYC and ß-catenin. BMC Cancer. 2016;16(1):730. https://doi.org/10.1186/s12885-016-2770-7.
Lee K, et al. c-MYC copy-number gain is an independent prognostic factor in patients with colorectal cancer. PLoS One. 2015;10(10): e0139727. https://doi.org/10.1371/journal.pone.0139727.
Toon C, et al. Immunohistochemistry for myc predicts survival in colorectal cancer. PLoS One. 2014;9(2): e87456. https://doi.org/10.1371/journal.pone.0087456.
Kriegl L, et al. Up-regulation of c-MYC and SIRT1 expression correlates with malignant transformation in the serrated route to colorectal cancer. Oncotarget. 2012;3(10):1182–93. https://doi.org/10.18632/oncotarget.628.
Masramon L, et al. Moderate amplifications of the c-myc gene correlate with molecular and clinicopathological parameters in colorectal cancer. Br J Cancer. 1998;77(12):2349–56. https://doi.org/10.1038/bjc.1998.390.
Rochlitz C, Herrmann R, de Kant E. Overexpression and amplification of c-myc during progression of human colorectal cancer. Oncology. 1996;53(6):448–54. https://doi.org/10.1159/000227619.
Yang J, et al. Higher expression of oncoproteins c-myc, c-erb B-2/neu, PCNA, and p53 in metastasizing colorectal cancer than in nonmetastasizing tumors. Ann Surg Oncol. 1996;3(6):574–9. https://doi.org/10.1007/bf02306092.
Kozma L, et al. Investigation of c-myc oncogene amplification in colorectal cancer. Cancer Lett. 1994;81(2):165–9. https://doi.org/10.1016/0304-3835(94)90198-8.
Sato K, et al. c-myc mRNA overexpression is associated with lymph node metastasis in colorectal cancer. Eur J Cancer (Oxford, England: 1990). 1994. https://doi.org/10.1016/0959-8049(94)90468-5.
Sharrard R, et al. Patterns of methylation of the c-myc gene in human colorectal cancer progression. Br J Cancer. 1992;65(5):667–72. https://doi.org/10.1038/bjc.1992.142.
Rowley S, et al. Comparison of deoxyribonucleic acid ploidy and nuclear expressed p62 c-myc oncogene in the prognosis of colorectal cancer. World J Surg. 1990;14(4):545–50. https://doi.org/10.1007/bf01658688 (discussion 551).
Shah M, et al. A dynamic exchange of TCF3 and TCF4 transcription factors controls MYC expression in colorectal cancer cells. Cell Cycle (Georgetown, Tex). 2015;14(3):323–32. https://doi.org/10.4161/15384101.2014.980643.
Jung J, et al. Zinc finger protein 746 promotes colorectal cancer progression via c-Myc stability mediated by glycogen synthase kinase 3β and F-box and WD repeat domain-containing 7. Oncogene. 2018;37(27):3715–28. https://doi.org/10.1038/s41388-018-0225-0.
Wiegering A, et al. Targeting translation initiation bypasses signaling crosstalk mechanisms that maintain high MYC levels in colorectal cancer. Cancer Discov. 2015;5(7):768–81. https://doi.org/10.1158/2159-8290.Cd-14-1040.
Sun Y, et al. Inhibitor of DNA binding 1 (Id1) mediates stemness of colorectal cancer cells through the Id1-c-Myc-PLAC8 axis via the Wnt/β-catenin and Shh signaling pathways. Cancer Manag Res. 2019;11:6855–69. https://doi.org/10.2147/cmar.S207167.
Knight J, et al. KRASMNK inhibition sensitizes-mutant colorectal cancer to mTORC1 inhibition by reducing eIF4E phosphorylation and c-MYC expression. Cancer Discov. 2021;11(5):1228–47. https://doi.org/10.1158/2159-8290.Cd-20-0652.
Taha-Mehlitz S, et al. Adenylosuccinate lyase is oncogenic in colorectal cancer by causing mitochondrial dysfunction and independent activation of NRF2 and mTOR-MYC-axis. Theranostics. 2021;11(9):4011–29. https://doi.org/10.7150/thno.50051.
Guo Z, et al. Rictor regulates FBXW7-dependent c-Myc and cyclin E degradation in colorectal cancer cells. Biochem Biophys Res Commun. 2012;418(2):426–32. https://doi.org/10.1016/j.bbrc.2012.01.054.
Kim D, et al. Inhibition of phosphodiesterase 4D decreases the malignant properties of DLD-1 colorectal cancer cells by repressing the AKT/mTOR/Myc signaling pathway. Oncol Lett. 2019;17(3):3589–98. https://doi.org/10.3892/ol.2019.9996.
Tan J, et al. B55β-associated PP2A complex controls PDK1-directed myc signaling and modulates rapamycin sensitivity in colorectal cancer. Cancer Cell. 2010;18(5):459–71. https://doi.org/10.1016/j.ccr.2010.10.021.
Yu Y, et al. FoxO3a confers cetuximab resistance in RAS wild-type metastatic colorectal cancer through c-Myc. Oncotarget. 2016;7(49):80888–900. https://doi.org/10.18632/oncotarget.13105.
Shi Y, et al. A novel mechanism of endoplasmic reticulum stress- and c-Myc-degradation-mediated therapeutic benefits of antineurokinin-1 receptor drugs in colorectal cancer. Adv Sci (Weinh). 2021;8(21): e2101936. https://doi.org/10.1002/advs.202101936.
Liu Z, et al. The IκB family member Bcl-3 stabilizes c-Myc in colorectal cancer. J Mol Cell Biol. 2013;5(4):280–2. https://doi.org/10.1093/jmcb/mjt020.
Mansour M, et al. SATB1 and SATB2 play opposing roles in c-Myc expression and progression of colorectal cancer. Oncotarget. 2016;7(4):4993–5006. https://doi.org/10.18632/oncotarget.6651.
Shen Z, et al. SNX16 activates c-Myc signaling by inhibiting ubiquitin-mediated proteasomal degradation of eEF1A2 in colorectal cancer development. Mol Oncol. 2020;14(2):387–406. https://doi.org/10.1002/1878-0261.12626.
Coni S, et al. Blockade of EIF5A hypusination limits colorectal cancer growth by inhibiting MYC elongation. Cell Death Dis. 2020;11(12):1045. https://doi.org/10.1038/s41419-020-03174-6.
Ebert MS, Sharp PA. Roles for microRNAs in conferring robustness to biological processes. Cell. 2012;149(3):515–24. https://doi.org/10.1016/j.cell.2012.04.005.
Hayes J, Peruzzi PP, Lawler S. MicroRNAs in cancer: biomarkers, functions and therapy. Trends Mol Med. 2014;20(8):460–9. https://doi.org/10.1016/j.molmed.2014.06.005.
Adams BD, Kasinski AL, Slack FJ. Aberrant regulation and function of microRNAs in cancer. Curr Biol. 2014;24(16):R762–76. https://doi.org/10.1016/j.cub.2014.06.043.
Rupaimoole R, Slack FJ. MicroRNA therapeutics: towards a new era for the management of cancer and other diseases. Nat Rev Drug Discov. 2017;16(3):203–22. https://doi.org/10.1038/nrd.2016.246.
Wang H, et al. miR-320b suppresses cell proliferation by targeting c-Myc in human colorectal cancer cells. BMC Cancer. 2015;15:748. https://doi.org/10.1186/s12885-015-1728-5.
Chen X, et al. DNA methylation-regulated and tumor-suppressive roles of miR-487b in colorectal cancer via targeting MYC, SUZ12, and KRAS. Cancer Med. 2019;8(4):1694–709. https://doi.org/10.1002/cam4.2032.
Park W, et al. miR193a-5p mediated ZNF746 and c-Myc signaling axis is critically involved in morusin induced apoptosis in colorectal cancer cells. Cells. 2021. https://doi.org/10.3390/cells10082065.
Guo X, et al. miR-181d and c-myc-mediated inhibition of CRY2 and FBXL3 reprograms metabolism in colorectal cancer. Cell Death Dis. 2017;8(7): e2958. https://doi.org/10.1038/cddis.2017.300.
Yue C, et al. microRNA-96 promotes occurrence and progression of colorectal cancer via regulation of the AMPKα2-FTO-m6A/MYC axis. J Exp Clin Cancer Res CR. 2020;39(1):240. https://doi.org/10.1186/s13046-020-01731-7.
Zhao J, Lin H, Huang K. Mesenchymal stem cell-derived extracellular vesicles transmitting microRNA-34a-5p suppress tumorigenesis of colorectal cancer through c-MYC/DNMT3a/PTEN Axis. Mol Neurobiol. 2021. https://doi.org/10.1007/s12035-021-02431-9.
Zhang Y, et al. Cancer-associated fibroblasts-derived exosomal miR-17-5p promotes colorectal cancer aggressive phenotype by initiating a RUNX3/MYC/TGF-β1 positive feedback loop. Cancer Lett. 2020;491:22–35. https://doi.org/10.1016/j.canlet.2020.07.023.
Gu H, et al. Long non-coding RNA MILNR1 retards colorectal cancer growth by inhibiting c-Myc. Cancer Commun (London, England). 2020;40(9):456–60. https://doi.org/10.1002/cac2.12079.
Shigeyasu K, et al. The PVT1 lncRNA is a novel epigenetic enhancer of MYC, and a promising risk-stratification biomarker in colorectal cancer. Mol Cancer. 2020;19(1):155. https://doi.org/10.1186/s12943-020-01277-4.
Qiao L, et al. Knockdown of long non-coding RNA prostate cancer-associated ncRNA transcript 1 inhibits multidrug resistance and c-Myc-dependent aggressiveness in colorectal cancer Caco-2 and HT-29 cells. Mol Cell Biochem. 2018;441:99–108. https://doi.org/10.1007/s11010-017-3177-8.
Xiang J, et al. Human colorectal cancer-specific CCAT1-L lncRNA regulates long-range chromatin interactions at the MYC locus. Cell Res. 2014;24(5):513–31. https://doi.org/10.1038/cr.2014.35.
Huang W, et al. SNHG11 promotes cell proliferation in colorectal cancer by forming a positive regulatory loop with c-Myc. Biochem Biophys Res Commun. 2020;527(4):985–92. https://doi.org/10.1016/j.bbrc.2020.05.002.
Gao Q, et al. Long noncoding RNA CMPK2 promotes colorectal cancer progression by activating the FUBP3-c-Myc axis. Oncogene. 2020;39(19):3926–38. https://doi.org/10.1038/s41388-020-1266-8.
Wu R, et al. The long noncoding RNA LUCAT1 promotes colorectal cancer cell proliferation by antagonizing Nucleolin to regulate MYC expression. Cell Death Dis. 2020;11(10):908. https://doi.org/10.1038/s41419-020-03095-4.
Izumi D, et al. Colorectal cancer stem cells acquire chemoresistance through the upregulation of F-Box/WD repeat-containing protein 7 and the consequent degradation of c-Myc. Stem Cells (Dayton, Ohio). 2017;35(9):2027–36. https://doi.org/10.1002/stem.2668.
Zuo S, et al. MEG3Long non-coding RNA activated by vitamin D suppresses glycolysis in colorectal cancer promoting c-Myc degradation. Front Oncol. 2020;10:274. https://doi.org/10.3389/fonc.2020.00274.
Matsushita K, et al. Strong HLA-DR antigen expression on cancer cells relates to better prognosis of colorectal cancer patients: possible involvement of c-myc suppression by interferon-gamma in situ. Cancer Sci. 2006;97(1):57–63. https://doi.org/10.1111/j.1349-7006.2006.00137.x.
Gao Q, Wang S, Zhang Z. E3 ubiquitin ligase SMURF2 prevents colorectal cancer by reducing the stability of the YY1 protein and inhibiting the SENP1/c-myc axis. Gene Ther. 2021. https://doi.org/10.1038/s41434-021-00289-z.
Masuo T, et al. Cyclosporine A inhibits colorectal cancer proliferation probably by regulating expression levels of c-Myc, p21(WAF1/CIP1) and proliferating cell nuclear antigen. Cancer Lett. 2009;285(1):66–72. https://doi.org/10.1016/j.canlet.2009.05.001.
Zhai D, et al. Sterol regulatory element-binding protein 1 cooperates with c-Myc to promote epithelial-mesenchymal transition in colorectal cancer. Oncol Lett. 2018;15(4):5959–65. https://doi.org/10.3892/ol.2018.8058.
Martinez-Useros J, et al. UNR/CSDE1 expression is critical to maintain invasive phenotype of colorectal cancer through regulation of c-MYC and epithelial-to-mesenchymal transition. J Clin Med. 2019. https://doi.org/10.3390/jcm8040560.
Sato K, et al. Novel oncogene 5MP1 reprograms c-Myc translation initiation to drive malignant phenotypes in colorectal cancer. EBioMedicine. 2019;44:387–402. https://doi.org/10.1016/j.ebiom.2019.05.058.
Yang L, et al. Silencing or inhibition of H3K79 methyltransferase DOT1L induces cell cycle arrest by epigenetically modulating c-Myc expression in colorectal cancer. Clin Epigenet. 2019;11(1):199. https://doi.org/10.1186/s13148-019-0778-y.
Xiang S, et al. N6-methyladenosine methyltransferase METTL3 promotes colorectal cancer cell proliferation through enhancing MYC expression. Am J Transl Res. 2020;12(5):1789–806.
Hu Y, et al. Arginine methyltransferase PRMT3 promote tumorigenesis through regulating c-MYC stabilization in colorectal cancer. Gene. 2021;791: 145718. https://doi.org/10.1016/j.gene.2021.145718.
Böckelman C, et al. CIP2A overexpression is associated with c-Myc expression in colorectal cancer. Cancer Biol Ther. 2012;13(5):289–95. https://doi.org/10.4161/cbt.18922.
Denk S, et al. CIP2A regulates MYC translation (via its 5’UTR) in colorectal cancer. Int J Colorectal Dis. 2021;36(5):911–8. https://doi.org/10.1007/s00384-020-03772-y.
Li C, et al. PTPN18 promotes colorectal cancer progression by regulating the c-MYC-CDK4 axis. Genes Dis. 2021;8(6):838–48. https://doi.org/10.1016/j.gendis.2020.08.001.
Hu J, Duan W, Liu Y. Ketamine inhibits aerobic glycolysis in colorectal cancer cells by blocking the NMDA receptor-CaMK II-c-Myc pathway. Clin Exp Pharmacol Physiol. 2020;47(5):848–56. https://doi.org/10.1111/1440-1681.13248.
Shi J, Christopher RV. The mechanisms behind the therapeutic activity of BET bromodomain inhibition. Mol Cell. 2014;54(5):728–36. https://doi.org/10.1016/j.molcel.2014.05.016.
Otto C, et al. Targeting bromodomain-containing protein 4 (BRD4) inhibits MYC expression in colorectal cancer cells. Neoplasia (New York, NY). 2019;21(11):1110–20. https://doi.org/10.1016/j.neo.2019.10.003.
Zhao R, et al. BRD7 promotes cell proliferation and tumor growth through stabilization of c-Myc in colorectal cancer. Front Cell Dev Biol. 2021;9: 659392. https://doi.org/10.3389/fcell.2021.659392.
Wu T, et al. Co-inhibition of BET proteins and NF-κB as a potential therapy for colorectal cancer through synergistic inhibiting MYC and FOXM1 expressions. Cell Death Dis. 2018;9(3):315. https://doi.org/10.1038/s41419-018-0354-y.
Cao K, Tait SWG. Apoptosis and cancer: force awakens, phantom menace, or both? Int Rev Cell Mol Biol. 2018;337:135–52. https://doi.org/10.1016/bs.ircmb.2017.12.003.
Li YJ, et al. Autophagy and multidrug resistance in cancer. Chin J Cancer. 2017;36(1):52. https://doi.org/10.1186/s40880-017-0219-2.
Liu K, et al. A novel mechanism of the c-Myc/NEAT1 axis mediating colorectal cancer cell response to photodynamic therapy treatment. Front Oncol. 2021;11: 652831. https://doi.org/10.3389/fonc.2021.652831.
Saeinasab M. SNHG15 is a bifunctional MYC-regulated noncoding locus encoding a lncRNA that promotes cell proliferation, invasion and drug resistance in colorectal cancer by interacting with AIF. J Exp Clin Cancer Res. 2019. https://doi.org/10.1186/s13046-019-1169-0.
Oh E, et al. Brusatol-mediated inhibition of c-Myc increases HIF-1α degradation and causes cell death in colorectal cancer under hypoxia. Theranostics. 2017;7(14):3415–31. https://doi.org/10.7150/thno.20861.
Wu Z, et al. The BET-Bromodomain Inhibitor JQ1 synergized ABT-263 against colorectal cancer cells through suppressing c-Myc-induced miR-1271-5p expression. Biomed Pharmacother (Biomedecine & pharmacotherapie). 2017;95:1574–9. https://doi.org/10.1016/j.biopha.2017.09.087.
Guo Y, et al. Spermine synthase and MYC cooperate to maintain colorectal cancer cell survival by repressing Bim expression. Nat Commun. 2020;11(1):3243. https://doi.org/10.1038/s41467-020-17067-x.
Wu Z, et al. Dioscin inhibited glycolysis and induced cell apoptosis in colorectal cancer via promoting c-myc ubiquitination and subsequent hexokinase-2 suppression. Onco Targets Ther. 2020;13:31–44. https://doi.org/10.2147/ott.S224062.
Shi C, et al. Bromodomain and extra-terminal motif (BET) inhibition is synthetic lethal with loss of SMAD4 in colorectal cancer cells via restoring the loss of MYC repression. Oncogene. 2021;40(5):937–50. https://doi.org/10.1038/s41388-020-01580-w.
Shi H, et al. Suppression of N-myc downstream-regulated gene 2 is associated with induction of Myc in colorectal cancer and correlates closely with differentiation. Biol Pharm Bull. 2009;32(6):968–75. https://doi.org/10.1248/bpb.32.968.
Boudjadi S, et al. Integrin α1β1 expression is controlled by c-MYC in colorectal cancer cells. Oncogene. 2016;35(13):1671–8. https://doi.org/10.1038/onc.2015.231.
Groulx J, Boudjadi S, Beaulieu J. MYC regulates α6 integrin subunit expression and splicing under its pro-proliferative ITGA6A form in colorectal cancer cells. Cancers. 2018. https://doi.org/10.3390/cancers10020042.
Wu Q, et al. MNX1-AS1MYC-activated LncRNA promotes the progression of colorectal cancer by stabilizing YB1. Can Res. 2021;81(10):2636–50. https://doi.org/10.1158/0008-5472.Can-20-3747.
Ying Y, et al. Oncogenic HOXB8 is driven by MYC-regulated super-enhancer and potentiates colorectal cancer invasiveness via BACH1. Oncogene. 2020;39(5):1004–17. https://doi.org/10.1038/s41388-019-1013-1.
Lv Z, et al. Disruption of the c-Myc/miR-200b-3p/PRDX2 regulatory loop enhances tumor metastasis and chemotherapeutic resistance in colorectal cancer. J Transl Med. 2017;15(1):257. https://doi.org/10.1186/s12967-017-1357-7.
Sun W, et al. The c-Myc/miR-27b-3p/ATG10 regulatory axis regulates chemoresistance in colorectal cancer. Theranostics. 2020;10(5):1981–96. https://doi.org/10.7150/thno.37621.
Brandl L, et al. The c-MYC/NAMPT/SIRT1 feedback loop is activated in early classical and serrated route colorectal cancer and represents a therapeutic target. Med Oncol (Northwood, London, England). 2018;36(1):5. https://doi.org/10.1007/s12032-018-1225-1.
Lu W, et al. The CARM1-p300-c-Myc-Max (CPCM) transcriptional complex regulates the expression of CUL4A/4B and affects the stability of CRL4 E3 ligases in colorectal cancer. Int J Biol Sci. 2020;16(6):1071–85. https://doi.org/10.7150/ijbs.41230.
Nishizawa Y, et al. Oncogene c-Myc promotes epitranscriptome mA reader YTHDF1 expression in colorectal cancer. Oncotarget. 2018;9(7):7476–86. https://doi.org/10.18632/oncotarget.23554.
Zhang J, et al. Human UTP14a promotes colorectal cancer progression by forming a positive regulation loop with c-Myc. Cancer Lett. 2019. https://doi.org/10.1016/j.canlet.2018.10.010.
Felsher DW, Bishop JM. Reversible tumorigenesis by MYC in hematopoietic lineages. Mol Cell. 1999;4(2):199–207. https://doi.org/10.1016/S1097-2765(00)80367-6.
Soucek L, et al. Modelling Myc inhibition as a cancer therapy. Nature. 2008;455(7213):679–83. https://doi.org/10.1038/nature07260.
Sabò A, et al. Selective transcriptional regulation by Myc in cellular growth control and lymphomagenesis. Nature. 2014;511(7510):488–492. https://doi.org/10.1038/nature13537.
Desbarats L, et al. Discrimination between different E-box-binding proteins at an endogenous target gene of c-myc. Gene Dev. 1996;10(4):447–60. https://doi.org/101101/gad.10.4.447.
Blackwood EM, Eisenman RN. Max: a helix-loop-helix zipper protein that forms a sequence-specific DNA-binding complex with Myc. Science. 1991;251(4998):1211–7. https://doi.org/10.1126/science.2006410.
Blackwell TK, et al. Sequence-specific DNA binding by the c-Myc protein. Science. 1990;250(4984):1149–51. https://doi.org/10.1126/science.2251503.
Clausen DM, et al. In vitro cytotoxicity and in vivo efficacy, pharmacokinetics, and metabolism of 10074–G5, a novel small-molecule inhibitor of c-Myc/max dimerization. J Pharmacol Exp Ther. 2010;335(3):715–27. https://doi.org/10.1124/jpet.110.170555.
Weinhouse S. On respiratory impairment in cancer cells. Science. 1956;124(3215):267–9. https://doi.org/10.1126/science.124.3215.267.
Demaria M, et al. Cellular Senescence Promotes Adverse Effects of Chemotherapy and Cancer Relapse. Cancer Discov. 2017;7(2):165–176. https://doi.org/10.1158/2159-8290.Cd-16-0241.
Dörr JR, et al. Synthetic lethal metabolic targeting of cellular senescence in cancer therapy. Nature. 2013;501(7467):421–5. https://doi.org/10.1038/nature12437.
Ren T, et al. MCU-dependent mitochondrial Ca2+ inhibits NAD+/SIRT3/SOD2 pathway to promote ROS production and metastasis of HCC cells. Oncogene. 2017;36(42):5897–909. https://doi.org/10.1038/onc.2017.167.
Fu L, et al. SIRT4 inhibits malignancy progression of NSCLCs, through mitochondrial dynamics mediated by the ERK-Drp1 pathway. Oncogene. 2017;36(19):2724–36. https://doi.org/10.1038/onc.2016.425.
Guerra F, Arbini AA, Moro L. Mitochondria and cancer chemoresistance. Biochimica et Biophysica Acta (BBA) Bioenergetics. 2017;1858(8):686–99. https://doi.org/10.1016/j.bbabio.2017.01.012.
Lheureux S, et al. Molecularly targeted therapies in cancer: a guide for the nuclear medicine physician. Eur J Nucl Med Mol Imaging. 2017;44(1):41–54. https://doi.org/10.1007/s00259-017-3695-3.
Vasudev NS, Reynolds AR. Anti-angiogenic therapy for cancer: current progress, unresolved questions and future directions. Angiogenesis. 2014;17(3):471–94. https://doi.org/10.1007/s10456-014-9420-y.
Acknowledgements
Many thanks for all the authors of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Author information
Authors and Affiliations
Contributions
Agreeing to be accountable for all aspects of the work, all authors participated in perioperative literature search and conclusion, manuscript drafting or revising, and decision making to which journal the manuscript would be submitted.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tan, L., Peng, D. & Cheng, Y. Significant position of C-myc in colorectal cancer: a promising therapeutic target. Clin Transl Oncol 24, 2295–2304 (2022). https://doi.org/10.1007/s12094-022-02910-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-022-02910-y