Abstract
Background
Cuproptosis is a recently discovered mechanism of programmed cell death caused by intracellular aggregation of mitochondrial lipoylated proteins and destabilization of iron-sulfur proteins triggered by copper. Hepatocellular carcinoma (HCC) is a common malignant tumor with a poor prognosis. We aimed to predict the survival of patients with HCC using the cuproptosis-related gene (CRG) expression.
Methods
We analyzed the expression, methylation, and mutation status of CRGs in 538 HCC patients and correlated the date with clinical prognosis. HCC patients were divided into two clusters based on their CRG expression. The relationship between CRGs, risk genes, and the immune microenvironment was analyzed using the CIBERSORT algorithm and the single-cell data analysis method. A cuproptosis risk model was constructed according to the five risk genes using the LASSO COX method. To facilitate the clinical applicability of the proposed risk model, we constructed a nomogram and conducted an antineoplastic drug sensitivity analysis.
Results
Our results suggest that the expression levels of CRGs in HCC are regulated by methylation. The prognoses were significantly different between the patients of the two clusters. The prognostic risk score positively correlated with memory T cell activation and negatively correlated with natural killer (NK) and regulatory T cell activation.
Conclusion
Our findings indicate the involvement of CRG regulation in HCC and provide new insights into prognosis assessment. Drug sensitivity analysis predicted drug candidates for the treatment of patients with different HCC subtypes.
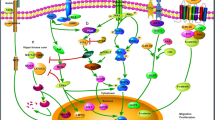
Graphical abstract







Similar content being viewed by others
Availability of data and material
The datasets analyzed for this study can be found in the TCGA (http://www.cancer.gov/tcga) and GEO (https://www.ncbi.nlm.nih.gov/geo).
Code availability
The code data are available from the corresponding author on request.
References
Villanueva A. Hepatocellular carcinoma. N Engl J Med 2019;380(15):1450–1462
Piñero F, Dirchwolf M, Pessôa MG. Biomarkers in hepatocellular carcinoma: diagnosis, prognosis and treatment response assessment. Cells 2020;9(6):1370
Tsvetkov P, et al. Copper induces cell death by targeting lipoylated TCA cycle proteins. Science 2022;375(6586):1254–1261
Bock FJ, Tait SWG. Mitochondria as multifaceted regulators of cell death. Nat Rev Mol Cell Biol 2020;21(2):85–100
Zischka H, et al. Liver mitochondrial membrane crosslinking and destruction in a rat model of Wilson disease. J Clin Invest 2011;121(4):1508–1518
Oliveri V. Selective targeting of cancer cells by copper ionophores: an overview. Front Mol Biosci 2022;9: 841814
Tang D, Chen X, Kroemer G. Cuproptosis: a copper-triggered modality of mitochondrial cell death. Cell Res 2022;32:417–418
Chandrashekar DS, et al. UALCAN: a portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia 2017;19(8):649–658
Papatheodorou I, et al. Expression Atlas update: from tissues to single cells. Nucleic Acids Res 2020;48(D1):D77–D83
Menyhárt O, Nagy Á, Győrffy B. Determining consistent prognostic biomarkers of overall survival and vascular invasion in hepatocellular carcinoma. R Soc Open Sci 2018;5(12):181006
Newman AM, et al. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods 2015;12(5):453–457
Rooney MS, et al. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 2015;160(1–2):48–61
Iasonos A, et al. How to build and interpret a nomogram for cancer prognosis. J Clin Oncol 2008;26(8):1364–1370
Xu LX, et al. Genomic and transcriptional heterogeneity of multifocal hepatocellular carcinoma. Ann Oncol 2019;30(6):990–997
Aubert L, et al. Copper bioavailability is a KRAS-specific vulnerability in colorectal cancer. Nat Commun 2020;11(1):3701
Michniewicz F, et al. Copper: an intracellular achilles’ heel allowing the targeting of epigenetics, kinase pathways, and cell metabolism in cancer therapeutics. ChemMedChem 2021;16(15):2315–2329
Chen F, et al. Serum copper and zinc levels and the risk of oral cancer: A new insight based on large-scale case-control study. Oral Dis 2019;25(1):80–86
Saleh SAK, et al. Serum levels of selenium, zinc, copper, manganese, and iron in prostate cancer patients. Curr Urol 2020;14(1):44–49
Mossmann D, Park S, Hall MN. mTOR signalling and cellular metabolism are mutual determinants in cancer. Nat Rev Cancer 2018;18(12):744–757
Yu M, et al. PLCγ-dependent mTOR signalling controls IL-7-mediated early B cell development. Nat Commun 2017;8(1):1457
Percival SS. Copper and immunity. Am J Clin Nutr 1998;67(5 Suppl):1064s–1068s
Djoko KY, et al. The role of copper and zinc toxicity in innate immune defense against bacterial pathogens. J Biol Chem 2015;290(31):18954–18961
Zhang Z, et al. Gasdermin E suppresses tumour growth by activating anti-tumour immunity. Nature 2020;579(7799):415–420
Vivier E, et al. Targeting natural killer cells and natural killer T cells in cancer. Nat Rev Immunol 2012;12(4):239–252
Kondratova M, et al. A multiscale signalling network map of innate immune response in cancer reveals cell heterogeneity signatures. Nat Commun 2019;10(1):4808
Ludewig B, et al. Protective antiviral cytotoxic T cell memory is most efficiently maintained by restimulation via dendritic cells. J Immunol 1999;163(4):1839–1844
Wherry EJ, et al. Lineage relationship and protective immunity of memory CD8 T cell subsets. Nat Immunol 2003;4(3):225–234
Klebanoff CA, et al. Central memory self/tumor-reactive CD8+ T cells confer superior antitumor immunity compared with effector memory T cells. Proc Natl Acad Sci USA 2005;102(27):9571–9576
van Panhuys N, et al. Effector lymphoid tissue and its crucial role in protective immunity. Trends Immunol 2005;26(5):242–247
Cao J, et al. Screening and identifying immune-related cells and genes in the tumor microenvironment of bladder urothelial carcinoma: based on TCGA Database and Bioinformatics. Front Oncol 2019;9:1533
Saito T, et al. Two FOXP3(+)CD4(+) T cell subpopulations distinctly control the prognosis of colorectal cancers. Nat Med 2016;22(6):679–684
Funding
This study was supported by grants from the Clinical Research Plan of SHDC (SHDC2020CR4018), the National Natural Science Foundation of China (81902907 and 81874182) and Shanghai Pujiang Program (2019PJD008).
Author information
Authors and Affiliations
Contributions
WY, WL, ZT: contributed to the conception of the study; WY, ZY, WL: performed the experiment; XW, ZN, ZY: contributed significantly to analysis and manuscript preparation; WY, ZJ: performed the data analyses and wrote the manuscript; XW, ZW: helped perform the analysis with constructive discussions.
Corresponding authors
Ethics declarations
Conflict of interest
All authors declare that there are no conficts of interest.
Consent for publication
All authors approved the manuscript for publication.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, Y., Zhang, Y., Wang, L. et al. Development and experimental verification of a prognosis model for cuproptosis-related subtypes in HCC. Hepatol Int 16, 1435–1447 (2022). https://doi.org/10.1007/s12072-022-10381-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12072-022-10381-0