Abstract
Ginkgo biloba is utilized as food, medicine, wood, and street trees among other things. The objective of this study was to develop a loop-mediated isothermal amplification (LAMP) assay for gender distinction of G. biloba. Male-specific SCAR gene can be utilized to identify G. biloba gender using LAMP. The optimized LAMP conditions, temperature 60 °C, 2-mM MgSO4, and [F3/B3]:[FIP/BIP] primer ratio of 1:4 were selected as final conditions. The G. biloba SCAR LAMP displayed a sensitivity of 10 ng when amplified by concentration under the optimum conditions. Additionally, it demonstrated a particular response in male with SYBR Green I in LAMP analysis that can be a more powerful tool for field and scale-up applications. Our work represents a first attempt to identify G. biloba gender using LAMP and offers an efficient and reliable tool for roadside landscaping.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ginkgo biloba, a dioecious perennial gymnosperm (65 million years ago), is an extant representative of the order Ginkgoales in the Tertiary Period [1, 2]. It is known as the “living fossil.” In terms of practical applicability, male and female G. biloba have different values [3,4,5]. Male trees are typically utilized as street and landscaping trees. Female plants are not desirable for this purpose since their seeds emit a foul odor [6,7,8]. Although ginkgo leaf extracts might provide some health benefits, recent studies have revealed that components of ginkgo seed have the potential to cause severe allergic reactions and carcinogenic activities [8,9,10,11,12]. Amplification of nucleic acid is the most useful method in nucleic acid detection. Nucleic acid amplification is being applied to detecting and diagnosing bacteria and viruses having nucleic acids [13,14,15]. Gender distinction of G. biloba using sequence characterized amplified region (SCAR) markers by PCR application has been reported [16]. However, PCR-based approach has significant limitations such as requirement of sophisticated equipment and up to a 2–4 h of process [17, 18].
Loop-mediated isothermal amplification (LAMP) method is a relatively recent and quick molecular biology method that uses four to six distinct primers to bind to eight target areas, resulting in excellent sensitivity and specificity [19, 20]. For in vitro enzymatic DNA synthesis under isothermal circumstances, LAMP employs autocycling strand displacement [21, 22]. Bst DNA polymerase from Geobacillus stearothermophilus is used in most applications, although comparable thermophilic DNA polymerases from other bacterial sources are also used [23, 24]. LAMP technology is not limited to targets composed of DNA, but by combining a reverse transcription step. LAMP procedure has recently been used to identify RNA viruses, such as SARS coronavirus and influenza virus [25, 26]. Different targets, including bacteria, viroid, and fungi, can be detect using LAMP assay technique [25, 27, 28]. LAMP assay can be used to identify target genetic material in the field and of rapidly amplify it at the same temperature without the need for a separate temperature control apparatus [29]. Also, since LAMP uses fluorescent reagents, additional steps such as gel electrophoresis are not required. Despite its popularity in genetic verification, LAMP possesses “inherent drawback,” such as non-specific primer annealing. This drawback has later paved the way for the use of several innovated nanoprobes, such as DNA-barcode [26] and multiplex real-time LAMP [30].
The objective of this study was to develop a LAMP assay for gender distinction of G. biloba. G. biloba male-specific primers were designed to amplify DNA-based male-determinant, 5′-truncated SCAR fragment. Here, we successfully optimized LAMP conditions for distinction of male trees (Fig. 1) [16, 31]. Gender determination of G. biloba was carried out in this study for the first-time using LAMP technology.
Materials and Methods
Plant Materials of G. biloba and Leaf Genomic DNA Isolation
G. biloba grew under natural condition in the field at Chungbuk national University. After determining the gender of the plant, individual leaf samples were collected and stored in a − 80 °C deep-freezer. To identify male and female trees, two G. biloba trees planted in Chungbuk National University (36° 37′ 42.9 N, 127° 27′ 27.3 E) were used. Information of all trees used in the study is presented in Supplementary Fig. 1. Genomic DNA (gDNA) of each tree leaf samples was extracted with a DNeasy Plant Mini Kit (50) (Qiagen, Germany) according to the manufacturer’s protocol [32]. Concentration of extracted gDNA was measured using a Nanodrop 2000 (Thermo Fisher Scientific, USA).
PCR Amplification Using SCAR-GBM Primers and Alignment of DNA Amplified Region
To amplify gDNA of G. biloba, SCAR-GBM primer was used for PCR amplification. PCR were performed in a 50 μL of reaction volume containing 1 × Ex Taq buffer (20-mM Tris–HCl, pH 8.3, 50-mM KCl, 2 mM MgSO4, 10-mM(NH4)2SO4, 0.1% Triton X-100), 0.2-mM dNTPs, 0.4 uM of each primer, 1.25 U of Taq DNA polymerase, and 10 ng of DNA library. The PCR program was as follows: an initial pre-denaturation step at 95 °C for 4 min, three cycles of denaturation at 95 °C for 30 s, annealing at 55 °C for 30 s, extension 72 °C for 50 s, followed by 35 cycles, and a final post-extension step at 72 °C for 7 min. Results of PCR were analyzed using agarose gel electrophoresis. PCR amplification product using SCAR-GBM primer was cloned into T-Blunt™ using a PCR cloning kit (Solgent, Korea). T-blunt vector, PCR product, and T-blunt buffer were incubated at RT for 5 min. The mixture was transformed into E. coli DH5α (Solgent, Korea) by heat shock method. After that, the mixture was mixed with 900-μL LB broth and incubated in a 37 °C shaking incubator for 40 min. The product was spread onto LB agar plates containing ampicillin (50 μg/mL), kanamycin (50 μg/mL), IPTG (0.2 mM), and X-Gal (40 μg/mL) and then incubated at 37 °C in an incubator overnight. White colonies were sequenced by Solgent (Daejeon, Korea) using M13-20F and M13-20R primers. Sequencing data were aligned using Clustal Omega software (https://www.ebi.ac.uk/Tools/msa/clustalo/).
LAMP Primer Design and PCR Amplification of LAMP Primer
LAMP primer sets were designed for determination of G. biloba gender using PrimerExplorer V5 software (http://primerexplorer.jp/lampv5e/index.html). LAMP primer set was selected from a gender heterologous sequence region to efficiently distinguish G. biloba gender. Detailed sequence information of each primer is listed in Table 1. LAMP primers included two outer primers (F3 and B3), two inner primers (forward inner primer; FIP and backward inner primer; BIP), and one loop primer (loop backward, LB). All primers were synthesized by Bioneer (Korea). To confirm the LAMP target, outer primers (F3 and B3) were used. Results of PCR were analyzed using 2% agarose gel electrophoresis.
Optimization of LAMP Conditions
The basic LAMP reaction mixture contained 2.5-μL 10X isothermal amplification buffer [1X contained 20-mM Tris–HCl, 10-mM (NH4)2SO4, 150-mM KCl, 2-mM MgSO4, 0.1% Tween® 20], 1 μL each outer primer (F3, B3; 5 uM), inner primer (FIP, BIP; 40 uM), and loop primer (LF, LB; 10 uM), 2-μL 10 mM dNTP, 1.5-μL 100-mM MgSO4 (total 8 mM), and up to 23-μL distilled water. After addition of 1-μL dsDNA template, the mixture was heated at 95 °C for 5 min and then cooled to room temperature for 5 min. Next, 1 μL (eight units) of Bst polymerase was added to the mixture. LAMP reaction was performed at 60–65 °C for 60 min. It was then heated to 80 °C for 10 min for termination.
Optimized LAMP reactions proceeded with a total volume of 25 μL containing the following: 2.5-μL 10 × Isothermal amplification buffer, 1.4-mM dNTPs, extra 6-mM MgSO4, 1.6-uM inner primer (FIP and BIP), 0.4-uM outer primer (F3 and B3), 0.4-uM loop primer (LB), 0.32 U of Bst 2.0 polymerase, and 100 ng of DNA. To identify optimal LAMP conditions, various temperature conditions (60, 61, 62, 63, 64, 65 °C) were firstly analyzed. Various extra MgSO4 concentration conditions (0, 2, 4, 6, and 8 mM) were then analyzed. Next, different primer ratios ([F3/B3]:[FIP/BIP], 1:2, 1:4, and 1:8) were analyzed. After that, different amounts (0, 10, 50, 100, 200, and 500 ng) of template were analyzed.
Visual Inspection of LAMP Products Using SYBR Green I
LAMP results were visualized by naked eyes and determined by adding SYBR Green I to reaction tubes. The immediate observation of the color change from orange to green can be achieved. The final reaction mixture of the LAMP product's fluorescent intensity was measured using a SpectraMax M2 Mode Microplate Reader (Molecular Devices, USA) at a wavelength of 497–515 nm. The final reaction mixture with a volume of 25 μL was diluted to a volume of 200 μL using nuclease-free water. The resulting diluted mixture was utilized to record fluorescence.
Results
Gender Determinant of G. biloba, LAMP Assay, and Development of DNA-Based Male Determinant
An interesting exception was SCAR-GBM, which amplified products of similar sizes from male and female G. biloba individuals, providing false-positive value (Fig. 2a, Supplementary Fig. 2 and Supplementary Table 2). Note that, by developing male-specific determinant, it is therefore sufficient to discriminate male G. biloba. The amplicon formed with the SCAR-GBM primer showed a difference according to the gender of G. biloba. Thus, sequences of amplicons were compared. Comparison of sequences of SCAR-GBM PCR amplicons from male and female gDNAs of G. biloba consistently detected the 5’ truncated SCAR fragments (SCAR fragment 485–676) as a DNA-based male determinant. In order to evaluate the specific determinant of male G. biloba, a new primer set was designed (Fig. 2a). Sequence details are shown in Table 1. F3/B3 primers amplified truncated SCAR fragments only from male G. biloba, but not from female G. biloba, which were contributed to LAMP amplification (Supplementary Fig. 3). In the present study, LAMP for the truncated SCAR fragment was conducted using a set of four basic primers (F3, B3, FIP, BIP) and loop backward (LB) primer. As shown in Fig. 2b, typical smear pattern of the amplified LAMP products was only observed for target-positive samples.
Assessment of Ginkgo biloba SCAR-GBM and LAMP detection of male Ginkgo biloba. a Alignment of male and female SCAR-GBM sequences. Multiple sequences alignment of G. biloba amplicons using SCAR-GBM primers (see square bracket, []) are shown. A comparison of sequences derived from ten representative individual clones showed that males matched 100% with the reference sequence. However, females showed different results for each of five samples analyzed. Positions of male-specific LAMP target and primers are indicated by colors: F3 and B3, yellow; F2 and F1c, green; B2 and B1c, blue; and loop backward (LB), purple. b Agarose gel electrophoresis of LAMP products based on presence or absence of gDNA of male G. biloba. Lane M, DNA Ladder (100–10,000 bp); Lanes 1 and 3, LAMP amplification without template DNA; Lanes 2 and 4, LAMP amplification with template DNA
Optimization of LAMP Reaction
To determine optimum LAMP conditions, LAMP reactions were performed by varying the temperature, MgSO4 concentration, primer ratio, and template amount to determine optimal amplification conditions. LAMP reaction for male G. biloba was amplified at temperatures ranging from 60 to 65 °C for 1 h. Band patterns revealed a ladder of bands on an agarose gel. Bands became clearer at lower reaction temperature. The optimal temperature of LAMP reaction was set at 60 °C (Fig. 3a–f). LAMP reactions were then performed with various MgSO4 concentrations ranging from 2 to 10 mM. High efficiency of LAMP was observed when 2-mM MgSO4 was used (Fig. 3g). After setting the optimal concentration of MgSO4 at 2 mM, different ratios (1:2, 1:4, and 1:8) of primers [F3/B3]:[FIP/BIP] were then analyzed. The band pattern on the agarose gel was the weakest at a primer ratio of 1:2. However, the ladder shape was clearly seen when the primer ratio was 1:4 or 1:8 (Fig. 3h). Thus, optimal temperature, ion concentration, and primer ratio were temperature of 60 °C, 2-mM MgSO4, and primer ratio of 1:4.
Optimization of LAMP conditions for amplification of male gDNA at different temperatures, MgSO4 concentration, and primer ratio. Results of 1% agarose gel electrophoresis of G. biloba gDNA LAMP detection using different reaction temperatures: a 60 °C, b 61 °C, c 62 °C, d 63 °C, e 64 °C, and f 65 °C. Lane M, DNA Ladder (100–10,000 bp); Lanes 1 and 3, LAMP amplification of male gDNA; Lanes 2 and 4, negative control, non-template contained. g. Results of 1% agarose gel electrophoresis of G. biloba gDNA LAMP detection using different concentrations of MgSO4. Lane M, DNA Ladder (100–10,000 bp); Lanes 1, 3, 5, 7, and 9, LAMP amplification of male gDNA with different concentrations (2, 4, 6, 8, and 10 mM) of MgSO4. Lanes 2, 4, 6, 8, and 10, negative control without containing gDNA template. h. Results of 1% agarose gel electrophoresis of G. biloba gDNA LAMP detection using different primer ratios. Lane M, DNA Ladder (100–10,000 bp); Lanes 1, 3, and 5, LAMP amplification of male gDNA using different ratios of primers (F3/B3: FIP/BIP = 1:2, 1:4, and 1:8). Lane 2, 4, and 6, negative control, containing no gDNA template
Sensitivity Test G. biloba LAMP Assay
We attempted to amplify endpoint-diluted DNA extracted from G. biloba in order to establish the detection limit of the optimized LAMP method. To measure the limit of optimized LAMP conditions, we used G. biloba gDNA at various concentrations (0–500 ng). LAMP reaction was carried out under optimized conditions. Amplification result was analyzed by 1% agarose gel electrophoresis. Electrophoresis bands showed equal intensities when template amounts were 500 ng and 200 ng. Band strength decreased when the amount of template was increased. It was found that amplification was feasible when at least 10 ng of the template was used. When template amount was less than 10 ng, smear-shaped bands were observed (Fig. 4a). Using Image Lab software (Version 6.1.0), the strength of the band amplified by the LAMP assay was measured. The band appeared when 10 ng or more of the template was used (Fig. 4b). Thus, the LAMP assay could be used to identify the gender of G. biloba using a template at a concentration of 10 ng or more.
LAMP detection results using different concentrations of male Ginkgo biloba gDNA. a. Results of 1% agarose gel electrophoresis of G. biloba gDNA LAMP detection using different gDNA concentrations. Lane M, DNA Ladder (100–10,000 bp); Lane 1, 500 ng/μL; Lane 2, 200 ng/μL; Lane 3, 100 ng/μL; Lane 4, 50 ng/μL; Lane 5, 10 ng/μL; and Lane 6, 0 ng/μL. b. Band volume quantity of agarose gel electrophoresis of G. biloba gDNA LAMP detection
Specific Amplification and Visual Inspection of LAMP Products Using SYBR Green I
The specificity of LAMP assay was examined using gDNAs extracted from G. biloba and control roadside trees. G. biloba male-specific positive amplification signal was exclusively present in male G. biloba samples. Female G. biloba and other species did not exhibit an amplification signal (Fig. 5a). Addition of 1 μL of SYBR green I to the reaction tube after LAMP reaction enabled naked eye inspection (Fig. 5b) or UV irradiation (Fig. 5c). Besides, only male tree tube changed color when the tube was exposed to UV light. Its fluorescence was then quantitatively evaluated. Using 15 male and 15 female G. biloba street trees (Supplementary Fig. 4) as samples, the specificity and accuracy of the LAMP assay were examined. Fourteen out of fifteen male trees produced positive results using the aforementioned optimal LAMP conditions. However, one out of fifteen female trees produced positive finding as a false negative (Fig. 6). Investigations revealed that the specificity and sensitivity were 93.3%. These results were found to be highly consistent with Cohen’s kappa value of 0.92 (see Table 2).
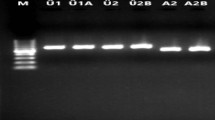
Visual inspection LAMP products using SYBR Green I. a Gel electrophoresis of LAMP for G. biloba. Only males were particularly amplified by agarose electrophoresis. b Naked eye inspection of LAMP products using SYBR Green I. SYBR Green I treatment only caused male tree samples to turn green, indicating a positive LAMP reaction. On the other hand, the orange color indicated a negative LAMP reaction. c and d Fluorometric values of LAMP assay obtained using SYBR Green I dye plotted for different samples. Lane M, DNA Ladder (100–10,000 bp); Lanes 1 and 2, LAMP amplification of male gDNA; Lane 3 and 4, LAMP amplification of female gDNA; Lane C1 and C2, LAMP amplification using gDNAs of control street trees (Prunus subg. Cerasus)
Large-scale gender distinguishing test by LAMP assay using SYBR Green I. Results of LAMP assay detection of G. biloba street trees. LAMP reactions were inspected by adding SYBR Green I dye. a LAMP detection of different male samples of G. biloba (1–15). b LAMP detection of different female samples of G. biloba (1–15)
Discussion
SCAR markers are gDNA fragments that are identified by PCR amplification with pairs of specific oligonucleotide primers [33, 34]. SCAR-GBM primers were derived from G. biloba SCAR markers [13]. They are expected to be straightforward, reliable, and easy to detect male G. biloba. However, contrary to our expectation, the PCR using SCAR-GBM primers produced amplicons in both genders tested. It is important to note that SCAR-GBM PCR occurrences are not in agreement with previous studies [16], resulting in loss of specificity and discriminative ability. For verifying the false-positive result of SCAR-GBM, sequence analysis was conducted for male and female PCR amplicons. The sequence of the amplicon amplified with the SCAR-GBM primer was identical to the male, but varied in size in the female, indicating that SCAR-GBM produced ambiguous polymorphic fragments in female G. biloba (see Supplementary Table 2).
An interesting exception was that SCAR-GBM, which amplified products of similar sizes from male and female G. biloba individuals, provided a false-positive value. Note that, by developing male-specific determinant, it is sufficient to discriminate male G. biloba. LAMP relies on an autocycling strand displacement of DNA amplification using four to six primers [35]. PCR and LAMP amplification proved that the 5’-truncated SCAR and LAMP primers could be used for more reliable and accurate gender-specific discrimination between male and female for G. biloba.
We confirmed that adding SYBR green I after the LAMP reaction could visually evaluate LAMP-based ginkgo gender distinction. When SYBR green I was added, development of green color indicated a positive response, while orange indicated a negative response. To enhance field applicability, simple gDNA extraction from plants and a ready-to-use LAMP mixture should be incorporated. High sensitivity and specificity were observed when samples of 30 G. biloba street trees were analyzed. Through this study, it will be possible to apply LAMP to distinguish the gender of G. biloba.
Concluding Remarks
Our new approach could provide a remedy for gender distinction difficulties of G. biloba. As unique sequence regions, 5’-truncated SCAR fragments of male G. biloba were used to design primers for specific detection. PCR successfully amplified single monomorphic SCAR fragments of 191 bp, indicating that 5’-truncated SCAR fragment had the same genetic identity. Moreover, the first effort of highly sensitive and specific LAMP assay for G. biloba was reported. The LAMP reaction was evaluated by agarose gel electrophoresis, UV inspection, and naked eye with SYBR Green I visualization. Due to its relatively shorten performance time, isothermal amplification condition, and better visible judgment of positive resultant, LAMP can be a more powerful tool for field and scale-up applications, thereby improving both distinction of G. biloba gender and prevention of harmful phytochemicals from G. biloba seeds.
Abbreviations
- BIP:
-
Backward inner primer
- FIP:
-
Forward inner primer
- G. biloba :
-
Ginkgo biloba
- gDNA:
-
Genomic DNA
- LAMP:
-
Loop-mediated isothermal amplification
- LB:
-
Loop backward primers
- LF:
-
Loop forward primer
- NPV:
-
Negative predictive value
- PPV:
-
Positive predictive value
- SCAR:
-
Sequence characterized amplified region
- UV:
-
Ultraviolet
References
Jacobs, B. P., & Browner, W. S. (2000). Ginkgo biloba: A living fossil. The American Journal of Medicine, 108, 341–342.
Hsieh, L. (1992). Origin and distribution of Ginkgo biloba. The Forestry Chronicle, 68, 612–613.
Singh, B., Kaur, P., Singh, R., & Ahuja, P. (2008). Biology and chemistry of Ginkgo biloba. Fitoterapia, 79, 401–418.
Major, R. T. (1967). The ginkgo, the most ancient living tree: The resistance of Ginkgo biloba L. to pests accounts in part for the longevity of this species. Science, 157, 1270–1273.
Šmarda, P., Horová, L., Knápek, O., Dieck, H., Dieck, M., Ražná, K., Hrubík, P., Orlóci, L., Papp, L., & Veselá, K. (2018). Multiple haploids, triploids, and tetraploids found in modern-day “living fossil” Ginkgo biloba. Horticulture Research. https://doi.org/10.1038/s41438-018-0055-9
Shen, D., Labreche, F., Wu, C., Fan, G., Li, T., Shi, H., & Ye, C. (2021). Preparation and aroma analysis of flavonoid-rich ginkgo seeds fermented using rice wine starter. Food Bioscience, 44, 101459.
Boateng, I. D., Yang, X.-M., Tahany, A. A., & Li, Y.-Y. (2021). Drying methods affect organoleptic and physicochemical properties of rehydrated ginkgo seed slices. Industrial Crops and Products, 160, 113166.
Tomova, T., Slavova, I., Tomov, D., Kirova, G., & Argirova, M. D. (2021). Ginkgo biloba seeds—An environmental pollutant or a functional food. Horticulturae, 7, 218.
Wang, H.-Y., & Zhang, Y.-Q. (2019). The main active constituents and detoxification process of Ginkgo biloba seeds and their potential use in functional health foods. Journal of Food Composition and Analysis, 83, 103247.
Mei, N., Guo, X., Ren, Z., Kobayashi, D., Wada, K., & Guo, L. (2017). Review of Ginkgo biloba-induced toxicity, from experimental studies to human case reports. Journal of Environmental Science and Health, Part C, 35, 1–28.
Koch, E., Jaggy, H., & Chatterjee, S. (2000). Evidence for immunotoxic effects of crude Ginkgo biloba L. leaf extracts using the popliteal lymph node assay in the mouse. International Journal of Immunopharmacology, 22, 229–236.
Dehiri, M., Diafat, A., Fatmi, W., Bouaziz, F., Khalil, R., & Bahloul, A. (2022). Toxicity evaluation of Algerian Peganum harmala seed hydromethanolic extract. Toxicology and Environmental Health Sciences, 14(4), 351–359. https://doi.org/10.1007/s13530-022-00149-2
Wan, Z., Umer, M., Lobino, M., Thiel, D., Nguyen, N.-T., Trinchi, A., Shiddiky, M. J., Gao, Y., & Li, Q. (2020). Laser induced self-N-doped porous graphene as an electrochemical biosensor for femtomolar miRNA detection. Carbon, 163, 385–394.
Perumal, V., Saheed, M. S. M., Mohamed, N. M., Saheed, M. S. M., Murthe, S. S., Gopinath, S. C., & Chiu, J.-M. (2018). Gold nanorod embedded novel 3D graphene nanocomposite for selective bio-capture in rapid detection of Mycobacterium tuberculosis. Biosensors and Bioelectronics, 116, 116–122.
Low, S. S., Loh, H.-S., Boey, J. S., Khiew, P. S., Chiu, W. S., & Tan, M. T. (2017). Sensitivity enhancement of graphene/zinc oxide nanocomposite-based electrochemical impedance genosensor for single stranded RNA detection. Biosensors and Bioelectronics, 94, 365–373.
Hong, Y. P., & Lee, J. W. (2016). Development of SCAR marker for identifying male trees of ginkgo biloba using multiplex PCR. Journal of Korean Society of Forest Science, 105, 422–428.
Lorenz, T. C. (2012). Polymerase chain reaction: Basic protocol plus troubleshooting and optimization strategies. JoVE Journal of Visualized Experiments, 63, e3998.
Kim, S. Y., Lee, J.-P., Shin, W.-R., Oh, I.-H., Ahn, J.-Y., & Kim, Y.-H. (2022). Cardiac biomarkers and detection methods for myocardial infarction. Molecular & Cellular Toxicology, 18(4), 443–455. https://doi.org/10.1007/s13273-022-00287-1
Lin, P., Wang, H., Cheng, Y., Song, S., Sun, Y., Zhang, M., Guo, L., Yi, L., Tong, M., & Cao, Z. (2018). Loop-mediated isothermal amplification-single nucleotide polymorphism analysis for detection and differentiation of wild-type and vaccine strains of mink enteritis virus. Scientific Reports, 8, 1–8.
Yuan, D., Kong, J., Li, X., Fang, X., & Chen, Q. (2018). Colorimetric LAMP microfluidic chip for detecting three allergens: Peanut, sesame and soybean. Scientific Reports, 8, 1–8.
Singh, M. D., Singh, H., Singh, N. K., Singh, N. K., Sood, N. K., & Rath, S. S. (2019). Evaluation of a loop-mediated isothermal amplification technique for the rapid visual detection of Hepatozoon canis infection. Acta Parasitologica, 65, 151.
Lee, J.-E., Mun, H., Kim, S.-R., Kim, M.-G., Chang, J.-Y., & Shim, W.-B. (2020). A colorimetric Loop-mediated isothermal amplification (LAMP) assay based on HRP-mimicking molecular beacon for the rapid detection of Vibrio parahaemolyticus. Biosensors and Bioelectronics, 151, 111968.
Ma, Y., Zhang, B., Wang, M., Ou, Y., Wang, J., & Li, S. (2016). Enhancement of polymerase activity of the large fragment in DNA polymerase I from Geobacillus stearothermophilus by site-directed mutagenesis at the active site. BioMed Research International. https://doi.org/10.1155/2016/2906484
Oscorbin, I. P., Belousova, E. A., Boyarskikh, U. A., Zakabunin, A. I., Khrapov, E. A., & Filipenko, M. L. (2017). Derivatives of Bst-like Gss-polymerase with improved processivity and inhibitor tolerance. Nucleic acids Research, 45, 9595–9610.
Lee, S. H., Lee, S.-H., Won, K., Kim, M.-S., Ryu, H., Kim, Y.-H., & Ahn, J.-Y. (2020). Loop-mediated isothermal amplification (LAMP)-based turn on fluorescent paper (ToFP) device for detecting Rosellinia necatrix. Journal of Biomedical Nanotechnology, 16, 166–178.
Lee, S. H., Ahn, G., Kim, M.-S., Jeong, O. C., Lee, J. H., Kwon, H. G., Kim, Y.-H., & Ahn, J.-Y. (2018). Poly-adenine-coupled LAMP barcoding to detect apple scar skin viroid. ACS Combinatorial Science, 20, 472–481.
Ahn, G., Lee, S. H., Song, M.-S., Han, B.-K., Kim, Y.-H., & Ahn, J.-Y. (2021). JEV-nanobarcode and colorimetric reverse transcription loop-mediated isothermal amplification (cRT-LAMP). Microchimica Acta, 188, 1–11.
Ahn, G., Lee, S., Lee, S. H., Baek, Y. H., Song, M.-S., Kim, Y.-H., & Ahn, J.-Y. (2021). Zika virus lateral flow assays using reverse transcription-loop-mediated isothermal amplification. RSC Advances, 11, 17800–17808.
Foo, P. C., Nurul Najian, A., Muhamad, N. A., Ahamad, M., Mohamed, M., Yean Yean, C., & Lim, B. H. (2020). Loop-mediated isothermal amplification (LAMP) reaction as viable PCR substitute for diagnostic applications: A comparative analysis study of LAMP, conventional PCR, nested PCR (nPCR) and real-time PCR (qPCR) based on Entamoeba histolytica DNA derived from faecal sample. BMC Biotechnology, 20, 1–15.
Gadkar, V. J., Goldfarb, D. M., Gantt, S., & Tilley, P. A. (2018). Real-time detection and monitoring of loop mediated amplification (LAMP) reaction using self-quenching and de-quenching fluorogenic probes. Scientific Reports, 8, 1–10.
Longdou, L., Quibai, X., Meili, J., Wujun, G., & Ruili, L. (2006) Identification of RAPD makers linked to the male sex in Dioecious ginkgo biloba L. Analele Stiintifice ale Universitatii" Alexandru Ioan Cuza" din Iasi Sec II a Genetica si Biologie Moleculara 7.
Oh, T. w., Cho, W.-K., Park, K. I., & Ma, J. Y. (2022). Protective effect of Acer palmatum Thunb. leaf extract on mice with steroid-induced ocular hypertension. Molecular & Cellular Toxicology, 18(1), 71–79. https://doi.org/10.1007/s13273-021-00173-2
Park, I., Yang, S., Kim, W. J., Noh, P., Lee, H. O., & Moon, B. C. (2018). Authentication of herbal medicines Dipsacus asper and Phlomoides umbrosa using DNA barcodes, chloroplast genome, and sequence characterized amplified region (SCAR) marker. Molecules, 23, 1748.
Al-Qurainy, F., Al-Ameri, A. A., Khan, S., Nadeem, M., Gaafar, A.-R.Z., & Tarroum, M. (2018). SCAR marker for gender identification in date palm (Phoenix dactylifera L.) at the seedling stage. International Journal of Genomics. https://doi.org/10.1155/2018/3035406
Yin, J., Wang, Q., Wang, Y., Li, Y., Zeng, W., Wu, J., Ren, Y., Tang, Y., Gao, C., & Hu, H. (2019). Development of a simple and rapid reverse transcription–loopmediated isothermal amplification (RT-LAMP) assay for sensitive detection of tilapia lake virus. Journal of Fish Diseases, 42, 817–824.
Acknowledgements
This work was supported by Korea Institute of Planning and Evaluation for Technology in Food, Agriculture and Forestry (IPET) through Crop Viruses and Pests Response Industry Technology Development Program, funded by Ministry of Agriculture, Food and Rural Affairs (MAFRA) (321108-04) and National Research Foundation of Korea (NRF) funded by the Ministry of Education (2020R1A6A1A06046235). This work was partially supported by a funding for the academic research program of Chungbuk National University in 2022.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
None of the authors have any relevant financial relationships to disclose.
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lee, JP., Woo, JA., Shin, WR. et al. Distinction of Male and Female Trees of Ginkgo biloba Using LAMP. Mol Biotechnol 65, 1693–1703 (2023). https://doi.org/10.1007/s12033-023-00673-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-023-00673-7