Abstract
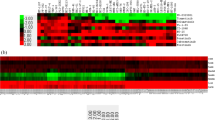
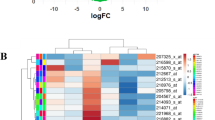
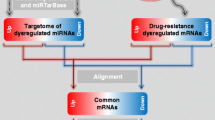
Cyclophosphamide (CP), an important alkylating agent which is used in the treatment therapy for chronic myeloid leukemia (CML). However, acquired drug resistance owing to the inactivation of its active metabolite aldophosphamide via tumoral-overexpressing aldehyde dehydrogenase (ALDH1A1) is one of the major issues with the CP therapy. However, the underlying mechanism of ALDH1A1 overexpression in cancer cells remains poorly defined. Therefore, the current study focused on analyzing the ALDH1A1-overexpressing microarray data for CP resistance and CP-sensitive CML cell lines. In this study, the microarray dataset was obtained from Gene Expression Omnibus GEO. The GEO2R tool was used to identify Differentially Expressing Genes (DEGs). Further, protein–protein interaction (PPI) network of DEGs were constructed using STRING database. Finally, Hub gene-miRNA-TFs interaction were constructed using miRNet tool. A total of 749 DEGs including 387 upregulated and 225 downregulated genes were identified from this pool of microarray data. The construction of DEGs network resulted in identification of three genes including ZEB2, EZH2, and MUC1 were found to be majorly responsible for ALDH1A1 overexpression. miRNA analysis identified that, hsa-mir-16-5p and hsa-mir-26a-5p as hub miRNA which are commonly interacting with maximum target genes. Additionally, drug-gene interaction analysis was performed to identify drugs which are responsible for ALDH1A1 expression. The entire study may provide a deeper understanding about ALDH1A1 regulatory genes responsible for its overexpression in CP resistance cancer. This understanding may be further explore for developing possible co-therapy to avoid the ALDH1A1-mediated CP resistance.





Similar content being viewed by others
References
Arrigoni E, Del Re M, Galimberti S, Restante G, Rofi E, Crucitta S, Baratè C, Petrini M, Danesi R, Di Paolo A. Concise review: chronic myeloid leukemia: stem cell niche and response to pharmacologic treatment. Stem Cells Transl Med. 2018. https://doi.org/10.1002/sctm.17-0175.
Jabbour E, Kantarjian H. Chronic myeloid leukemia: 2020 update on diagnosis, therapy and monitoring. Am J Hematol. 2020. https://doi.org/10.1002/ajh.25792.
Cortes JE, Talpaz M, Kantarjian H. Chronic myelogenous leukemia: a review. Am J Med. 1996. https://doi.org/10.1016/s0002-9343(96)00061-7.
Palejwala AH, O’Connor KP, Shi H, Villeneuve L, Scordino T, Glenn CA. Chronic myeloid leukemia manifested as myeloid sarcoma: review of literature and case report. J Clin Neurosci. 2019. https://doi.org/10.1016/j.jocn.2019.04.011.
Hochhaus A, Breccia M, Saglio G, García-Gutiérrez V, Réa D, Janssen J, Apperley J. Expert opinion-management of chronic myeloid leukemia after resistance to second-generation tyrosine kinase inhibitors. Leukemia. 2020. https://doi.org/10.1038/s41375-020-0842-9.
García-Gutiérrez V, Hernández-Boluda JC. Tyrosine kinase inhibitors available for chronic myeloid leukemia: efficacy and safety. Front Oncol. 2019. https://doi.org/10.3389/fonc.2019.00603.
Carofiglio F, Lopalco A, Lopedota A, Cutrignelli A, Nicolotti O, Denora N, Stefanachi A, Leonetti F. Bcr-Abl tyrosine kinase inhibitors in the treatment of pediatric CML. Int J Mol Sci. 2020. https://doi.org/10.3390/ijms21124469.
Kong JH, Winton EF, Heffner LT, Gaddh M, Hill B, Neely J, Hatcher A, Joseph M, Arellano M, El-Rassi F, Kim A. Outcomes of chronic phase chronic myeloid leukemia after treatment with multiple tyrosine kinase inhibitors. J Clin Med. 2020. https://doi.org/10.3390/jcm9051542.
Biggs JC, Szer J, Crilley P, Atkinson K, Downs K, Dodds A, Concannon AJ, Avalos B, Tutschka P, Kapoor N. Treatment of chronic myeloid leukemia with allogeneic bone marrow transplantation after preparation with BuCy2. Blood. 1992;80:1352–7.
Mueller MC, Cervantes F, Hjorth-Hansen H, Janssen JJ, Milojkovic D, Rea D, Rosti G. Ponatinib in chronic myeloid leukemia (CML): Consensus on patient treatment and management from a European expert panel. Crit Rev Oncol Hematol. 2017. https://doi.org/10.1016/j.critrevonc.2017.10.002.
Andersson BS, Mroue M, Britten RA, Farquhar D, Murray D. Mechanisms of cyclophosphamide resistance in a human myeloid leukemia cell line. Acta Oncol. 1995. https://doi.org/10.3109/02841869509093963.
Mansoori B, Mohammadi A, Davudian S, Shirjang S, Baradaran B. The different mechanisms of cancer drug resistance: a brief review. Adv Pharm Bull. 2017. https://doi.org/10.15171/apb.2017.041.
Zhang J, Tian Q, Yung Chan S, Chuen Li S, Zhou S, Duan W, Zhu YZ. Metabolism and transport of oxazaphosphorines and the clinical implications. Drug Metab Rev. 2005. https://doi.org/10.1080/03602530500364023.
Gerber JM, Qin L, Kowalski J, Smith BD, Griffin CA, Vala MS, Collector MI, Perkins B, Zahurak M, Matsui W, Gocke CD. Characterization of chronic myeloid leukemia stem cells. Am J Hematol. 2011. https://doi.org/10.1002/ajh.21915.
He X, Deng Y, Yue W. Investigating critical genes and gene interaction networks that mediate cyclophosphamide sensitivity in chronic myelogenous leukemia. Mol Med Rep. 2017. https://doi.org/10.3892/mmr.2017.6636.
Verma H, Silakari O. Investigating the Role of Missense SNPs on ALDH 1A1 mediated pharmacokinetic resistance to cyclophosphamide. Biol Med. 2020. https://doi.org/10.1016/j.compbiomed.2020.103979.
Hobert O. Gene regulation by transcription factors and microRNAs. Science. 2008. https://doi.org/10.1126/science.1151651.
Liu L, Cai S, Han C, Banerjee A, Wu D, Cui T, Xie G, Zhang J, Zhang X, McLaughlin E, Yin M. ALDH1A1 contributes to PARP inhibitor resistance via enhancing DNA repair in BRCA2−/− ovarian cancer cells. Mol Cancer Ther. 2020. https://doi.org/10.1158/1535-7163.MCT-19-0242.
Clough E, Barrett T. The gene expression omnibus database. 2016. Stat Genom. https://doi.org/10.1007/978-1-4939-3578-9_5.
Bao F, Polk P, Nordberg ML, Veillon DM, Sun A, Deininger M, Murray D, Andersson BS, Munker R. Comparative gene expression analysis of a chronic myelogenous leukemia cell line resistant to cyclophosphamide using oligonucleotide arrays and response to tyrosine kinase inhibitors. Leuk Res. 2007. https://doi.org/10.1016/j.leukres.2007.03.002.
Barrett T, Wilhite SE, Ledoux P, Evangelista C, Kim IF, Tomashevsky M, Marshall KA, Phillippy KH, Sherman PM, Holko M, Yefanov A. NCBI GEO: archive for functional genomics data sets-update. Nucleic Acids Res. 2012;41:D991–5.
Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc B. 1995. https://doi.org/10.2307/2346101.
Xu Z, Zhou Y, Cao Y, Dinh TLA, Wan J, Zhao M. Identification of candidate biomarkers and analysis of prognostic values in ovarian cancer by integrated bioinformatics analysis. J R Stat Soc B. 1995. https://doi.org/10.2307/2346101.
Szklarczyk D, Morris JH, Cook H, Kuhn M, Wyder S, Simonovic M, Santos A, Doncheva NT, Roth A, Bork P, Jensen LJ. The STRING database in 2017: quality-controlled protein–protein association networks, made broadly accessible. Nucleic Acids Res. 2016. https://doi.org/10.1093/nar/gkw937.
Otasek D, Morris JH, Bouças J, Pico AR, Demchak B. Cytoscape automation: empowering workflow-based network analysis. Genome Biol. 2019. https://doi.org/10.1186/s13059-019-1758-4.
Chang L, Zhou G, Soufan O, Xia J. miRNet 2.0: network-based visual analytics for miRNA functional analysis and systems biology. Nucleic Acids Res. 2020. https://doi.org/10.1093/nar/gkaa467.
Davis AP, Grondin CJ, Johnson RJ, Sciaky D, Wiegers J, Mattingly WTC, CJ. . Comparative toxicogenomics database (CTD): update. Nucleic Acids Res. 2021. https://doi.org/10.1093/nar/gkaa891.
Fleming RA. An overview of cyclophosphamide and ifosfamide pharmacology. Pharmacotherapy. 1997. https://doi.org/10.1002/j.1875-9114.1997.tb03817.x.
Verma H, Singh Bahia M, Choudhary S, Kumar Singh P, Silakari O. Drug metabolizing enzymes-associated chemo resistance and strategies to overcome it. Drug Metab Rev. 2019. https://doi.org/10.1080/03602532.2019.1632886.
Alam M, Rajabi H, Ahmad R, Jin C, Kufe D. Targeting the MUC1-C oncoprotein inhibits self-renewal capacity of breast cancer cells. Oncotarget. 2014. https://doi.org/10.18632/oncotarget.1848.
Wang R, Yang L, Li S, Ye D, Yang L, Liu Q, Zhao Z, Cai Q, Tan J, Li X. Quercetin inhibits breast cancer stem cells via downregulation of aldehyde dehydrogenase 1A1 (ALDH1A1), chemokine receptor type 4 (CXCR4), mucin 1 (MUC1), and epithelial cell adhesion molecule (EpCAM). Med Sci Monit. 2018;24:412–20. https://doi.org/10.12659/msm.908022.
Li H, Bitler BG, Vathipadiekal V, Maradeo ME, Slifker M, Creasy CL, Tummino PJ, Cairns P, Birrer MJ, Zhang R. ALDH1A1 is a novel EZH2 target gene in epithelial ovarian cancer identified by genome-wide approaches. Cancer Prev Res. 2012. https://doi.org/10.1158/1940-6207.CAPR-11-0414.
Gorodetska I, Lukiyanchuk V, Peitzsch C, Kozeretska I, Dubrovska A. BRCA1 and EZH2 cooperate in regulation of prostate cancer stem cell phenotype. Int J Cancer. 2019. https://doi.org/10.1002/ijc.32323.
Peitzsch C, Cojoc M, Hein L, Kurth I, Mäbert K, Trautmann F, Klink B, Schröck E, Wirth MP, Krause M, Stakhovsky EA. An epigenetic reprogramming strategy to resensitize radioresistant prostate cancer cells. Cancer Res. 2016. https://doi.org/10.1158/0008-5472.CAN-15-2116.
Li Q, Liu KY, Liu Q, Wang G, Jiang W, Meng Q, Yi Y, Yang Y, Wang R, Zhu S, Li C. Antihistamine drug ebastine inhibits cancer growth by targeting polycomb group protein EZH2. Mol Cancer Ther. 2020. https://doi.org/10.1158/1535-7163.MCT-20-0250.
Momparler RL, Côté S. Targeting of cancer stem cells by inhibitors of DNA and histone methylation. Expert Opin Investig Drugs. 2015. https://doi.org/10.1517/13543784.2015.1051220.
Balasubramanian V, Iyer P, Arora S, Troyer P, Normant E. CPI-169, a novel and potent EZH2 inhibitor, synergizes with CHOP in vivo and achieves complete regression in lymphoma xenograft models. AACR. 2014. https://doi.org/10.1158/1538-7445.AM2014-1697.
Yan Y, He M, Yu Z, Sun M, Zhao L, Zhao H, Yao W, Wei M. Combined expression of ZEB2 and ALDH1A1 is correlated with poor prognosis of breast cancer patients. Int J Clin Exp Med. 2018;11:1994–2003.
Colacino JA, Azizi E, Brooks MD, Harouaka R, Fouladdel S, McDermott SP, Lee M, Hill D, Madden J, Boerner J, Cote ML. Heterogeneity of human breast stem and progenitor cells as revealed by transcriptional profiling. BioRxiv. 2017. https://doi.org/10.1101/109751.
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–97.
Liu T, Cai J, Cai J, Wang Z, Cai L. EZH2-miRNA positive feedback promotes tumor growth in ovarian cancer. Front Oncol. 2021. https://doi.org/10.3389/fonc.2020.608393.
Vishnubalaji R, Hamam R, Abdulla MH, Mohammed MAV, Kassem M, Al-Obeed O, Aldahmash A, Alajez NM. Genome-wide mRNA and miRNA expression profiling reveal multiple regulatory networks in colorectal cancer. Cell Death Dis. 2015. https://doi.org/10.1038/cddis.2014.556.
Reza G, Neda M, Jahanbakhsh A, Nazila N, Seyed Javad M, Yaghoub Y, Ashraf M, Hossein P, Sakari K, Reza M. Downregulation of plasma MiR-142–3p and MiR-26a-5p in patients with colorectal carcinoma. Iran J Cancer Prev. 2015. https://doi.org/10.17795/ijcp2329.
Navid F, Baker SD, McCarville MB, Stewart CF, Billups CA, Wu J, Davidoff AM, Spunt SL, Furman WL, McGregor LM, Hu S. Phase I and clinical pharmacology study of bevacizumab, sorafenib, and low-dose cyclophosphamide in children and young adults with refractory/recurrent solid tumors. Clin Cancer Res. 2013. https://doi.org/10.1158/1078-0432.CCR-12-1897.
Ye Y, Zhang S, Chen Y, Wang X, Wang P. High ALDH1A1 expression indicates a poor prognosis in gastric neuroendocrine carcinoma. Pathol Res Pract. 2018. https://doi.org/10.1016/j.prp.2017.10.015.
Fukuda M, Yamaguchi S, Ohta T, Nakayama Y, Ogata H, Shimizu K, Nishikawa T, Adachi Y, Fukuma E. Combination therapy for advanced breast cancer: cyclophosphamide, doxorubicin, UFT, and tamoxifen. Oncology. 1999;13:77–81.
Vishnubalaji R, Manikandan M, Fahad M, Hamam R, Alfayez M, Kassem M, Aldahmash A, Alajez NM. Molecular profiling of ALDH1+ colorectal cancer stem cells reveals preferential activation of MAPK, FAK, and oxidative stress pro-survival signalling pathways. Oncotarget. 2018. https://doi.org/10.18632/oncotarget.24420.
Funding
This work was supported by the Indian Council of Medical Research (ICMR), New Delhi; under sanction No. ISRM/12(10)/2019.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Narendra, G., Raju, B., Verma, H. et al. Identification of potential genes associated with ALDH1A1 overexpression and cyclophosphamide resistance in chronic myelogenous leukemia using network analysis. Med Oncol 38, 123 (2021). https://doi.org/10.1007/s12032-021-01569-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-021-01569-9